Characterization and quantification of a number of most cancers phenotypes of the panel of mutant p53 protein-expressing MCF10A cell strains
We chosen to check the ten most prevalent TP53 missense mutations present in breast most cancers sufferers (Supplementary Fig. 1a), amongst which 4 mutations (R248Q, R248W, R273C, and R273H) happen at residues that contact DNA (DNA contact mutations, hereafter), and the remaining six mutations (G245S, H179R, R175H, Y163C, Y220C, and Y234C) have an effect on the general construction of the DBD (structural mutants, hereafter). Along with the variations among the many particular person mutations, it was of curiosity to see whether or not there are generalities in how these two teams of mutations have an effect on mobile phenotypes, gene expression, and DNA binding properties. For the research, MCF10A cell strains stably expressing missense mutant p53 proteins and endogenous WT TP53, have been established. Aside from p53 mutant Y234C, western blots confirmed elevated ranges of all mutant p53 proteins (Supplementary Fig. 1b), thus doubtless exerting DN results on the endogenous WT p53 proteins. The mutant protein ranges have been corresponding to these of breast most cancers cell strains harboring missense TP53 mutations akin to HCC70 (TP53R248Q), MDA-MB-468 (TP53R273H), MDA-MB-231(TP53R280K), AU565 (TP53R175H), and SK-BR-3 (TP53 R175H).
To research quantitatively how the missense mutant p53 proteins have an effect on mobile phenotypes, we carried out cell-based practical assays for measuring main most cancers hallmarks: 1) cell viability in absence of progress components; 2) resistance to apoptosis; 3) cell migration; 4) cell invasion; 5) resistance to anoikis; and 6) modifications in mammosphere morphology and polarity. Along with the parental MCF10A cells with WT TP53 (WT cells, hereafter), we included a management MCF10A cell line overexpressing the WT p53 protein (WTOE) to normalize for the protein-overexpression results, and the phenotypic values in mutant p53-expressing cells have been normalized as log2-transformed fold modifications over the WTOE values (Supplementary Desk 1). One other management cell line with knocked-down WT p53 (WTKD) utilizing shRNA constructs was additionally employed to underscore LOF vs. GOF actions of the missense mutants. To verify the knock-down, as endogenous p53 proteins have been hardly seen below regular rising situations, we induced and stabilized the p53 protein ranges by treating with doxorubicin and performing western blots. Supplementary Fig. 1c exhibits a transparent discount of p53 proteins in WTKD cells in comparison with the parental MCF10A (or WT) cells, whereas WTOE cells expressed a really excessive degree of V5-tagged p53 proteins.
Progress issue (GF)-independent survival
Most cancers cells survive and maintain excessive cell proliferation charges even within the absence of progress components. The cell line panel was incubated for 72 h in minimal medium with or with out serum (S) and/or epidermal progress issue (EGF or E hereafter) in combos (+S+E, +S−E, −S+E −S−E), and the viability was decided by the CellTiter-glo assay. Underneath regular tradition situation (+S+E), no important distinction in cell viability was noticed throughout the cell strains (Supplementary Fig. 2a). Within the absence of serum and EGF (−S−E), WTKD cells survived higher than the WT or WTOE cells (Fig. 1a). The R248Q mutant confirmed considerably larger viability than WTKD cells, indicating neomorphic GOF exercise over the LOF results, and R273C, R248W, R175H, and R273H led to elevated cell viability however to a lesser extent. The G245S cells didn’t survive with out important progress components. In settlement with proof that mammary epithelial cells require EGF for the initiation of survival signaling42, withdrawal of EGF (+S-E) decreased the viability of all different p53 mutant cell strains, besides R248Q that additionally survived higher than WTOE cells with out serum (Supplementary Fig. 2a). Underneath the GF-deprived situation (−S−E), enhanced proliferation was noticed for the mutants R175H and R248Q in comparison with WTOE cells (Supplementary Fig. 2b), demonstrating an elevated functionality of self-sustained proliferation. Likewise, mouse embryonic fibroblasts (MEFs) extracted from the p53R172H/+ (equal to human R175H) mice have been extra proliferative than p53+/− MEFs17.
