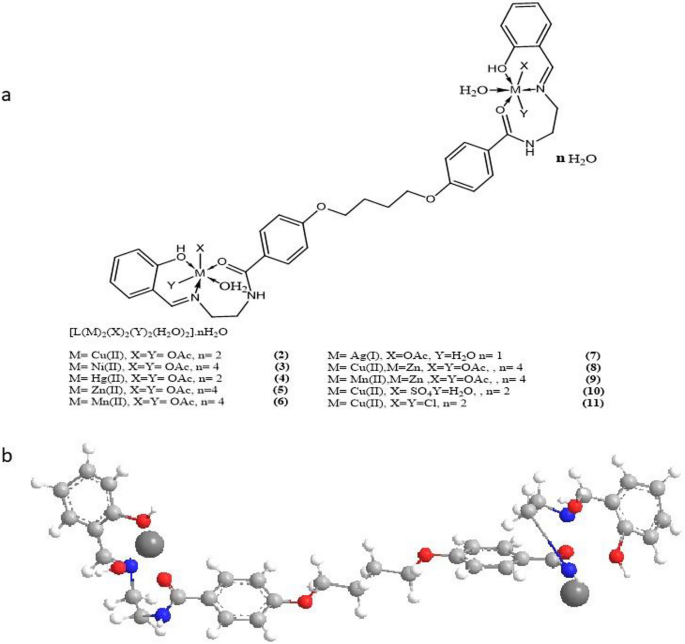
The entire complexes are nanoparticles which are steady at room temperature, non-hydroscopic, insoluble in water, and barely soluble in frequent natural solvents like CHCl3, however soluble in DMF and DMSO. The analytical and bodily knowledge of the ligand and its complexes had been supplied in Desk 1, and the spectral knowledge (Tables 2, 3, 4, 5, 6) had been appropriate with the proposed buildings, as proven in Figs. 2 and 3. Desk 1 reveals that the molar conductance in DMF resolution had been within the 6.3–16.4 Ω−1cm2mol−1 vary, indicating a non-electrolytic nature19, some complexes excessive values confirmed partial dissociation in DMF. Complexes (2)–(11) had been fashioned by reacting (1) with metallic salts in ethanol at (1L: 2M) molar ratios. The complexes had been composed of diatomic complexes.
The comparability of the infrared (IR) spectra of the ligand (1) and its metallic complexes (2)–(11) offers perception into the bonding mechanism between the ligand and the metallic ion. The presence of two sorts of hydrogen bonding, each intra- and intermolecular, is indicated by bands noticed within the 3560–3350 cm−1 and 3340–2980 cm−1 ranges of the ligand20. The upper frequency vary corresponds to a weaker hydrogen bond, whereas the decrease frequency band is related to a stronger hydrogen bond. Moreover, a medium band at 3140 cm−1 is attributed to the v(NH) group21,22. Curiously, the v(NH) group within the complexes seems on the similar location as within the free ligand, indicating that the NH group doesn’t take part in metallic ion coordination23. The v(NH) band is found within the metallic complexes inside the 3013–2546 cm−1 vary. Moreover, the v(OH) group is liable for a powerful band noticed at 3360 cm−1, which seems within the complexes inside the 3480–3420 cm−1 vary. The v(C=O) group causes a powerful band at 1690 cm−1, noticed within the complexes inside the 1685–1644 cm−1 vary. Bands noticed inside the ranges of 3560–3210 cm−1 and 3370–3820 cm−1 within the complexes are attributed to hydrated, coordinated water molecules and hydrogen bonding. One other robust band at 1625 cm−1 is attributable to the v(C=N) band, which happens all through the complicated between 1601 and 1620 cm−1. The acetate ion band v(Ac) is noticed inside the vary of 1449–1260 cm−1, whereas the sulphate ion band v(Ac) emerges at 1160, 1164, 1040, and 690 cm−1. Fragrant bands are noticed in two areas: the primary inside the vary of 1570–1460 cm−1 and the second inside the vary of 853–739 cm−1 for the ligand. Within the complexes, fragrant bands are noticed in two areas as nicely: the primary inside the vary of 1519–1402 cm−1 and the second inside the vary of 771–617 cm−1. Particularly, the band at 405 cm−1 is attributed to v(Cu–Cl) complicated (11)24,25. Complexes with bands inside the vary of 676–610 cm−1 are assigned to v(M–O), whereas bands inside the vary of 512–510 cm−1 are assigned to v(M–N)26,27. The 1H nuclear magnetic resonance (NMR) spectra of the ligand (1) and its Zn(II) complicated (5) reveal vital info. Within the ligand’s 1H-NMR spectra, a peak at 7.7 ppm is noticed as a consequence of an NH group proton, whereas peaks inside the vary of three.5–4.37 ppm are attributed to methylene group protons. Protons from the OH fragrant group are detected at 10.8 ppm. Because of the presence of robust hydrogen bonding, the protons of fragrant rings are noticed at 6.6–7.3 ppm, and the protons of the methine group seem at 8.8 ppm28,29. Within the 1H-NMR spectrum of the complexes (5), NH protons are noticed at 7.8 ppm, and methylene protons are noticed inside the vary of three.56–4.47 ppm. Peaks at 10.5 ppm are as a consequence of fragrant OH, whereas fragrant ring protons emerge inside the vary of 6.4–7.15 ppm. Lastly, peaks at 8.3 and 9.0 ppm are noticed as a consequence of methane group protons30,31. When evaluating the 1H-NMR of the ligand and the spectra of complicated (5), a major downfield shift of the proton sign in relation to the free ligand is noticed, indicating that the metallic ions are coordinated to the ligand. This shift could possibly be attributed to the formation of a coordination bond (NM)32,33,34. The proposed formulations of the ligand (1), its Ni(II) complexes (3), and Mn(II) complicated (6) are supported by the mass spectra. The spectrum of the ligand reveals molecular ion peaks (m/z) at 622 amu, which is per the molecular weight of the ligand (622). Moreover, the noticed fragments at m/z = 65, 93, 121, 152, 196, 268, 329, 415, 532, and 622 amu correspond to N2H5O2, C2H9N2O2, C3H9N2O3, C4H12N2O4, C7H20N2O4, C11H28N2O5, C15H28N3O5, C21H28N4O5, C29H33N4O6, and C36H38N4O6 moieties, respectively. Nonetheless, the Ni(II) complicated (3) displays a peak (m/z) at 1048 amu. Moreover, the noticed peaks at m/z 58, 121, 324, 500, 581, 669, 726, 802, 926, and 1084 amu are attributed to C3NO2, C4H11NO3, C15H18NO7, C24H24N2O10, C29H29N2O11, C35H29N2O12, C38H29N3O12, C40H30N3O15, C42H37N4O17Ni, and C44H58N4O18Ni2 moieties, respectively. Lastly, the Mn(II) complicated (6) displays a peak (m/z) at 960 amu. The noticed fragments (m/z) at 65, 121, 179, 330, 445, 583, 649, 795, and 960 amu are as a consequence of Assignments C3HN2, C5H3N2O2, C7HN2O4, C14H4NO8, C20H15NO10, C24H18NO12Mn, C27H20N3O14Mn, C33H30N3O18Mn, and C37H44N4O18Mn2, respectively. The information is introduced in Desk 3(i, ii, and iii).
