PHF6 loss will increase leukemia initiating cell frequency in Hoxa9-driven AML
To find out the prognostic significance of PHF6 mutations in human AML, we used publicly out there mutational and survival knowledge from the BEAT AML dataset [26]. Of 805 AML sufferers, 22 (2.7%) had PHF6 mutations. PHF6 mutations had been related to lowered total survival in opposed threat sufferers (Fig. 1A) PHF6 gene mutational lessons (frameshift, nonsense, missense) confirmed comparable poor survival curves relative to one another (Fig. S1A–C).
A Kaplan-Meier survival curve for PHF6 mutated and unmutated opposed threat grownup AML sufferers (ELN classification) from the BEAT AML dataset. B Experimental design for Hoxa9 retroviral transduction of Ctrl and cKO marrow, adopted by colony forming unit assay (CFU), AML induction in mice, and serial transplantation of reworked leukemic cells. C Bar graph displaying variety of colony forming items (CFUs) obtained from 8 rounds of serial methylcellulose replating of 500 cells/plate of Ctrl+Hoxa9 and cKO+Hoxa9 reworked mouse bone marrow. (n = 6–12 organic replicates). D Left Bar graph displaying common variety of cells per colony (colony measurement) obtained after 7 rounds of serial methylcellulose replating of 500 cells/plate of Ctrl+Hoxa9 and cKO+Hoxa9 reworked mouse bone marrow. (n = 6–12 organic replicates) Proper Consultant images of colonies on the third plating. Scale bar represents 500 μm. E Kaplan–Meier survival curves of Ctrl+Hoxa9 and cKO+Hoxa9 major transplant recipients receiving 400 Ok, 100 Ok, or 30 Ok GFP+ cells. (n = 7–21 mice per cohort). F Kaplan–Meier survival curve of Ctrl+Hoxa9 and cKO+Hoxa9 secondary transplant recipients, receiving 100 Ok or 30 Ok GFP+ cells harvested from bone marrow of major recipients 8 weeks after transplantation. (n = 7–10 mice per cohort). G Kaplan–Meier survival curve of Ctrl+Hoxa9 and cKO+Hoxa9 tertiary transplant recipients, receiving 100 Ok GFP+ cells harvested from bone marrow of secondary recipients 8 weeks after transplantation. (n = 7–11 mice per cohort). H, I Limiting dilution evaluation for LIC calculation for (H) freshly Hoxa9-transduced cells, (I) 8 weeks major transplanted leukemic cells. All bar graphs present imply ± SEM and statistical significance was calculated utilizing the Scholar t-test. For all survival curves, statistical significance was calculated utilizing the Log-rank (Mantel-Cox) take a look at. *p < 0.05, **p < 0.01, ***p < 0.001; ****p < 0.0001, ns = not important.
To evaluate the function of PHF6 loss in mouse AML, we generated conditional hematopoietic Phf6 knockout Vav-CreCre/+Phf6fl/y (cKO) mice (Fig. S1D) and in contrast them to their Vav-CreCre/+Phf6+/y (Ctrl) littermates. Revealed research of hematopoietic Phf6 loss have reported no proof of leukemic transformation [17,18,19, 27]. We additionally discovered no blood depend abnormalities in cKO mice as much as 9 months (Fig. S1E–Ok), additional indicating that Phf6 loss alone is probably going inadequate to provoke leukemia.
We subsequent induced AML utilizing the Hoxa9 retroviral transduction mannequin. We picked this mannequin as a result of its skill to provide AML with a comparatively longer latency (lethality in ~3–6 months) [24], permitting us to check a possible function for Phf6 loss in accelerating AML kinetics. We transduced complete bone marrow from 5-FU-treated Ctrl and cKO mice with MSCV Hoxa9–IRES-GFP retrovirus, and investigated the impact of Phf6 loss on the flexibility of Hoxa9-transformed cells to type colonies in methylcellulose (Fig. 1B). We noticed that Ctrl+Hoxa9 cells had been practically exhausted after 4 platings, whereas cKO+Hoxa9 cells demonstrated persistent colony-forming skill as much as 8 platings (Fig. 1C), with bigger colonies (Fig. 1D). Thus, Phf6 loss provides a replating benefit to Hoxa9-transformed marrow and delays its in vitro exhaustion. Conversely, MLL-AF9–IRES-GFP reworked marrow confirmed no proof of exhaustion with replating, and no proof of additional acceleration with Phf6 loss (Fig. S2A, B). On condition that 70% of human AMLs present excessive HOXA9 ranges [25], we deemed the Hoxa9 overexpression mouse mannequin as being broadly related to human leukemia biology, versus MLL-AF9, which doesn’t co-occur with PHF6 mutation in people. We subsequently proceeded with the Hoxa9-driven mannequin for our research.