Phenotypes have been measured in three unbiased experiments with replicates, and the outcomes have been normalized to the log2 fold change over the phenotype of p53 WTOE cells. The error bars characterize the usual error of imply (SEM) values, and important variations (two-sided single-sample t-test) from the WTOE imply are indicated by pink asterisks for indicated p values. In all plots, cells have been sorted by every phenotype, from much less (left) to extra (proper) aggressive. a Survivability of the cell line panel in absence of progress components (serum and EGF) have been measured by the CellTiter-glo assay. b Cell apoptosis was detected by the extent of exercise of caspase 3 and caspase 7, and the reciprocal of luminescence have been used to measure the extent of resistance to apoptosis. c Migration was assessed utilizing the transwell assay. d Cell invasion was assessed utilizing the transwell assay with Matrigel-coated chamber membranes. e To measure anoikis, the loss of life index (ratio of useless cells and stay cells) was calculated for every cell line, and the reciprocal values have been used to measure the extent of resistance to anoikis.
Resistance to apoptosis
Triggered by p53 upon DNA harm, apoptosis is a barrier to most cancers improvement, however mutations in TP53 abolish apoptosis43 inflicting resistance to medication akin to doxorubicin. To look at doxorubicin-induced apoptosis, we measured the caspases 3/7 actions after 24-hour remedy with a variety of concentrations (0.3, 0.7, 1.0, and three.0 µM) of doxorubicin across the reported IC50 for breast most cancers cells of 1.0 µM44. Practically all cells died at 3.0 µM, and the outcomes have been excluded. Apparently, when normalized to the WTOE values, the traits of apoptosis ranges throughout the cell strains have been totally different for various doses of doxorubicin. As proven in Supplementary Fig. 3a, the development at 0.3 µM was much like that of 0.7 µM (Pearson’s R = 0.54) however not 1.0 µM (R = −0.40), and the response differentials between cell strains have been a lot bigger at 0.3 µM, indicating that totally different p53 mutations have an effect on the sensitivity largely at decrease degree of DNA harm. To summarize the relative apoptotic resistance throughout a number of doses of doxorubicin as a single consultant worth for every cell line, we employed a scoring methodology of evaluating the world between the dose response curves of every cell line and the management WTOE (Supplementary Fig. 3b). Overexpression of WT p53 didn’t have an effect on considerably the sensitivity to apoptosis from the parental MCF10A cells (WT). The R175H cells have been considerably extra resistant than WTOE, whereas G245S, R273H, R273C, R248W, and R248Q have been extra delicate (Fig. 1b). Apparently, viability below GF deprivation didn’t all the time correlate with apoptosis resistance. Mutants like R248Q, R273C and R248W have been extra viable than WTOE below GF deprivation however delicate to cell apoptosis, whereas Y163C was much less viable however extra immune to apoptosis.
Cell migration and invasion
Throughout most cancers development, adherent and polarized epithelial cells bear epithelial-to-mesenchymal transition (EMT) that enables the cells to chop by way of the extracellular matrix within the basement membrane45. WT p53 suppresses EMT, whereas mutations in p53 proteins trigger destabilization of cell-cell junctions, delocalization of β-catenin into cell cytoplasm46, and cell migration. When migratory potential of the p53 mutant cells was measured utilizing the transwell plates (Fig. 1c and Supplementary Fig. 4a), Y220C, R273C, R248W, and Y163C cells have been considerably extra migratory than WTOE cells, whereas the least migratory mutants have been G245S, Y234C and R273H. Immunofluorescence staining for β-catenin and vimentin on the least (G245S), reasonably (R175H), and most (Y220C and R273C) migratory cells confirmed that Y220C and R273C cells displayed typical mesenchymal traits together with a variety “fan-like” morphology, delocalized β-catenin within the cytoplasm, and outstanding vimentin (a mesenchymal cell marker) staining (Supplementary Fig. 4b).
Transwell chamber membranes coated with Matrigel have been then used to measure cell invasion (Fig. 1d and Supplementary Fig. 5a). There was a transparent distinction between weakly invasive Y234C, H179R, and G245S cells and extra invasive R273C, Y220C, and R248W cells, which basically correlated with the degrees of migration. Apparently, the R175H cells confirmed a reasonably elevated migration however no change in invasiveness, demonstrating that selling migration just isn’t enough for making cells invasive. Additional confirming this, when two of the migratory mutants R175H (reasonably migratory however not invasive) and R248W (extremely migratory and invasive) cells have been seeded on a 3D matrix of the kind I collagen-coated wells with a Matrigel overlay, R248W cells have been detected as much as 600 µm away from the bottom of the properly, whereas only a few R175H cells might invade the Matrigel overlay (Supplementary Fig. 5b). When in comparison with the metastatic MDA-MB-231 TNBC cells, essentially the most migratory and invasive mutant p53-expressing cell strains, Y220C and R273C, have been much less migratory however equally invasive (Supplementary Fig. 6), placing these aggressive p53 mutations at an EMT induction degree much like absolutely developed metastatic most cancers cells.