Thermal analyses together with differential thermal evaluation (DTA) and thermogravimetric evaluation (TGA) had been carried out with a view to validate the presence of water molecules, as indicated by the IR spectra. The thermal stability of complexes (2), (6), (9), and (11) was confirmed inside the temperature vary of 27–700 °C, with no vital modifications noticed as much as 45 °C. The disruption of hydrogen bonds was noticed as an endothermic peak at temperatures starting from 45 to 55 °C, as proven in Desk 4. Subsequently, endothermic peaks had been detected inside the temperature vary of 80–120 °C, equivalent to the lack of hydrated water molecules. Moreover, the removing of coordinated water molecules occurred within the 140–260 °C vary35,36,37. The thermogram of the Cu(II) complicated (2) exhibited a disintegration course of that happened in six phases. The primary part occurred at 55 °C, leading to a minor weight reduction accompanied by an endothermic peak, presumably attributed to the breaking of hydrogen bonds. The second part occurred at 135 °C with a weight lack of 1.68% (Calculated: 1.70%) and an endothermic peak, which could possibly be related to the removing of two hydrated water molecules. The elimination of coordinated water molecules happened throughout the breakdown course of at 260 °C, inflicting a weight lack of 3.85% (Calculated: 3.46%). The removing of 4 acetate teams generated an endothermic peak at 375 °C with a weight lack of 23.42% (Calculated: 23.50%). The complicated displayed an endothermic peak at 540 °C as a consequence of its melting level. Lastly, exothermic peaks had been noticed at 415, 490, 680, and 840 °C, indicating a gradual oxidative thermal breakdown, leading to a weight lack of 20.65% (Calculated: 20.70%), forsaking 2CuO30. Equally, the thermogram of the Mn(II) complicated (6) revealed a disintegration course of occurring in six phases. The primary stage happened at 50 °C, leading to a minor weight reduction accompanied by an endothermic peak, probably related to the disruption of hydrogen bonds. The second stage occurred at 120 °C, resulting in a weight lack of 4.98% (Calculated: 5.09%) as a result of removing of two hydrated water molecules. The breakdown course of concerned the elimination of coordinated water molecules at 210 °C, inflicting a weight lack of 1.98% (Calculated: 2.05%). The removing of 4 acetate teams generated an endothermic peak at 370 °C with a weight lack of 24.29% (Calculated: 24.38%). The complicated exhibited an endothermic peak at 580 °C as a consequence of its melting level. Lastly, exothermic peaks had been noticed at 410, 465, 650, and 830 °C, indicating a sluggish oxidative thermal breakdown, leading to a weight lack of 22.88% (Calculated: 23.33%) and yielding 2MnO37. The thermogram of the Mn(II)/Zn(II) mixture (9) indicated a disintegration course of occurring in six phases. The primary part occurred at 48°C, with no weight reduction noticed, however accompanied by an endothermic peak, probably attributed to the breaking of hydrogen bonds. The second part occurred at 85 °C, leading to a weight lack of 3.36% (Calculated: 3.42%) and an endothermic peak, which could possibly be associated to the removing of two hydrated water molecules. The breakdown stage, attributable to the removing of coordinated water molecules, occurred at 140 °C, leading to a weight lack of 3.50% (Calculated: 3.55%)37. The removing of 4 acetate teams may clarify the endothermic peak reported at 280 with 24.02% weight reduction (Calc. 24.11%). Due to its melting level, the complicated displayed an endothermic peak at 380°C. Lastly, exothermic peaks come up at 415, 550, 680, and 745 °C, equivalent to oxidative thermal breakdown that progresses slowly, leaving with 20.38% weight reduction (Calc 20.47%) of MnO and ZnO38,39. In accordance with the thermogram of Cu(II) complicated (11) the complexes disintegrated in six phases. The primary occurred at 48 °C with no weight reduction as an endothermic peak, which could possibly be attributed to hydrogen bonds breaking. The second part occurred at 85 °C, with a weight lack of 3.38% (Calc. 3.74%) as an endothermic peak, which could be attributed to the removing of two hydrated water molecules. The elimination of coordinated water molecules occurred throughout the breakdown course of, which occurred at 190 °C with 3.81% weight reduction (Calc. 3.88%). The removing of 4 acetate teams resulted in an endothermic peak at 315Co with 15.85% weight reduction (Calc. 15.92%). Due to its melting level, the complicated displayed an endothermic peak at 380 °C. Lastly, exothermic peaks developed at 439, 460, and 580 °C, equivalent to gradual oxidative thermal breakdown, leaving 2CuO with a weight lack of 21.15% (Calc. 21.21%)40,41,42. The thermal knowledge are current in Desk 4.