To check the function of Phf6 loss within the growth of AML in vivo, we transplanted Hoxa9-transduced marrow into lethally irradiated syngeneic recipients (Fig. 1B). We confirmed that Ctrl+Hoxa9 marrow produced lethality in recipient mice in ~3–5 months after transplantation (Fig. 1E) as a result of AML characterised by >20% blasts within the marrow (Fig. S2C), peripheral leukocytosis (Fig. S2D), splenomegaly with effacement of splenic structure (Fig. S2E), and infiltration of leukemic cells within the liver (Fig. S2F). Survival was comparable between Ctrl+Hoxa9 and cKO+Hoxa9 teams in major recipients transplanted with a number of doses (400 Ok, 100 Ok, or 30 Ok cells) (Fig. 1E), with comparable levels of leukemic infiltrate (Fig. S3A–F) and splenomegaly at morbidity (Fig. S3G). Nevertheless, secondary and tertiary transplantation of marrow confirmed progressively accelerated lethality in cKO+Hoxa9 in comparison with Ctrl+Hoxa9 (Fig. 1F, G, S3H). Thus, Phf6 loss accelerates Hoxa9-pushed mouse AML on serial transplantation.
We subsequent sought to find out the impact of Phf6 loss on the frequency of leukemia initiating cells (LIC), the sub-population of reworked marrow able to initiating leukemia. We carried out limiting dilution transplantation assays (LD) on freshly transduced marrow (GFP+ cells sorted 2 days after retroviral transduction) in addition to on marrow from recipients (GFP+ bone marrow cells sorted from recipients 8 weeks after major transplantation). We picked the 8-week time level primarily based on the initiation of lethality on this mannequin at ~12 weeks (Fig. 1E). LD of freshly transduced marrow confirmed that, at baseline, cKO+Hoxa9 cells had a 2-fold larger frequency of cells able to leukemic transformation (Fig. 1H). LD on marrow extracted 8 weeks post-transplantation confirmed an roughly 5-fold larger frequency of leukemia initiating cells (LICs) in cKO+Hoxa9 marrow (Fig. 1I). Thus, Phf6 loss will increase LIC frequency in Hoxa9-driven AML, with the rise occurring throughout in vivo AML evolution.
Phf6 loss will increase leukemic illness burden
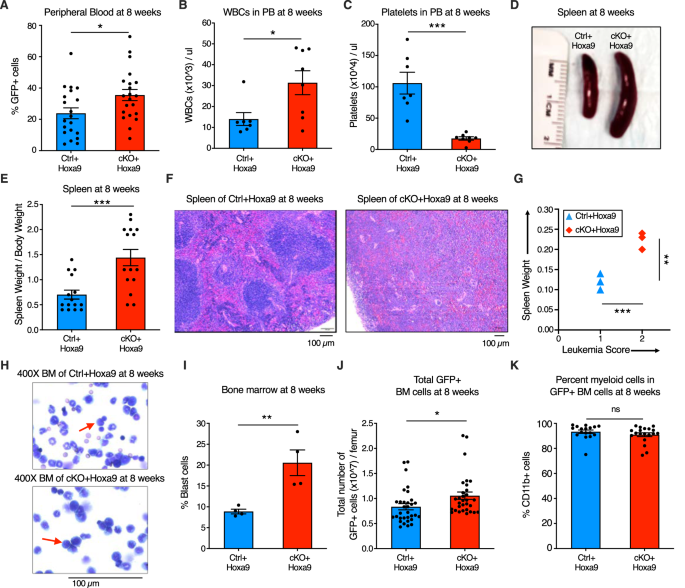
To characterize the impact of Phf6 loss additional, we analyzed peripheral blood, splenic structure, and bone marrow leukemic cell burden of major recipients at 8 weeks after transplantation. Mice transplanted with cKO+Hoxa9 cells confirmed a better frequency of GFP+ cells in peripheral blood at 8 weeks than mice receiving Ctrl+Hoxa9 cells (Fig. 2A). The cKO+Hoxa9 group additionally had larger leukocytosis (Fig. 2B) and extra extreme thrombocytopenia (Fig. 2C). Mice in each teams displayed comparable ranges of anemia (Fig. S4A, B). The cKO+Hoxa9 group had elevated spleen measurement and weight in comparison with the Ctrl+Hoxa9 group (Fig. 2D, E), and histopathological evaluation confirmed larger effacement of splenic structure (Fig. 2F). Splenic infiltration was quantified utilizing a beforehand described leukemia infiltration rating [28], and was discovered to be larger in cKO+Hoxa9 mice in comparison with Ctrl+Hoxa9 (Fig. 2G). Giemsa-stained cytospin preparations confirmed larger blast percentages in cKO+Hoxa9 on the 8-week timepoint (Fig. 2H, I), and circulation cytometry confirmed larger absolute and p.c GFP+ cells (Fig. 2J, S4C). All GFP+ cells had been myeloid for each teams (Fig. 2K, S4D). Thus, whereas mice from each teams succumbed at comparable instances after major transplant (Fig. 1F), analyses at matched time factors earlier than the onset of mortality revealed larger illness burden in cKO+Hoxa9 mice in comparison with Ctrl+Hoxa9.