Resistance to anoikis
To measure resistance to anoikis, which reinforces tumor metastasis47 and is commonly brought on by lack of practical p5348, the cell strains have been grown in low-binding plates, mimicking the detachment from the basement membrane, for 7 days to type cell aggregates. The aggregates have been stained with ethidium bromide (EtBr) and Vybrant violet to establish useless (i.e., EtBr-permeable) and stay cells, respectively, (Supplementary Fig. 7) and the loss of life index (the ratio of useless to stay cell counts) was calculated (Fig. 1e). With increased loss of life indices, Y234C and G245S cells had larger sensitivity to anoikis than WTOE, whereas the opposite mutant cells have been extra anoikis resistant, R273C and R248W being essentially the most resistant and forming spherical aggregates of stay cells (blue cells) indifferent from the 3D matrix (Supplementary Fig. 7). Notably, anoikis resistant R248W and R273C have been extra delicate to apoptosis with doxorubicin remedy (Fig. 1b), indicating opposing results of the mutations on DNA damage- vs. cell detachment-induced apoptotic applications.
The 3D mammosphere morphology
The MCF10A cells type growth-arrested, polarized hole spheres in 3D Matrigel matrix, referred to as mammospheres, which resemble the glandular epithelial structure of mammary acini41. It was reported that over-expression of WT p53 didn’t have an effect on this phenotype, however cells with out p53 (null or WTKD) and cells with just a few hotspot TP53 mutations led to depolarized, disorganized cell clusters with both stuffed or partially cleared lumen49. For top-throughput evaluation of mammosphere formation, we developed a modified ‘on-top’ methodology in a 96-well format, the place cells have been seeded on high of a 3D Matrigel layer permitting mammosphere formation inside a focal aircraft for automated confocal Z-stack imaging in a lowered time (~7–9 days).
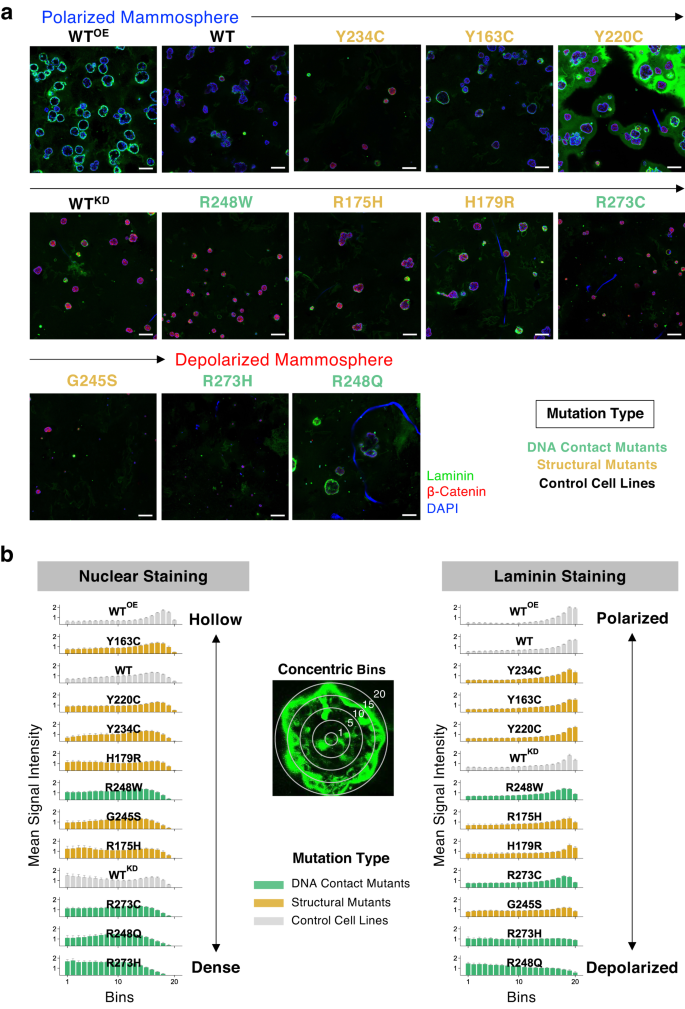
When the world of the equatorial cross-section of mammospheres derived from the cell strains (Fig. 2a) was measured, the WT and WTKD cells fashioned mammospheres with comparable imply areas of 471 and 449 µm2, respectively, whereas the WTOE cells fashioned largest hole spheres with a imply space of 785 µm2 (Supplementary Fig. 8). All mutant p53-expressing cells fashioned smaller mammospheres with various sizes, with the R273H cells forming the smallest spheroids with a imply space of simply 65 µm2, and the utmost measurement variation was noticed for R248Q, R248W, and G245S. Apparently, mammospheres fashioned by the 2 most invasive mutants R273C and Y220C confirmed a big measurement distinction with imply areas of 198 µm2 and 475 µm2, respectively.