Magnetic moments
Desk 5 shows the magnetic moments of the metallic complexes (2)–(11) at room temperature. Copper(II) complexes (2), (8), (10) and (11) have values within the 1.69–1.71 B.M. vary, indicating the existence of 1 unpaired electron in an octahedral construction42,43. The nickel(II) complicated (3) revealed values within the 3.12 B.M vary, indicating an octahedral nickel(II) complicated44. The low complicated values had been attributable to spin–spin interactions between metallic(II) ions45. The diamagnetic properties of Zn(II) complexes (5), (8), and (9), Ag(I) complicated (7), and Hg(II) complicated (4) had been noticed44 Mn(II) complexes (6) and (9) had BM values of 5.61 and 4.92, respectively, confirming octahedral geometry round Mn(II) ions46.
Digital spectra
Desk 5 summarizes the digital spectrum knowledge for the ligand (1) and its metallic complexes in DMF resolution. Ligand (1) in DMF resolution confirmed two bands at 280 nm (log€ = 3. 8 × 10–3 mol−1 cm−1) and 315 nm (log€ = 4.25 × 10–3 mol−1 cm−1)46. Copper(II) complexes (2), (8), (10) and (11) confirmed bands within the 265–267 and 280–312 nm ranges, these bands had been as a consequence of intraligand transitions, nevertheless, the bands appeared within the 462–478, 570–578 and 602–610 nm ranges, had been assigned to O → Cu, cost switch, 2B1 → 2E and 2B1 → 2B2 transitions, indicating a distorted octahedral construction47,48. Zinc(II) complexes (5), (8), and (9), Silver (I) complicated (7), and Hg(II) complicated (4) confirmed bands had been as a consequence of intraligand transitions. Nonetheless, nickel(II) complicated (3) confirmed bands at 268–285, 310–395, 475, 590,610 and 776 nm, the primary three bands had been inside the ligand and the opposite bands are attributable to O → Ni cost switch, 3A2g(F) → 3T1g(P)(ν3), 3A2g(F) → 3T1g(F)(ν2) and 3A2g(F) → 3T2g(F)(ν1) transitions respectively, indicating an octahedral Ni(II) geometry49,50. The ν2/ν1 ratio was 1.24, which was lower than the same old vary of 1.5–1.75, indicating a distorted octahedral Ni(II) complicated51,52. Mn(II) complexes (6) and (9) confirmed bands at 265, 280, 307, 462, 586, and 612 for complicated (6) which complicated (9) confirmed pands at 260, 283, 318, 420, 560, and 605 nm the primary bands had been intra-ligand transitions, nevertheless the opposite bands corresponded to A1g → Eg, A1g → T2g, and Ag → T2g transitions which had been appropriate to Mn(II) octahedral construction44,52.
Electron spin resonance (ESR)
The spectral knowledge for complexes (2), (6), (8), (9), (10), and (11) had been supplied in Desk 6. The spectra of the copper(II) complexes (2), (10), and (11) exhibited the attribute of a d9 configuration, with a floor state of d(x2 − y2), which is often noticed in copper (II) complexes.The complexes displayed g|| > g┴ > 2.0023, indicating an octahedral geometry across the copper(II) ion50,51. The g-values may be described by the equation G = (g||− 2)/ (g┴ − 2)50,52, the place G represents the change coupling interplay parameter. If G < 4.0, it signifies the presence of a major change coupling, whereas a G worth > 4.0 means that the native tetragonal axes are aligned parallel or barely misaligned. Complexes (2), (10), and (11) exhibited values of three.0 and a pair of.83, indicating spin-exchange interactions between Cu(II) ions. This phenomenon is additional confirmed by the values of the magnetic moments (see Desk 5). The g||/A|| worth can also be thought-about as a diagnostic time period for stereochemistry53. The values of g||/A|| had been 210 and 175.8, that are anticipated for distorted octahedral complexes. The g-values of the copper(II) complexes with a 2B1g floor state (g|| > g┴) may be expressed as K11254.
$$ {textual content{g}}_ = { 2}.00{2 }{-} , ({textual content{8K}}^{{2}} lambda^circ /Delta {textual content{Exy}}) $$
(1)
$$ {textual content{g}}_{ bot } = { 2}.00{2 }{-} , ({textual content{2K}}_{2 } bot lambda^circ /Delta {textual content{Exz}}) $$
(2)
the place okay|| and okay┴ are the parallel and perpendicular elements respectively of the orbital discount issue (Ok), λ° is the spin–orbit coupling fixed for the free copper, ΔExy and ΔExz are the electron transition energies of 2B1g → 2B2g and 2B1g → 2Eg. From the above relations, the orbital discount components (Ok||, Ok┴, Ok), that are measure phrases for covalency55,56, may be calculated. For an ionic atmosphere, Ok = 1; whereas for a covalent atmosphere, Ok < 1. The decrease the worth of Ok, the higher is the covalency.