A–C Bar graphs displaying peripheral blood evaluation at 8 weeks after transplantation of Ctrl+Hoxa9 and cKO+Hoxa9 cells. A Proportion of GFP+ cells in peripheral blood. B, C Counts of (B) WBCs and (C) platelets in peripheral blood. Regular vary for WBCs: 2000–10,000/µl. Regular vary for platelets: 900–1600 × 103/µl [41]. D Consultant {photograph} of spleens at 8 weeks after transplantation. Ruler depicts size in centimeters. E Bar graph displaying spleen weight (expressed as share of complete physique weight) of major recipients at 8 weeks after transplantation of Ctrl+Hoxa9 and cKO+Hoxa9 cells. F Consultant picture of H&E staining of spleen from Ctrl+Hoxa9 and cKO+Hoxa9 at 8 weeks after transplantation. Scale bar is 100 µm at 10X. G Spleen weight (y-axis) and leukemia rating (x-axis) from Ctrl+Hoxa9 and cKO+Hoxa9 major recipients at 8 weeks. X-axis represents a beforehand printed leukemia infiltration rating [28] calculated primarily based on splenic structure. Intact white and pink pulp was scored as 0, extramedullary hematopoiesis evident by aberrant cells in disturbed white pulp was scored as 1, whereas infiltration with leukemic blasts with excessive mitotic exercise was scored as 2. H Consultant picture of Wright–Giemsa staining of cytospin of Ctrl+Hoxa9 and cKO+Hoxa9 bone marrow cells at 8 weeks after transplantation. Scale bar is 100 µM at 400X. Purple arrows point out consultant blast cells. I Bar graph of share blast cells of complete nucleated cells in bone marrow at 8 weeks after transplant. J, Ok Bar graphs displaying (J) absolute variety of GFP+ cells in marrow and (Ok) share of CD11b+ myeloid cells amongst all GFP+ cells at 8 weeks after transplantation. All bar graphs present imply ± SEM and statistical significance was calculated utilizing the Scholar t-test. For all survival curves, statistical significance was calculated utilizing the Log-rank (Mantel-Cox) take a look at. *p < 0.05, **p < 0.01, ***p < 0.001; ****p < 0.0001, ns = not important.
Phf6 loss will increase the frequency of self-renewing, transplantable LICs
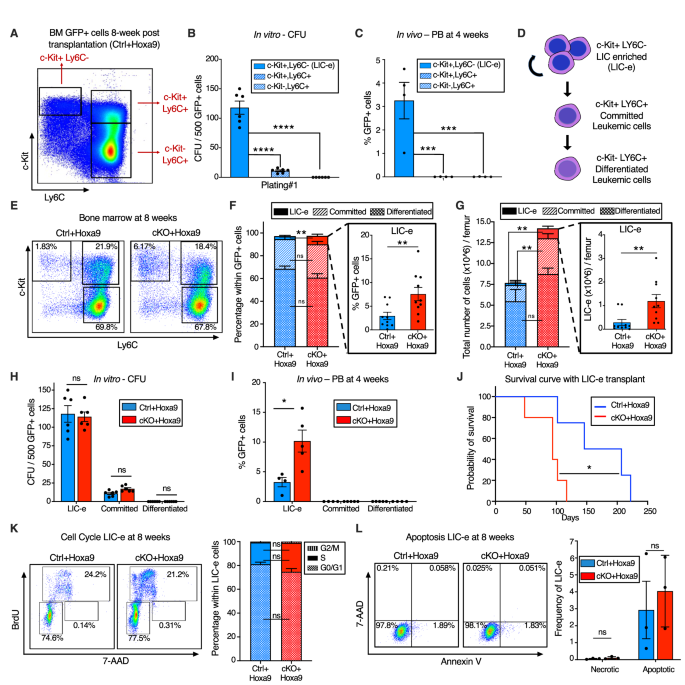
To characterize the immunophenotype of AML subpopulations (together with LICs), we additional analyzed the marrow of Ctrl+Hoxa9 recipients at 8 weeks after transplantation. The GFP+ cells didn’t specific B or T cell markers (Fig. S5A). Immature AML cells are recognized to have excessive c-Package expression [29], and leukemic stem cells (LSCs) within the MLL-AF9 retroviral mouse mannequin aberrantly specific mature myeloid lineage antigens comparable to Ly6C and CD11b [30]. To determine the corresponding subpopulation containing LICs in Hoxa9-only-driven AML, and to characterize the differentiation hierarchy of this mannequin, we settled on a technique utilizing c-Package and Ly6C expression to divide GFP+ marrow cells into three populations: (i) cKit+ Ly6C-, (ii) c-Package+ Ly6C+, and (iii) c-Package- Ly6C+ (Fig. 3A, S5B). The inhabitants on the prime of the hierarchy was the cKit+ Ly6C- inhabitants, an immature inhabitants with expression of cKit, CD34, and dim CD11b, with no expression of Ly6C, Ly6G, or Sca-1, and blended expression of CD16/32 (Fig. S5C). This inhabitants was able to giving rise to extra differentiated Ly6C+ cells inside 2 days of tradition (Fig. S5D), may produce colonies on methylcellulose plating (Fig. 3B), and will engraft into recipient mice (Fig. 3C). Based mostly on this subpopulation’s skill to engraft, however cognizant that not all cells inside it are LICs, we termed it the ‘LIC enriched’ (LIC-e) inhabitants (Fig. 3D). The second inhabitants was the c-Package+ Ly6C+ inhabitants, additionally expressing CD11b, CD34, and CD16/32, however not Sca-1 (Fig. S5C). On tradition, this inhabitants may solely give rise to Ly6C+ cells, however to not any Ly6C- cells (Fig. S5D), indicating that it’s irreversibly dedicated to differentiation. This inhabitants may produce a small variety of colonies in