a Low-magnification picture (scale bar: 100 μm) of the mammospheres fashioned by p53 missense mutants in Matrigel by on-top methodology stained for nucleus (DAPI, blue), laminin (inexperienced), and β-catenin (pink). The cell strains have been ordered by the diploma of polarization proven beneath in (b) which exhibits the hollowness and laminin staining of the mammospheres. The nuclear DAPI depth was measured in 20 concentric bins from the middle of the mammospheres (left panel). The suitable panel exhibits the laminin staining depth profiles throughout the bins, the place laminin sign depth within the inside bins point out disruption of cell polarity. The depth worth information was normalized to the overall variety of pixels for every spheroid.
The hollowness of mammospheres was quantified by the area-normalized DAPI sign intensities in 20 consecutive concentric bins (or rings) from the middle (Fig. 2b). The WTOE cells fashioned mammospheres with cleared lumen just like the WT cells, whereas WTKD and all of the p53 mutant cells besides Y163C didn’t clear cells on the middle. The mutants R273H, R248Q, and R273C, which fashioned smaller spheroids, had outstanding nuclear staining on the inside bins indicating stuffed lumens, whereas the remaining mutants had partially cleared lumens just like the WTKD cells.
An essential morphological attribute of regular mammospheres is the apical and basal polarization of the cells throughout the peripheral layer. When the area-normalized staining depth of laminin, a protein that usually strains the outer basal aspect of the spheroids, was measured throughout the 20 concentric bins (Fig. 2b), WTOE and WT cells had sturdy peripheral laminin staining, indicating regular epithelial cell polarity, whereas the laminin staining was detected contained in the mammospheres for WTKD cells, reflecting disrupted cell polarity (Supplementary Fig. 9). The R248Q cells confirmed the strongest laminin staining on the middle, whereas the laminin depth remained fixed throughout the bins for the mutants R273H, G245S, and R273C that fashioned the dense mammospheres. Notably, G245S-derived mammospheres confirmed very low laminin ranges general, and the mammospheres fashioned by the least invasive and essentially the most anoikis-sensitive Y234C cells had predominantly peripheral laminin staining.
Totally different missense mutant p53 proteins induce heterogeneous neomorphic phenotypes
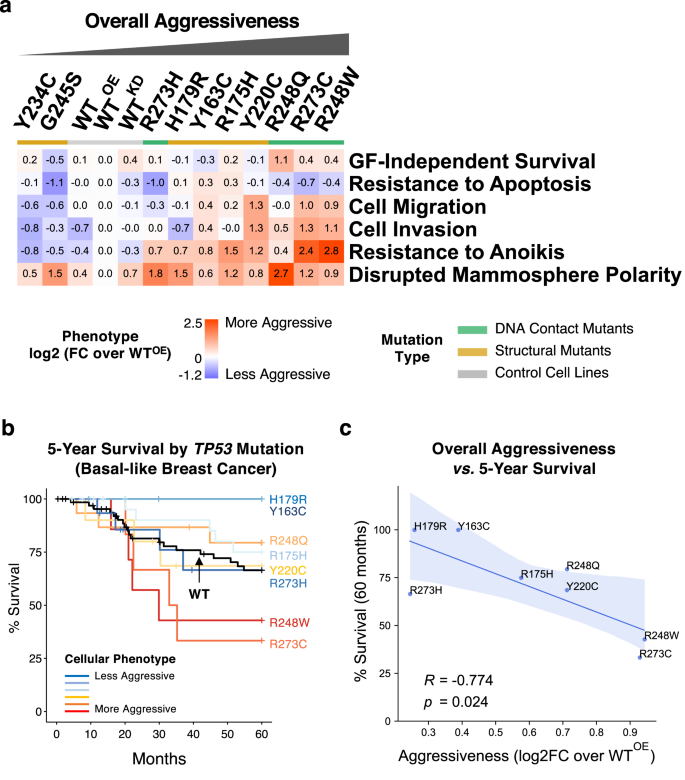
As summarized in Fig. 3a and Supplementary Desk 1, totally different missense p53 mutant proteins are functionally unequal, with distinct “neomorphic” phenotypes in contrast to these of the WTKD cells, and paying homage to the phenotypic heterogeneity noticed in TNBC tumors31,50. In comparison with the reference cell line WTOE, the ten p53 missense mutants organized into teams with much less or extra aggressive oncogenic behaviors, besides mammosphere polarity, the place all missense mutants led to partially or absolutely disrupted polarity. Throughout all phenotypes, p53 mutants R248W, R273C, and R248Q have been essentially the most aggressive, which confirmed elevated migration/invasion, anoikis resistance and lack of cell polarity in mammospheres, whereas the Y234C and G245S mutants have been the least aggressive. Because the expression ranges of mutant p53 proteins diverse (Supplementary Fig. 1b), we additionally examined whether or not the phenotypic variations have been affected by the relative protein ranges between the cells. Nevertheless, we noticed no important correlation between protein ranges and the phenotypic traits apart from a average, however not important, correlation to the Resistance to Apoptosis (Pearson’s R = 0.49, p = 0.224).