$$ {textual content{Ok}}_{ bot }^{{2}} = , left( {{textual content{g}}_{ bot } – { 2}.00{2}} proper) , Delta {textual content{Exz }}/{2}lambda {textual content{o}} $$
(3)
$$ {textual content{Ok}}^{{2}} {textual content{ = (g}} , – { 2}.00{2}) , Delta {textual content{Exy }}/{8}lambda {textual content{o}} $$
(4)
$$ {textual content{Ok}}^{{2}} = , left( {{textual content{Ok}}^{{2}} , + {textual content{2K}}_{ bot }^{{2}} } proper)/{3} $$
(5)
Ok values (Desk 6), for the copper(II) complexes (2) and (10) had been indicating for a covalent bond character57,58,59,60. Kivelson and Neiman famous that, for ionic atmosphere g|| ≥ 2.3 and for a covalent atmosphere g||< 2.357. Theoretical work by Smith18 appears to verify this view. The g-values reported right here (Desk 6) confirmed appreciable covalent bond character59. Additionally, the in-plane σ-covalency parameter, α2(Cu) was calculated by
$$ alpha^{{2}} left( {{textual content{Cu}}} proper) , = , left( {{textual content{A}}||/0.0{36}} proper) , + left( {{textual content{g}}|| – {2}.00{2}} proper) , + {3}/{7}left( {{textual content{g}}_{ bot } – {2}.00{2}} proper) , + 0.0{4} $$
(6)
The calculated values (Desk 6) instructed a covalent bonding55,56,58,59,60. The in-plane and out of- aircraft π- bonding coefficients β21 and β2 respectively, are dependent upon the values of ΔExy and ΔExz within the following equations45:
$$ alpha^{{2}} beta^{{2}} = , left( {{textual content{g}}_{ bot } – { 2}.00{2}} proper) , Delta {textual content{Exy}}/{2}lambda {textual content{o}} $$
(7)
$$ alpha^{{2}} beta_{{1}}^{{2}} = , left( {{textual content{g}}|| , – { 2}.00{2}} proper) , Delta {textual content{Exz}}/{8}lambda {textual content{o}} $$
(8)
On this work, the complexes (2), and (10) confirmed β12 values 0.82, and 0.69 indicating a covalency within the in-plane π-bonding54,58,59,60. β2 worth for complexes (2), and (11) confirmed 0.83 and 0.92 indicating ionict character of the out-of-plane35. It’s doable to calculate approximate orbital populations ford orbitals40 by
$$ {textual content{A}}|| , = {textual content{ A}}_{{{textual content{iso}}}} {-}{textual content{ 2B}}left[ {{1 } pm , left( {{7}/{4}} right) , Delta {text{g}}||} right] , Delta {textual content{g}}_ = {textual content{ g}}_ – {textual content{ ge}} $$
(9)
$$ {textual content{a}}^{{2}} {textual content{d }} = {textual content{2B}}/{textual content{ 2B}}^circ $$
(10)
The calculated dipolar coupling for unit occupancy of the d orbital is denoted as A° and 2B°, respectively. In the course of the knowledge evaluation, all signal mixtures of the Cu hyperfine coupling elements had been taken under consideration. Bodily significant outcomes are solely obtained when A|| and A^ are damaging. The ensuing isotropic coupling fixed can also be damaging, in addition to the parallel element of the dipolar coupling 2B (− 175.7 and − 177.2 G). These outcomes can solely be noticed for an orbital involving the (dx2 − y2) atomic orbital on copper. The worth for 2B is inside the regular vary for copper(II) complexes. The |Aiso| worth is comparatively small. By dividing the 2B worth by 2B° (the calculated dipolar coupling for unit occupancy of d(x2 − y2) (235.11 G), utilizing Eq. (10), it’s instructed that each one orbital populations are 74.75% and 94.41% d-orbital spin density, indicating the presence of the dx2 − y2 orbital for the unpaired electron47. Then again, the Co(II) complicated (4), Mn(II) complexes (6) and (9), and [Cu(II)/Zn (II)] complicated (8), in addition to the Cu(II) complicated (10), exhibit isotropic values of two.02, 2.03, and a pair of.13, respectively, indicating an octahedral construction with covalent bond character35.
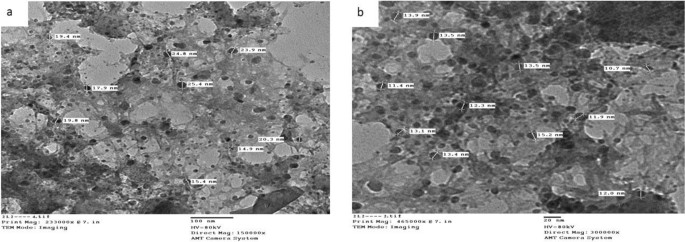
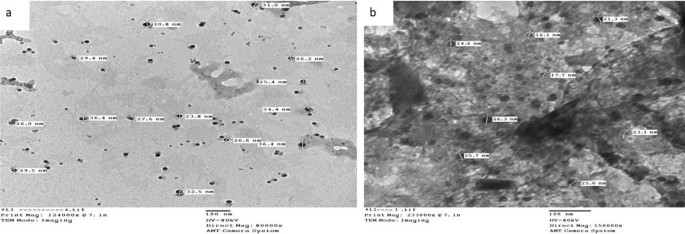
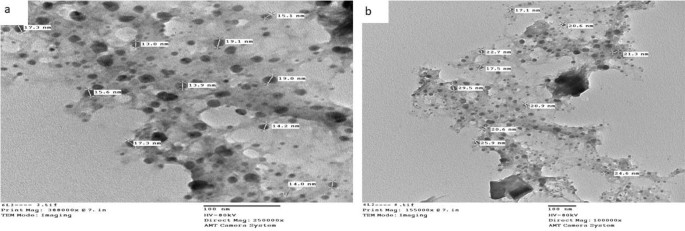
Transmission electron microscopic characterization (TEM)
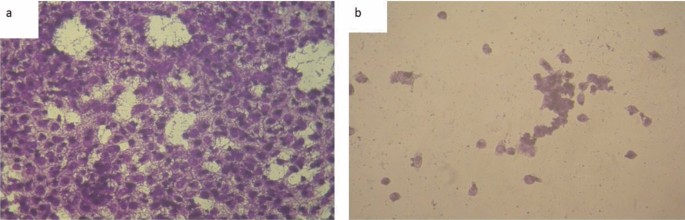
The typical diameter of the complexes particles beneath investigation was decided to be within the 13.0–36.4 nm vary, as proven in Figs. 4, 5, and 6. The particles vary in dimension from 1 to 100 nm. The complexes exhibited improved dimension dependent traits. Elevated lively agent floor space ends in sooner dissolution of the lively agent in an aqueous atmosphere such because the human physique, sooner dissolution, with gore61. The TEM photographs for Cu(II) complicated (2), Ag(I) complicated (7), and Cu(II)/Zn complicated (8) had been confirmed in Figs. 4, 5 and 6 respectively.