methylcellulose (Fig. 3B), however couldn’t engraft mice (Fig. 3C). We termed it the ‘dedicated’ leukemic cell inhabitants (Fig. 3D). The third inhabitants was the c-Package- Ly6C+ inhabitants, which expressed Ly6C, CD11b, CD34, and CD16/32, however not Sca-1, and had blended expression of Ly6G (Fig. S5C). It couldn’t produce any colonies (Fig. 3B) nor engraft (Fig. 3C). In vitro, this inhabitants solely gave rise to Ly6C+ cells (Fig. S5D). We termed it the “differentiated” leukemic cell inhabitants (Fig. 3D). We thus established the hierarchical group of Hoxa9-driven AML, and used it to guage the consequences of Phf6 loss.
A Consultant circulation cytometry plot of bone marrow GFP+ cells at 8 weeks after transplantation with Ctrl+Hoxa9 cells, compartmentalized into three subpopulations (i) LIC-e, (ii) Dedicated, and (iii) Differentiated leukemic cells utilizing c-Package and Ly6C expression. Observe: The identical circulation cytometry plot has been proven in Fig. S5 with detailed immunophenotypic markers. B Bar graph displaying variety of colony forming items (CFUs) obtained on methylcellulose plating of 500 cells of sorted subpopulations of Ctrl+Hoxa9 transplanted marrow. (n = 6 organic replicates). C Bar graph displaying frequencies of GFP+ cells in peripheral blood of secondary recipient mice at 4 weeks after transplantation of sorted subpopulations from Ctrl+Hoxa9 major recipient marrow. (n = 4–5 organic replicates). D Schematic of hierarchical group of leukemic cells (LIC-e, dedicated, and differentiated leukemic cells) inside AML produced by retroviral Hoxa9 transduction. E Consultant circulation cytometry plots depicting subpopulations of Ctrl+Hoxa9 and cKO+Hoxa9 leukemia marrow at 8 weeks after major transplant. F, G Stacked bar graphs displaying (F) frequencies, and (G) absolute quantity per femur of LIC-e, dedicated, and differentiated leukemic populations from Ctrl+Hoxa9 and cKO+Hoxa9 marrow at 8 weeks after transplantation. Insets present frequencies and absolute numbers of the LIC-e subpopulation. (n = 10–11). H Bar graph displaying variety of CFUs obtained on methylcellulose plating of 500 cells of sorted LIC-e, dedicated, and differentiated leukemic populations from Ctrl+Hoxa9 and cKO+Hoxa9 major recipient bone marrow at 8 weeks after transplantation. (n = 6 organic replicates). I Bar graph displaying frequencies of GFP+ cells within the peripheral blood of secondary recipient mice at 4 weeks after transplantation with sorted LIC-e, dedicated, and differentiated leukemic cell subpopulations from Ctrl+Hoxa9 and cKO+Hoxa9 major recipient bone marrow at 8 weeks after transplantation. (n = 5 organic replicates). Observe: The Ctrl+Hoxa9 samples in (H) and (I) are the identical as these depicted in (B) and (C)—the present figures are evaluating Ctrl and cKO. J Kaplan–Meier curve of secondary transplant recipients receiving 50 Ok sorted LIC-e cells from Ctrl+Hoxa9 and cKO+Hoxa9 major mouse bone marrow at 8 weeks after transplantation. (n = 4–5 organic replicates). Ok Left Consultant circulation cytometry plots for cell cycle evaluation of LIC-e cells, with BrdU (marking cells in S part) and 7-AAD (marking DNA) 2 h after BrdU injection into dwell mice at 8 weeks after transplantation. Proper Stacked bar graph signifies frequencies of in vivo LIC-e cells in G0/G1, S, and G2/M phases. (n = 13 organic replicates). L Left Consultant circulation cytometry plots of apoptotic (Annexin V+, 7AAD−), and necrotic (7AAD+) LIC-e cells. Proper Bar graph reveals frequencies of apoptotic and necrotic LIC-e cells from Ctrl+Hoxa9 and cKO+Hoxa9 major recipient bone marrow at 8 weeks after transplantation. All bar graphs present imply ± SEM and statistical significance was calculated utilizing the Scholar t-test. For all survival curves, statistical significance was calculated utilizing the Log-rank (Mantel-Cox) take a look at. *p < 0.05, **p < 0.01, ***p < 0.001; ****p < 0.0001, ns = non important.