a The phenotype measurements for every of the p53 mutant-expressing cells have been normalized (as log2-transformed imply fold modifications) over these of the management p53 WTOE cells and displayed as a warmth map. The cell strains have been sorted by the imply of the normalized values for all 6 phenotypes. The p53 mutants are color-coded by mutant sort as indicated. The detailed outcomes are in Supplementary Desk 1. b 5-year survival charges of basal-like breast most cancers sufferers with totally different missense TP53 mutations have been obtained from TCGA and METABRIC datasets and visualized by the Kaplan-Meier plot. Solely the mutations with 3 or extra corresponding samples have been used. The mutations have been color-coded by the phenotypic aggressiveness (pink: extra aggressive, blue: much less aggressive), and the WT TP53 curve is proven in black. c Correlation between the general aggressiveness and the 5-year survival charges for various p53 mutations is proven with the Pearson’s correlation coefficient (R) and the p worth. The blue bands characterize the 95% confidence intervals.
Affiliation of mobile phenotypes with general survival of breast most cancers sufferers with missense p53 mutations
To look at whether or not the general and particular person phenotypic traits have been related to medical phenotypes, we’ve got carried out bioinformatics analyses on publicly obtainable patient-based datasets of TCGA51 and METABRIC52. When the basal-like subtype of sufferers with both wild-type TP53 (n = 67) or the ten missense TP53 mutations (n = 90) utilized in our research have been chosen and grouped by the TP53 mutation sort, it was instantly noticeable that there have been solely two or fewer sufferers with both Y234C or G245S mutations, in settlement with their least aggressive behaviors in our phenotypic assays (Fig. 3a). When the 5-year general survival charges of sufferers with the remaining 8 TP53 mutations with 3 or extra obtainable samples have been then in contrast together with the WT p53 group, there was a transparent correlative development between the general phenotypic aggressiveness and the survival price (Fig. 3b and Supplementary Fig. 10a). Sufferers with essentially the most aggressive R248W and R273C mutations primarily based on our mobile assay had the bottom survival charges, whereas these with much less aggressive Y163C and R179R mutants confirmed increased survival charges with a marginal statistical significance (log-rank check p = 0.072) (Supplementary Fig. 10b). As well as, quantitative evaluation confirmed a powerful and important inverse correlation (Pearson’s R = −0.774, p = 0.024) between the general phenotypic aggressiveness and the survival charges (Fig. 3c). When the affiliation of particular person mobile phenotypes have been examined, Cell Invasion and Resistance to Anoikis each confirmed important inverse correlations with the survival charges (Supplementary Fig. 10c), indicating sturdy relevance of those cell-level phenotypes to the medical illustration of absolutely developed most cancers. Regardless of the restricted availability in medical pattern measurement, these counsel that the outcomes obtained through the use of our cell-based mannequin replicate reliably the medical phenotypic aggressiveness and that the medical phenotypes could also be predisposed by the kind of TP53 missense mutations and the accompanying mobile modifications acquired on the very early stage of most cancers improvement.
Transcriptomic profiling and integrative quantitative pathway evaluation recognized practical traits related to phenotypes of mutant p53-expressing cells
On condition that p53 is a transcription issue and that these missense mutations cluster within the DBD, we hypothesized that these various phenotypes may come up largely from disparate transcriptional regulatory features of every mutant p53 protein with downstream modifications in gene expression profiles and mobile pathways. Though non-transcriptional mechanisms, akin to altered protein–protein interactions, may also contribute to the neomorphic phenotypic modifications, on this research, we centered on the gene regulatory results (RNA-Seq) and the DNA binding properties (ChIP-Seq) of various p53 mutant proteins, adopted by complete bioinformatics analyses to deduce molecular mechanism underlying the phenotypic heterogeneity.