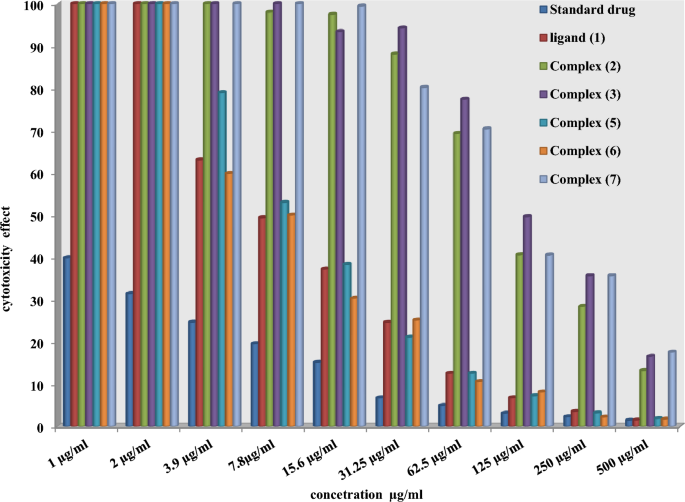
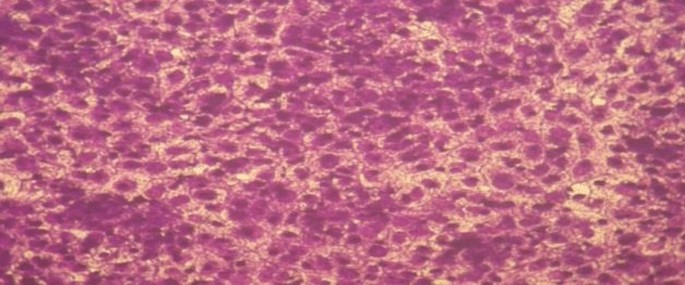
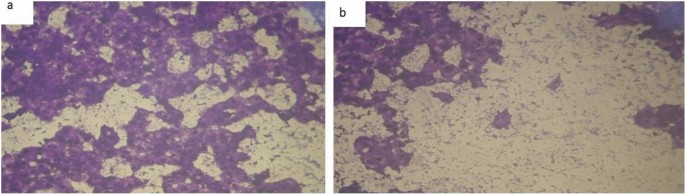
Chemotherapeutic investigations: The information introduced in Figs. 7, 8 and 13) depict the organic efficacy of ligand (1) and its metallic complexes (2), (3), (5), (6), and (7) towards HEPG-2 cell traces. On this examine, we goal to match the chemotherapeutic potential of the evaluated complexes with the reference remedy as proven in Fig. 7, Cisplatin. Altering the anion, coordination websites, and metallic ion kind appears to influence the organic habits by affecting the flexibility to bind DNA51,52,62. Gaetke and Chow beforehand proposed that metals could induce oxidative tissue harm by way of a free-radical mediated pathway much like the Fenton response. It has been demonstrated that Cu(I) complexes are able to forming GSSG (Glutathione disulfide) by way of the next reactions47,48,49:
$$ left[ {{text{Cu}}left( {text{I}} right) , left( {{text{ligand}}} right)} right] , + {textual content{ GSH }} to , left[ {{text{Cu}}left( {text{I}} right) , left( {{text{GS}}} right)} right] , + {textual content{ ligand }} + {textual content{H}}^{ + } $$
(11)
$$ left[ {{text{Cu}}left( {text{I}} right)left( {{text{GS}}} right)} right] , + {textual content{GS}} + , [{text{Cu }}left( {{text{II}}} right) + {text{GSSG]}} $$
(12)
The DNA-bound Cu(II) complicated undergoes oxidation within the presence of H2O2, probably producing Cu(II) (oxo/hydroxo) species54,55,56,57. As a consequence, the interplay between the Cu(II) complicated and nucleic acid happens by way of the Cu-oxo/hydroxy intermediate. Cu(II) complexes are well-known for his or her redox exercise, which appears to be concerned in organic processes58,59,60. The conversion of Cu(II) to Cu(I) by intracellular thiols, reminiscent of GSH (glutathione, a nonenzymatic antioxidant in environments containing oxygen), types the premise for the redox biking of Cu complexes61. Equations (13–15) illustrate the chemical pathway for [Cu(II) (ligand)] complexes. To summarize, Cu (II) complexes readily mix with GSH42, ensuing within the formation of Cu(I) complexes and GS+. This Cu(I) complicated can generate a superoxide anion within the presence of oxygen, which may induce ROS by way of a fenton-like response:
$$ left[ {{text{Cu }}left( {{text{II}}} right) , left( {{text{ligand}}} right)} right] , + {textual content{ GSH }} to , left[ {left( {{text{GS}}} right){text{ Cu}}left( {{text{II}}} right) , left( {{text{ligand}}} right)} right] , + {textual content{ H}}^{ + } $$
(13)
$$ left[ {left( {{text{GS}}} right){text{ Cu}}left( {{text{II}}} right) , left( {{text{ligand}}} right)} right] , to , left[ {{text{GS}}* , + , } right[{text{Cu}}left( {text{I}} right) , left( {{text{ligand}}} right)] $$
(14)
$$ left[ {{text{Cu}}left( {text{I}} right) , left( {{text{ligand}}} right)} right] , + {textual content{ O}}_{{2}} to {textual content{ O}}_{{2}} , + , left[ {{text{Cu}}left( {{text{II}}} right) , left( {{text{ligand}}} right)} right] $$
(15)
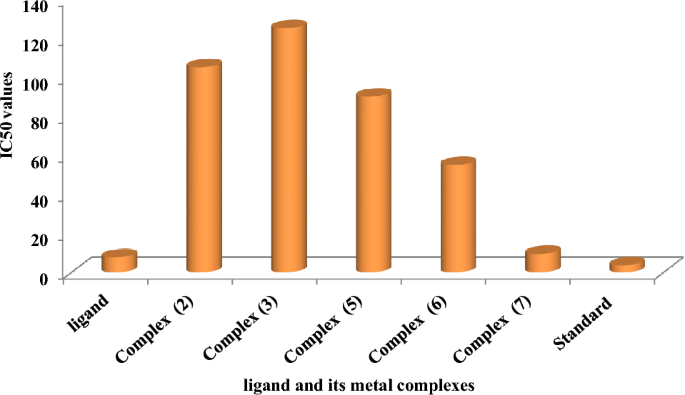
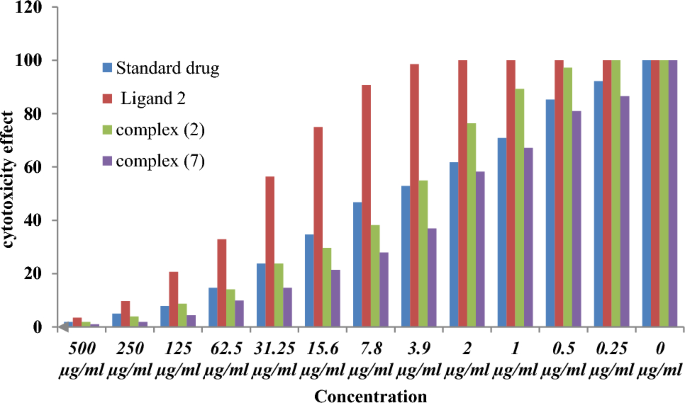
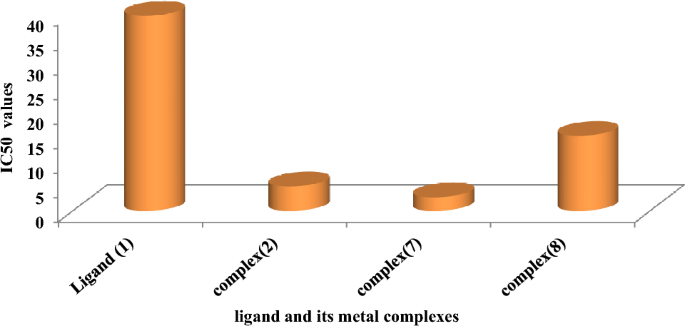
As beforehand described51, therapy with the completely different complexes in DMSO had an equal influence on the utilized tumoral cell line. The solvent dimethyl sulphoxide (DMSO) didn’t affect cell development. The ligand (1) exhibited a light inhibitory impact on the examined concentrations, whereas the complexes had a extra vital impact on HEPG-2 cell traces. The ratio of surviving fraction towards MCF-7 tumor cells elevated because the focus decreased inside the examined focus vary50. The cytotoxicity knowledge indicated that the complicated (7) beneath investigation had been efficient. complicated (7) was probably the most detrimental to cell traces, with a half-maximal inhibitory focus (IC50) worth of 9.5 M, adopted by complicated (6) with