The LIC-e inhabitants, although comprising a small minority of GFP+ cells, was expanded in 8-week cKO+Hoxa9 marrow in comparison with Ctrl+Hoxa9, whereas the relative fractions of dedicated and differentiated leukemic cells had been comparable (Fig. 3E, F). The distinction within the LIC-e inhabitants was much more pronounced when absolute cell numbers had been thought of, displaying a 5-fold enhance in cKO+Hoxa9 (Fig. 3G). To find out useful variations between Ctrl+Hoxa9 and cKO+Hoxa9 subpopulations, we sorted equal numbers of cells from every subpopulation at 8 weeks after transplant and carried out methylcellulose tradition and secondary transplantation into irradiated recipients. Dedicated and differentiated leukemic cells of both group fashioned few to no colonies, whereas LIC-e cells confirmed comparable colony-forming skill (Fig. 3H). cKO+Hoxa9 LIC-e cells confirmed larger engraftment in secondary recipients at 4 weeks (Fig. 3I) and led to extra speedy lethality than Ctrl+Hoxa9 LIC-e cells (Fig. 3J). We didn’t observe any distinction in cell cycle distribution or apoptosis of LIC-e cells (Fig. 3K, L). Thus, Phf6 loss led to an expanded and extra transplantable LIC-enriched AML subpopulation whose enhanced leukemic potential was not defined by variations in cell cycle or apoptosis.
Phf6 loss promotes a stemness gene community
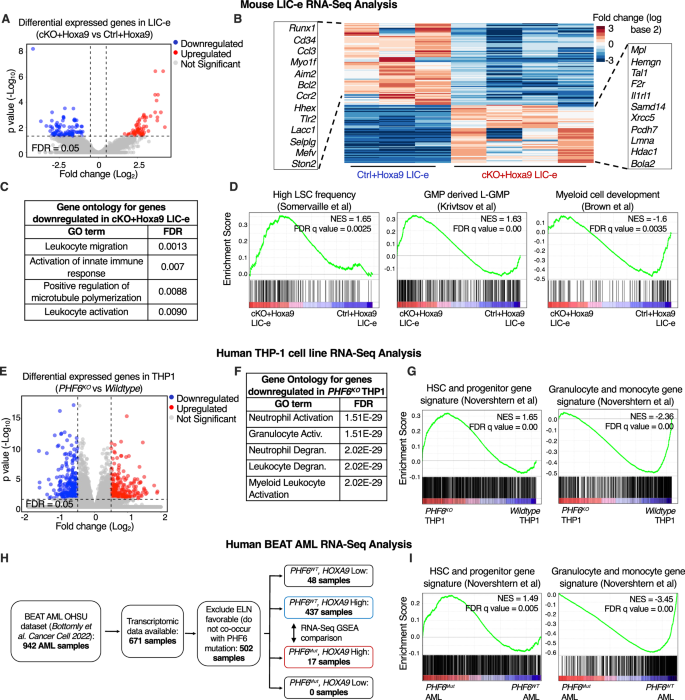
We decided the transcriptional penalties of Phf6 loss on Hoxa9-transformed marrow by performing RNA-Seq on LIC-e and dedicated leukemic cells from marrow of transplanted recipients at 8 weeks. Dedicated leukemic cells confirmed no change in gene expression with Phf6 loss (Fig. S6A, B, Desk S2), whereas the LIC-e inhabitants confirmed 91 downregulated and 65 upregulated genes in cKO+Hoxa9 in comparison with Ctrl+Hoxa9 (Fig. 4A, B, Desk S2). Genes downregulated in cKO+Hoxa9 LIC-e cells confirmed Gene Ontology (GO) enrichment for myeloid differentiation phrases (Fig. 4C). Gene set enrichment evaluation (GSEA) [31] confirmed that the cKO+Hoxa9 LIC-e transcriptome confirmed constructive enrichment for genesets associated to excessive LSC potential [32] and leukemic GMPs (L-GMPs) [33] and unfavorable enrichment for genesets associated to myeloid differentiation [34], and mature neutrophils and monocytes [35] (Fig. 4D, S6C).