As we used unprovoked cells for phenotype measurements, we profiled the baseline transcriptome (i.e., with out p53 inducers akin to doxorubicin or irradiation) utilizing RNA-Seq on the complete cell panel (Supplementary Desk 2a). Principal Part Evaluation (PCA) utilizing essentially the most variably expressed genes (n = 2383) throughout the cell strains positioned all p53 mutant-expressing cell strains individually from the management cell strains (Supplementary Fig. 11a), amongst which the R273H cell line was distantly positioned, indicating essentially the most distinct gene expression profile. When general aggressiveness was overlayed onto the PCA plot, essentially the most aggressive R248W, R273C, R248Q, and Y220C mutants clustered in proximity (i.e., sharing related gene expression profiles), whereas the much less aggressive Y234C and G245S mutants unfold out, indicating distinct gene expression profiles between them. No clear separation between totally different mutation varieties was noticed (Supplementary Fig. 11b). When particular person phenotypes have been examined, extra invasive, migratory, or anoikis-resistant p53 mutants shared comparatively related world gene expression profiles (Supplementary Fig. 11c).
To search out organic pathways that have been correlatively dysregulated with phenotypic traits among the many 13 cell strains, we developed a set of three pathway evaluation strategies (Supplementary Fig. 12) tailor-made particularly for quantifying the associations between two units of steady variables (i.e., phenotypic scores vs. gene expression profiles throughout the cell strains), in distinction to traditional strategies akin to Fisher’s precise check for analyzing discretized information (e.g., non-invasive vs. invasive cells). We looked for pathways that have been enriched among the many genes with a powerful expression-to-phenotype correlation throughout the 13 cell strains through the use of the Gene Set Enrichment Evaluation (GSEA)53 (Supplementary Desk 2a). We additionally computed the enrichment ranges of pathways in every cell line by the single-sample GSEA (ssGSEA)54 after which measured correlation of the enrichment ranges with the phenotypes. Lastly, we adopted a machine-learning strategy of Partial Least Squares Regression (PLSR), a strong regression modeling methodology appropriate for analyzing excessive dimensional information with small pattern sizes55,56. Multivariate PLSR fashions have been educated on expression values of a given pathway gene set to suit finest to the phenotypic traits, and the pathway-to-phenotype affiliation was assessed by measuring correlation between the noticed and the model-predicted phenotypic values. For sturdy measurement of mannequin efficiency, fivefold cross-validation was employed.
A complete of 2093 pathway phrases have been ranked for phenotype affiliation by the enrichment p values (GSEA, Supplementary Desk 2b), absolutely the phenotype correlation (ssGSEA, Supplementary Desk 2c), and the correlation between noticed and model-predicted phenotypes (PLSR, Supplementary Desk 2nd). When the highest phrases for every phenotype have been chosen through the use of the minimal percentile ranks of the three strategies as the general abstract metric, a number of phrases have been represented broadly throughout totally different phenotypes, together with a number of signaling and metabolic pathways (Fig. 4). Most notably, a number of Hippo/YAP/TAZ pathway-related phrases have been noticed for cell invasion, GF-independent survival, resistance to apoptosis, and disrupted mammosphere polarity, akin to “Hippo-YAP Signaling Pathway”, “YAP1_DN” (down-regulated genes upon overexpression of YAP1), and “Cordenonsi YAP Conserved Signature” (evolutionary conserved downstream goal gene signature of YAP). Particularly for invasion, the Hippo pathway was considerably enriched among the many extremely ranked gene units (GSEA p = 0.0004). As compared, a number of KRAS-related phrases have been additionally noticed, however the enrichment was not important (GSEA p = 0.64). We then quantified relative affiliation energy of every pathway to phenotypes by performing PLSR on ssGSEA values of total gene units (e.g., WikiPathways) with cross-validated function choice.
For the highest 15 phenotype-associated pathways for every phenotype chosen primarily based on the minimal rank in GSEA, ssGSEA, and PLSR outcomes, the percentile ranks are displayed as warmth maps. The pathways mentioned in texts are color-coded, as indicated. KP KEGG pathways, RP Reactome pathways, WP WikiPathways, and OS MSigDB Oncogenic Signatures. RNA-Seq and pathway evaluation outcomes are in Supplementary Desk 2.