an IC50 worth of 55 Advanced (7) exhibited cytotoxicity towards cell traces in any respect doses in comparison with the usual remedy (Cisplatin), which had an IC50 worth of 15.3 M. This may be attributed to the Cu(II) ion being certain to DNA. Since altering the anion and kind of metallic ion can influence organic habits as a consequence of variations in DNA binding capacity, the analysis of various complexes is especially intriguing from this angle. Tweedy’s chelation concept means that the chemotherapeutic exercise of the complexes may be traced again to the central metallic atom48. Moreover, the constructive cost of the metallic raises the acidity of the coordinated ligand containing protons, leading to stronger hydrogen bonds that improve organic exercise39. The cytotoxic impact of the ligand and a few of its metallic complexes on HEPG-2 cell traces is introduced in Desk 7, Fig. 8.
Chemotherapeutic investigations: The information introduced in Figs. 9, 10, 11, 12 depict the organic effectivity of ligand (1) and its metallic complexes (2), (7), and (8) towards MCF-7 cell traces. On this examine, we goal to match the chemotherapeutic potential of the evaluated complexes with the reference remedy, Cisplatin. Altering the anion, coordination websites, and metallic ion kind appears to influence the organic habits by affecting the flexibility to bind DNA51,52,62. Gaetke and Chow beforehand proposed that metals could induce oxidative tissue harm by way of a free-radical mediated pathway much like the Fenton response.
The DNA-bound Cu(II) complicated undergoes oxidation within the presence of H2O2, probably producing Cu(II) (oxo/ hydroxo) species54,55,56,57. As a consequence, the interplay between the Cu(II) complicated and nucleic acid happens by way of the Cu-oxo/hydroxy intermediate. Cu(II) complexes are well-known for his or her redox exercise, which appears to be concerned in organic processes58,59,60. The conversion of Cu(II) to Cu(I) by intracellular thiols, reminiscent of GSH (glutathione, a nonenzymatic antioxidant in environments containing oxygen), types the premise for the redox biking of Cu complexes61, as proven earlier than. Therapy with the completely different complexes in DMSO had an equal influence on the utilized tumoral cell line. The solvent dimethyl sulphoxide (DMSO) didn’t affect cell development. The ligand (2) exhibited a light inhibitory impact on the examined concentrations, whereas the complexes had a extra vital impact on MCF-7 cell traces. The ratio of surviving fraction towards MCF-7 tumor cells elevated because the focus decreased inside the examined focus vary50. The cytotoxicity knowledge indicated that the complexes (2) and (8) beneath investigation had been efficient, copper(II) complicated (2) was probably the most detrimental to cell traces, with an IC50 worth of 5.0 M, adopted by complicated (7) with an IC50 worth of two.8, complicated (7) exhibited cytotoxicity towards cell traces in any respect doses in comparison with the usual remedy (Cisplatin), Fig. 13, which had an IC50 worth of 15.3 M. The IC50 worth is 15.3 M Advanced (8), Figs. 8 and 9 additionally displayed cytotoxicity towards cell traces. This may be attributed to the Cu(II) ion being certain to DNA. Since altering the anion and kind of metallic ion can influence organic habits as a consequence of variations in DNA binding capacity, the analysis of various complexes is especially intriguing from this angle. Tweedy’s chelation concept means that the chemotherapeutic exercise of the complexes may be traced again to the central metallic atom48. Moreover, the constructive cost of the metallic raises the acidity of the coordinated ligand containing protons, leading to stronger hydrogen bonds that improve organic exercise39. The cytotoxic impact of the ligand and a few of its metallic complexes on MCF-7 cell traces is introduced in Desk 8, Figs. 9 and 10.