A Volcano plot displaying differentially expressed genes in LIC-e cells from cKO+Hoxa9 in comparison with Ctrl+Hoxa9 bone marrow at 8 weeks after transplantation. (n = 3–4 organic replicates). B Heatmap of differential expression between Ctrl+Hoxa9 and cKO+Hoxa9 LIC-e cells. Insets present chosen downregulated (left) and upregulated (proper) genes in cKO+Hoxa9 LIC-e in contrast with Ctrl+Hoxa9 LIC-e. C High Gene Ontology phrases enriched in genes downregulated in cKO+Hoxa9 LIC-e in contrast with Ctrl+Hoxa9 LIC-e. D Gene set enrichment evaluation (GSEA) plots of the cKO+Hoxa9 LIC-e transcriptome in comparison with Ctrl+Hoxa9. Plots present constructive enrichment of gene units associated to excessive LSC frequency (left) and leukemic GMPs (center), and unfavorable enrichment of a gene set associated to myeloid growth (proper). Normalized Enrichment scores (NES) and FDR q values are proven. E Volcano plot displaying differentially expressed genes in PHF6KO THP-1 cells in comparison with wildtype THP1 cells. (n = 4–5 clonal traces). F High Gene Ontology phrases enriched in genes downregulated in PHF6KO THP-1 cells in comparison with wildtype THP1 cells. G Gene set enrichment evaluation (GSEA) plots of the PHF6KO THP-1 transcriptome in comparison with wildtype clones. Plots present constructive enrichment of gene units associated to excessive HSC and Progenitors (left), and unfavorable enrichment of a gene set associated to granulocytes and monocytes (proper). H Circulation chart depicting selection of BEAT AML dataset samples picked for RNA-Seq analyses, with exclusion of instances labeled as ELN favorable and people with low HOXA9 mRNA ranges (which present uncommon/no PHF6 mutations). I Gene set enrichment evaluation (GSEA) plots of the transcriptome of PHF6 mutated AML sufferers with excessive HOXA9 expression in comparison with PHF6 wildtype AML sufferers with excessive HOXA9 expression. Plots present constructive enrichment of gene units associated to excessive HSC and Progenitors (left), and unfavorable enrichment of a gene set associated to granulocytes and monocytes (proper).
We subsequent sought to find out whether or not PHF6 loss produces comparable transcriptional modifications in human AML. We used CRISPR to generate PHF6 knockout (PHF6KO) clones of the THP-1 human AML cell line (Fig. S6D). RNA-Seq confirmed that genes downregulated in PHF6KO in comparison with wildtype clones had been additionally enriched for myeloid differentiation GO phrases (Fig. 4E, F, Desk S3). PHF6KO clones additionally confirmed constructive GSEA enrichment of HSC and progenitor signatures, and unfavorable enrichment of granulocyte and monocyte signatures [36] (Fig. 4G). We additional analyzed publicly out there RNA-Seq knowledge from the BEAT AML dataset [26]. A majority of samples had excessive HOXA9 mRNA ranges, and PHF6 mutated instances solely fell throughout the HOXA9 excessive (HOXA9Excessive) group (Fig. 4H). On evaluating the transcriptomes of PHF6 mutant (PHF6Mut) to PHF6 wildtype (PHF6WT) AMLs throughout the HOXA9Excessive group, we discovered comparable constructive GSEA enrichment of HSC and progenitor signatures, and unfavorable enrichment of granulocyte and monocyte signatures [36] (Fig. 4I). Thus, Phf6 loss or mutation in each mouse and human AML skews their transcriptomes to a extra stem-like and fewer differentiated state.
Phf6 loss prevents exhaustion of LIC-e cells by sustaining their self-renewal potential
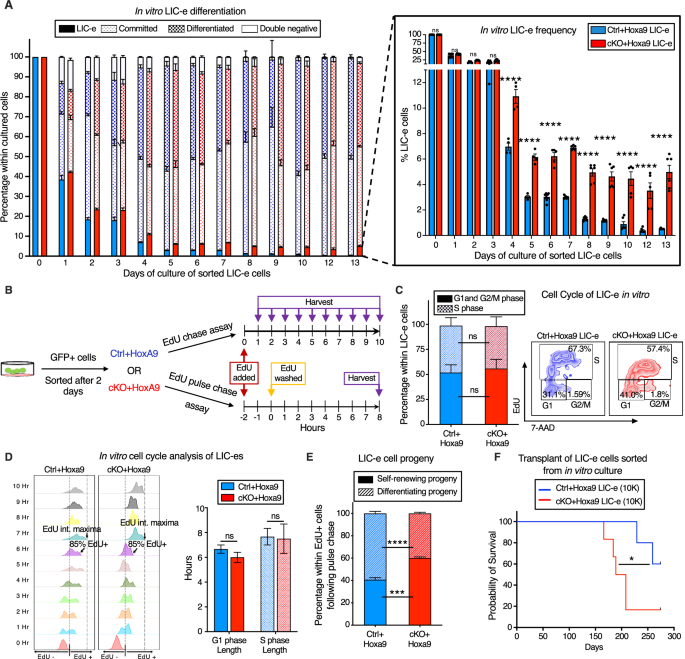
To find out the kinetics of the consequences of Phf6 loss on the habits of LIC-e cells, we cultured Hoxa9-transduced mouse bone marrow in cytokine-supplemented media. The expansion price of bulk tradition was comparable for Ctrl+Hoxa9 and cKO+Hoxa9 marrow (Fig. S7A). When sorted LIC-e cells had been cultured, most cells misplaced LIC-e identification inside days (Fig. 5A). Nevertheless, although each teams produced comparable fractions of dedicated (c-Package+ Ly6C+) and differentiated cells (c-Package- Ly6C+), the Ctrl+Hoxa9 tradition virtually fully depleted its LIC-e inhabitants (<1%), whereas the cKO+Hoxa9 tradition maintained this inhabitants, plateauing at 5–6% of the overall tradition after 5 days (Fig. 5A). Thus, Phf6 loss prevents exhaustion of the LIC-e inhabitants with out impairing the speed of proliferation or differentiation of the majority tradition, recapitulating the in vivo LIC-e accumulation phenotype proven earlier (Fig. 3E–G).