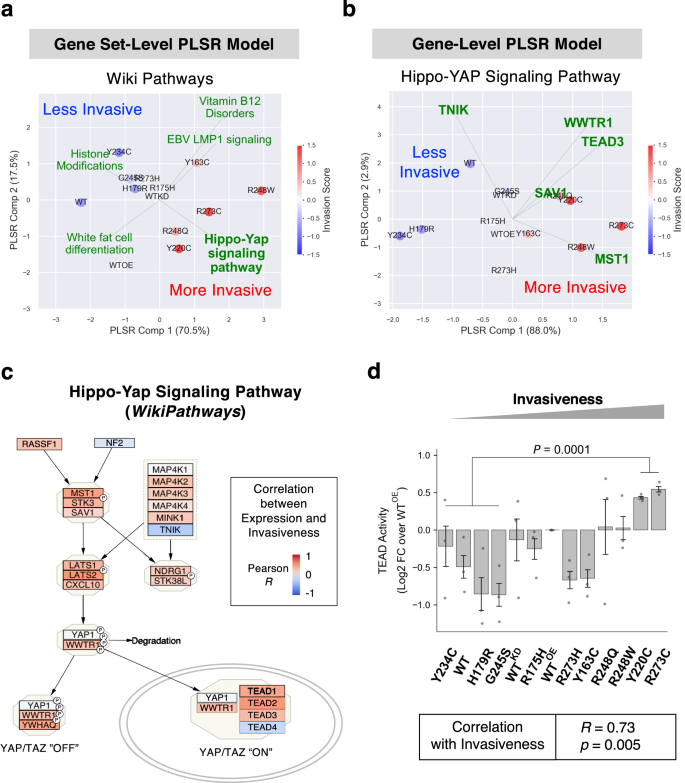
Determine 5a exhibits that the fitted PLSR mannequin with high 5 invasion-associated pathways have been in a position to clarify 88% of the phenotypic variance and challenge them in 2D house by their invasiveness from the highest left (much less invasive) to the underside proper (extra invasive). Additional, the “Hippo-YAP Signaling Pathway” was recognized as the highest contributing pathway to elevated invasiveness through the function choice course of. The Hippo/YAP/TAZ pathway regulates a variety of most cancers phenotypes together with cell proliferation, survival, EMT, anchorage unbiased progress, cell migration, invasion, and stemness57,58,59. As well as, top-ranked included pathways associated to SREBP (or SREBF) (Fig. 4), a grasp transcriptional regulator of mevalonate/ldl cholesterol metabolism and a identified upstream regulator of the Hippo pathway performing through Rho GTPases60. Determine 5b exhibits a 2D part projection (explaining >90% of phenotypic variance) of cell strains by a PLSR mannequin educated with the highest 5 feature-selected genes within the Hippo pathway, the place expression ranges of MST1 and TNIK contributed positively or negatively, respectively, to invasiveness within the totally different mutant p53 background. When expression of the Hippo pathway genes was examined, a lot of the genes have been extremely expressed in invasive cells, with MST1 (or STK4), LATS2, and TEAD2 strongly correlating with invasiveness (Fig. 5c and Supplementary Fig. 13a). As well as, expression of a big fraction of the downstream goal genes of YAP/TAZ activation within the Cordenonsi YAP Conserved Signature additionally positively correlated with invasiveness (Supplementary Fig. 13b). These counsel that coordinated transcriptional regulation of the Hippo pathway genes downstream of various mutant p53 proteins might govern invasiveness.
a Biplots for a PLSR mannequin educated on ssGSEA scores of the complete pathway phrases in WikiPathways database and the vector of invasiveness is proven. The mannequin was primarily based on high 5 pathway phrases that have been recognized by cross-validated ahead function choice. Cell strains (coloured by invasiveness) have been reworked and projected on a 2-component house, and the defined variance of invasiveness by every part is proven within the axis labels. The loadings of pathway phrases have been scaled to suit the info vary and displayed as inexperienced strains. b Biplot for a PLSR mannequin educated on expression values of genes within the Hippo-YAP Signaling Pathway (WikiPathways) and the vector of invasiveness is proven. Evaluation carried out as described for Fig. 5a. c Within the WikiPathways diagrams for the Hippo/YAP/TAZ pathways, particular person genes have been color-coded by the Pearson correlations between gene expression ranges and invasiveness throughout the 13 cell strains. d Transcriptional exercise of TEAD proteins was measured within the 13-cell line panel in triplicates by a cell-based luciferase reporter assay. The luminescence values have been normalized to the log2 fold modifications over the WTOE values. The distinction between a bunch of two extra invasive cells (R273C and Y220C) and 4 much less invasive cells (Y234C, WT, H179R, and G245S) was examined by the two-sided Pupil’s t-test. Pearson’s correlation between the normalized TEAD actions and the invasiveness throughout the cell strains was additionally calculated (backside field).