Molecular docking of some complexes with breast most cancers protein 3S7S
Molecular modeling, docking is a technique which predicts the popular orientation of 1 molecule to a second when a ligand and a goal are certain to one another to type a steady complicated. Data of the popular orientation in flip could also be used to foretell the energy of affiliation or binding affinity between two molecules utilizing, for instance, scoring features.The protein 3S7S, which has been chosen, serves as a illustration of the crystal construction of the chemical that’s thought-about to be physiologically lively within the human placenta. By means of the utilization of this method, one is ready to confirm the location at which the ligand-receptor interplay happens, in addition to the kind of interplay that takes place. Moreover, this technique additionally permits the calculation of the space between the ligand and the receptor inside the interplay grid. The scoring vitality obtained from the docking computations corresponds to the inhibitory impact exhibited by the related ligand. Throughout the context of the current investigation, the protein 3S7S mirrors the crystal construction of the human placental aromatase enzyme, which is liable for the synthesis of the estrogen hormone and contributes to estrogen-dependent breast cancer50. Based mostly on the scoring vitality, it may be noticed that each one ligands possess a substantial variety of interactions with the receptor protein. The outcomes of the examine serve to reveal the capability of the ligand to impede the protein 3S7S. Based mostly on the scoring vitality, it may be noticed that each one ligands possess a substantial variety of interactions with the receptor protein. The outcomes of the examine serve to reveal the capability of the ligand to impede the protein 3S7S.
Advanced 2
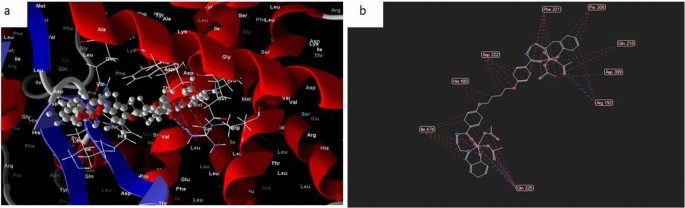
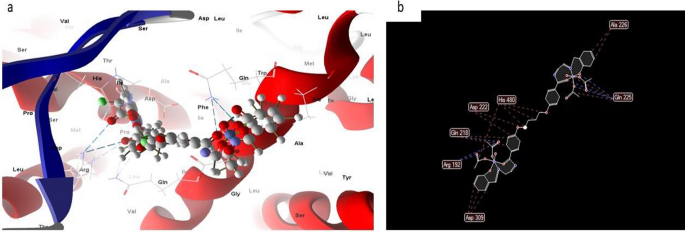
Within the case of the docked complicated 2, it could actually typically be noticed that the distances related to the efficient ligand-receptor interactions, as depicted docked and 2D construction in Fig. 14 in complicated (2) are roughly 3.5 A. This means the presence of typical precise bonds, thus signifying a major binding affinity. The closest interplay, for example, is recognized by way of H-donors with 3S7S, with a distance of two.18 A, whereas complex2 displays a scoring vitality (S) of 182,278 kcal. Moreover, it was decided that complex2 possesses 17 binding websites comprising varied amino acids (Gln 225, lIe 474, His 480, Asp 222, Phe 221, Professional 308, Gln 218, Asp 309, and Arg 192), suggesting their appreciable inhibitory impact.
Advanced (3)
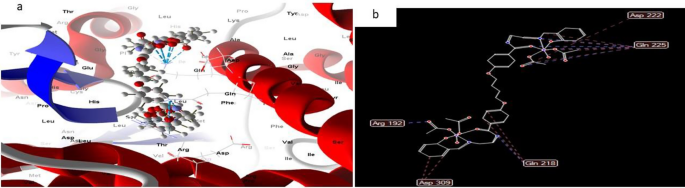
Whereas most docked and 2D construction of complicated (3), Fig. 15 had efficient ligand-receptor interplay lengths of three.5 A, indicating the presence of typical actual bonds and thus robust binding affinity. The closest interplay, for instance, is noticed through H-donors with 3S7S (2.34 A) and (complicated (3) With scoring vitality (S) 195,127 kcal. Moreover, seventeen binding websites of distinct amino acids (Ala 226,Gln 225, Asp 309, Arg 192, Gln 218, Asp 222, and His 480) with complicated(3) had been discovered, suggesting their increased inhibition than copper complicated (2).
Advanced 5
Nonetheless, most often, the docked and 2D construction of complicated (5), Fig. 16 have efficient ligand-receptor interplay distances of three.5 A, indicating the presence of typical precise bonds and thus robust binding affinity. The closest interplay, for instance, is noticed through H-donors with 3S7S (2.46 A) and complicated (5) With scoring vitality (S) 233,009 kcal. Moreover, seventeen binding websites for varied amino acids (Gln 218, Asp 309, Arg 192, Asp 222, and Gln 225) with complicated (5) had been found.