A Bar graph displaying frequencies of subpopulations ensuing from in vitro tradition of LIC-e cells sorted 4 days after Hoxa9 transduction of Ctrl and cKO bone marrow. Inset bar graph depicts solely LIC-e frequencies in the identical tradition. (n = 13 organic replicates). B Experimental design for research of in vitro cell cycle evaluation (prime) and self-renewal (backside) of LIC-e cells utilizing EdU chase and pulse-chase assay respectively. C Left, Bar graph displaying frequencies of G0/G1, S, and G2/M phases in Ctrl+Hoxa9 and cKO+Hoxa9 LIC-e cells in tradition 2 h after addition of EdU. (n = 4–5 organic replicates) Proper, Consultant circulation cytometry plots of identical, with EdU marking cells in S part and 7-AAD staining DNA. D Left, Consultant circulation cytometry plots displaying kinetics of uptake of EdU by Ctrl+Hoxa9 and cKO+Hoxa9 LIC-e cells over 10 h of tradition, carried out for calculation of cell cycle size. Time to complete inhabitants (>85%) EdU uptake represents G1 part size, and time to EdU depth maxima represents S part size. Proper, Bar graph represents the size of cell cycle phases of LIC-e cells in tradition. (n = 4–5 organic replicates) E Stacked bar graph displaying the share of self-renewing and differentiating progeny produced by Ctrl+Hoxa9 and cKO+Hoxa9 LIC-e cells in tradition. (n = 4–5 organic replicates). F Kaplan-Meier curve of major transplant recipients receiving 10 Ok sorted Ctrl+Hoxa9 and cKO+Hoxa9 LIC-e cells. (n = 5–6 organic replicates) All bar graphs present imply ± SEM and statistical significance was calculated utilizing the Scholar t-test. For all survival curves, statistical significance was calculated utilizing the Log-rank (Mantel-Cox) take a look at. *p < 0.05, **p < 0.01, ***p < 0.001; ****p < 0.0001, ns = non important.
To reconfirm that LIC-e accumulation wasn’t as a result of delicate biking variations, we carried out a 10-hour cell cycle evaluation in tradition by including EdU to sorted Ctrl+Hoxa9 and cKO+Hoxa9 LIC-e cells and harvesting them at serial time factors (Fig. 5B). We noticed that the distribution of Ctrl+Hoxa9 or cKO+Hoxa9 LIC-e cells in G1 and S phases confirmed no distinction initially of tradition, and confirmed no distinction within the kinetics or magnitude of EdU incorporation (Fig. 5C). Based mostly on beforehand printed rationale [37], we outlined the time required for 85% of every pattern to turn out to be EdU+ because the “G1 part size”, and the time required for every pattern to achieve a maximal EdU fluorescent depth because the ‘S part size’. We didn’t observe any distinction in G1 and S part lengths between Ctrl+Hoxa9 and cKO+Hoxa9 LIC-e cells (Fig. 5D). We subsequently hypothesized that cKO+Hoxa9 LIC-e cells, whereas able to producing differentiated progeny, might have a larger tendency to provide progeny with persistent LIC-e identification. To check this speculation, we carried out an EdU pulse-chase experiment by incubating sorted LIC-e cells with EdU for two hours (pulse), adopted by washing off the EdU and additional culturing cells for 8 extra hours (chase, for the size of S part) (Fig. 5B) to permit all cells that had been in S part throughout the preliminary EdU pulse to finish mitosis, so that each one EdU+ cells on the finish of the 8 hour EdU-free chase could be daughter cells/progeny of the unique EdU-uptaking cells. We decided the share of self-renewing progeny by calculating the share of complete progeny (complete EdU+ cells) that had LIC-e markers (Ly6C−EdU+), whereas the remainder (Ly6C+ EdU+) had been differentiating progeny. We noticed that whereas 40.6% of the progeny of Ctrl+Hoxa9 LIC-e cells had been additionally LIC-e cells, this fraction was elevated to 60.0% in cKO+Hoxa9 (Fig. 5E). To substantiate that in vitro LIC-e cells are functionally equal to their in vivo counterparts, we carried out major transplantation with 10 Ok LIC-e cells sorted after 2 days of tradition. Recipients of cKO+Hoxa9 LIC-e cells succumbed sooner than Ctrl+Hoxa9 LIC-e (Fig. 5F). Collectively, Phf6 loss prevents the exhaustion of LIC-e cells by rising the fraction of their progeny that retain persistent LIC-e identification.