Advanced 6
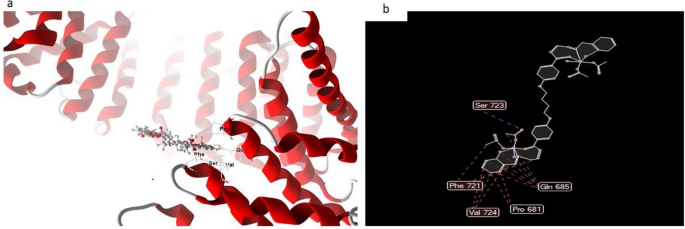
Normally, the docked and 2D construction of complicated (6) in Fig. 17, ligand-receptor interplay distances had been 3.5 A, exhibiting the presence of typical actual bonds and consequently appreciable binding affinity. For instance, the closest contact is discovered utilizing H-donors with 3S7S (3.28 A) and complicated (6) with scoring vitality (S) 230,220 kcal. Moreover, with compound (6), which is regarded to be the simplest complicated in the direction of 3S7S protein, seventeen binding websites of various amino acids (Gln 685, Professional 681, Val 724, Phe 721, and Ser 723) had been discovered.
Advanced 7
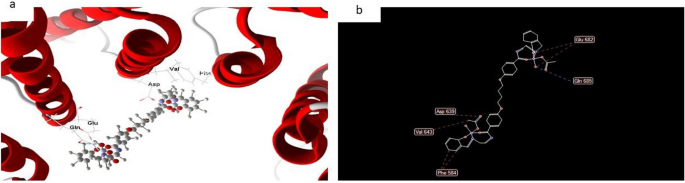
Nonetheless, the docked and 2D construction of complicated (7) in Fig. 18 have efficient ligand-receptor interplay distances had been ≤ 3.57 A most often, which signifies the presence of typical actual bonds and therefore excessive binding affinity. For instance, the closest interplay is noticed through H-donors with 3575 (3.35 A) and complicated (7) with scoring vitality (S) 173,468 kcal Moreover, seventeen binding websites had been noticed of various amino acids (Gln 685, Glu 682, Phe 584, Val 643 and Asp 639) with complicated (7).
Therefore we are able to conclude that the inhibition of complex5 > complicated 6 > complicated 3 > complicated 2 > complicated 7 to breast most cancers protein 3S7S.
Molecular docking of some complexes with liver most cancers protein 4OO6
4OO6 represents Crystal construction of human KAP-beta2 certain to the NLS of HCC1 (Hepato Mobile Carcinoma protein 1) This method elucidates the ligand-receptor website and kind of interactions. It additionally provides an estimation of the space between the ligand and the receptor contained in the interplay grid. The scoring vitality of every pose simulated by the docking calculations displays the diploma of inhibition impact of the corresponding ligand. Within the current examine.
Advanced 8
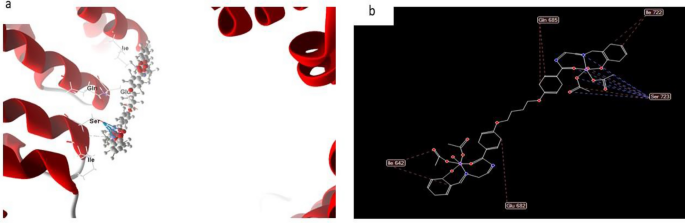
Normally, the docked and 2D construction of complicated (8), Fig. 19 had efficient ligand-receptor interplay distances of three.1 A, indicating the presence of typical precise bonds and thus robust binding affinity. The closest interplay, for instance, is noticed through H-donors with 4OO6 (2.17 A) and (complicated (8) With scoring vitality (S) 244,215 kcal. Moreover, twelve binding websites of distinct amino acids (Gln 685, lle 722, lle 642, Glu 682, and Ser 723) with complicated (8) had been discovered, suggesting their robust inhibition.
Advanced 9
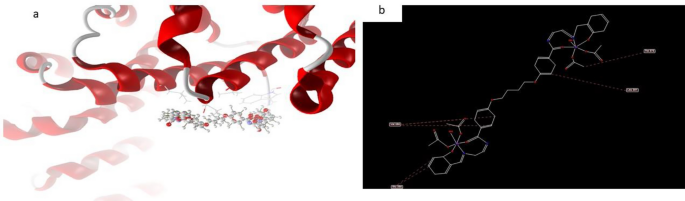
Nonetheless, the docked and 2D construction of complicated (9), Fig. 20 have efficient ligand-receptor interplay distances had been ≤ 3.1 A most often, which signifies the presence of typical actual bonds and therefore excessive binding affinity. For instance, the closest interplay is noticed through H-donors with 4OO6 (2.17 A) and complicated (9) with scoring vitality (S) 229,215 kcal Moreover, 17 binding websites had been noticed of various amino acids (Glu 286, Val 386, lle 642, Leu 281 and Trp 373) with complicated (9) demonstrating their excessive inhibition.
Advanced 10
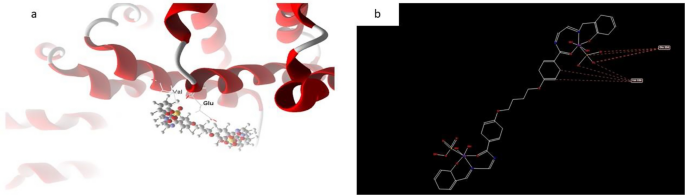
In distinction, the docked and 2D construction of complicated (10), Fig. 21 has efficient ligand-receptor interplay lengths of 4.15 A most often, indicating the presence of typical precise bonds and thus robust binding affinity. The closest interplay, for instance, is detected through H-donors with 4OO6 (3.85A) and (complex10), with scoring vitality (S) 195,131 kcal. Moreover, twelve binding websites of varied amino acids (Val 336 and Glu 286) had been found, with complex10 exhibiting their diminished inhibition52,53,54,55,57,58,61,62.
Therefore we are able to conclude that the inhibition of complex8 > complicated 9 > complicated 10 to Hepato Mobile Carcinoma protein.