Results of Phf6 loss on chromatin accessibility in LIC-e cells
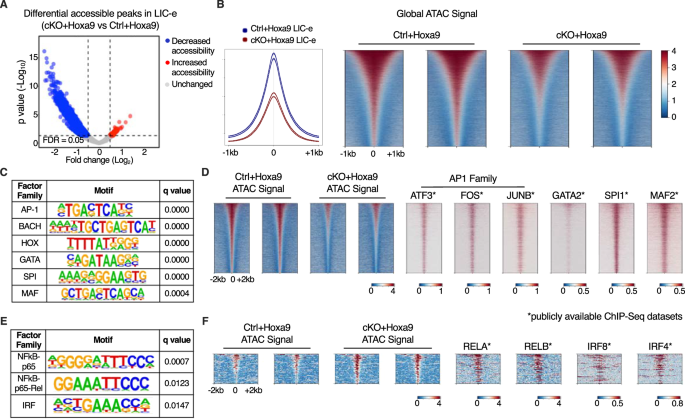
To profile the consequences of Phf6 loss on the accessibility panorama of LIC-e cells, we carried out ATAC-Seq on sorted LIC-e cells from freshly transduced cKO+Hoxa9 and Ctrl+Hoxa9 marrow. We noticed that cKO+Hoxa9 LIC-e cells confirmed a worldwide discount in chromatin accessibility in comparison with Ctrl+Hoxa9, with just a few areas displaying elevated accessibility (Fig. 6A, B). Areas that misplaced accessibility in cKO+Hoxa9 LIC-e cells confirmed enrichment for AP-1, HOX, SPI, and GATA household motifs, amongst others (Fig. 6C). Public ChIP-Seq tracks from leukemic or myeloid cells confirmed co-occupancy of those elements at ATAC peaks with lowered accessibility (Fig. 6D, Desk S4). Promoters of a number of genes like Runx1, Selplg, and Aim2, that are downregulated in cKO+Hoxa9 LIC-e cells (Fig. 4B), confirmed occupancy by these elements and confirmed lowered chromatin accessibility in cKO+Hoxa9 LIC-e cells (Fig. S7B). Conversely, the small variety of areas that gained accessibility in cKO+Hoxa9 LIC-e cells confirmed enrichment for NF-kB and IRF household motifs (Fig. 6E). Public ChIP-Seq tracks confirmed co-occupancy of NF-kB (RELA, RELB) and IRF8, IRF4 at areas of elevated accessibility (Fig. 6F, Desk S4). Total, Phf6 loss, probably through a mix of direct and oblique results, led to altered accessibility at websites sure by key hematopoietic transcription elements.
A Volcano plot displaying differentially accessible areas in LIC-e cells from cKO+Hoxa9 in contrast Ctrl+Hoxa9. (n = 3 organic replicates). B Consultant sign profile (left) and metagene plots (proper) displaying genome-wide depth of ATAC sign in Ctrl+Hoxa9 and cKO+Hoxa9 LIC-e cells. C HOMER evaluation for areas of decreased chromatin accessibility in cKO+Hoxa9 LIC-e cells, displaying enrichment of AP-1, HOX, GATA, SPI1, and MAF motifs. D Consultant metagene plots at areas of decreased chromatin accessibility in cKO+Hoxa9 LIC-e, present ChIP-Seq sign for choose proteins whose motifs are seen to be enriched by HOMER in (C). E HOMER evaluation for areas of elevated chromatin accessibility in cKO+Hoxa9 LIC-e cells displaying enrichment of NF-kB and IRF motifs. F Consultant metagene plots at areas of elevated chromatin accessibility in cKO+Hoxa9 LIC-e, present ChIP-Seq sign for choose proteins whose motifs are seen to be enriched by HOMER in (E). Publicly out there ChIP-Seq datasets in leukemia or myeloid cells had been utilized in metagene heatmaps (Desk S4). All plots had been centered round ATAC-Seq peaks. SeqPlots was used to attract all metagene plots.