Dysregulation of hippo pathway correlated with invasiveness of cells with totally different p53 mutants
We then sought practical affirmation if the Hippo pathway exercise was concordantly regulated with invasiveness of the p53 mutant cell strains, which was considerably related to survival charges of basal-like breast most cancers sufferers (Supplementary Fig. 10c). As depicted in Fig. 5c, the tumor suppressor exercise of the Hippo pathway results in phosphorylation of the YAP/TAZ proteins by LATS1/2 kinases, inflicting their cytoplasmic retention and degradation. In distinction, the unphosphorylated types of YAP/TAZ drive oncogenesis by partnering with TEAD transcription components to extend transcription of the related targets61. To check whether or not exercise of the Hippo pathway correlated with cell invasion, we measured the transcriptional exercise of TEAD protein (i.e., the practical output of YAP/TAZ activation) in cells by transfecting with the luciferase reporter plasmid with TEAD-binding sequence motifs. As proven in Fig. 5d, the TEAD-mediated transcriptional activation was considerably correlated with invasiveness (Pearson’s R = 0.73, p = 0.005) throughout the 13 cell strains. Additional, the 2 most invasive cell strains, R273C and Y220C confirmed considerably increased (two-sided t-test p = 0.0001) TEAD actions than these of the 4 least invasive cells. These demonstrated that the Hippo/YAP/TAZ/TEAD axis could also be a key contributor for phenotypic heterogeneity figuring out the invasiveness of the cells expressing totally different missense p53 mutations, in settlement with the reported function of TAZ in breast most cancers improvement and aggressive phenotypes in cell/animal fashions62 and breast carcinomas63. In our RNA-Seq information, expression of WWTR1 (the gene image of TAZ) however not YAP1 was additionally elevated in invasive cell strains (Fig. 5c and Supplementary Fig. 13a), implying the selective regulation of TAZ transcription by totally different mutant p53 proteins in selling cell invasion.
Affiliation of the Hippo pathway dysregulation with basal-like molecular subtype of breast tumors and survival charges
To research whether or not the dysregulated Hippo/YAP/TAZ pathway signature recognized from the MCF10A mannequin is related for different most cancers cell strains and tumors, we carried out bioinformatics evaluation on the Hippo pathway actions in a broad spectrum of breast most cancers cell strains in CCLE64 in addition to affected person tumor pattern information in TCGA and METABRIC databases. ssGSEA evaluation on RNA-Seq information confirmed that elevation of genes in a number of Hippo-related pathway was a definite signature of the basal-like breast most cancers subtype amongst cell strains (Supplementary Fig. 14a, left panel) in addition to tumors (Supplementary Fig. 14b, c, left panels). Apparently, when analyzed by PCA primarily based on the pathway enrichment scores, the Hippo pathway signature was extra strongly related to the basal-like subtypes of cell strains and tumors than different main cancer-associated pathways together with p53, PI-3K, or Ras pathways (Supplementary Fig. 14, proper panels), as indicated by the upper Calinski-Harabasz (CH) Indices of the Hippo pathway, which measured the energy of clustering of the basal-like pattern group relative to different subtypes. Subsequently, extra aggressive p53 mutants might push the cell phenotypes in the direction of extra basal-like by inducing selectively ectopic activation of the Hippo pathway.
We then examined whether or not the elevated Hippo/YAP/TAZ pathway signature was related to medical phenotypic aggressiveness. When the basal-like samples in TCGA and METABRIC have been chosen and clustered by ssGSEA enrichment scores for the Hippo-related pathways, three clusters with excessive, mid, and low ranges of enrichments have been noticed (Supplementary Fig. 15a). After we then in contrast the 5-year survival charges, the Hippo-high/mid teams had poor survival with a marginal statistical significance (log-rank check p = 0.08, Supplementary Fig. 15b, high panel). Importantly, when the identical methodology was utilized to different cancer-related pathways, we didn’t observe any distinction between the pattern teams with increased and decrease pathway enrichments (Supplementary Fig. 15b, backside panels). These counsel that dysregulation of the Hippo/YAP/TAZ pathway is a district signature of clinically aggressive tumors throughout the basal-like subtype.
ChIP-Seq uncovered heterogeneity in DNA binding capability and choice
The p53 protein is a transcription issue with roughly 300 identified goal genes, which binds to a well-conserved motif65. To check how totally different missense mutations have an effect on the DNA binding properties, notably relating to invasiveness, we carried out ChIP-Seq on two invasive R273C and Y220C and two much less invasive R273H and Y234C cell strains through the use of an antibody towards the V5 tag fused on the C-term of p53 mutant proteins. We additionally included the WTOE cells as a constructive management and the WT cells expressing solely endogenous WT p53 as a adverse management. To allow an built-in evaluation, we matched the RNA-Seq experimental situations utilizing cells below unstimulated tradition situations, which contrasts with different p53 ChIP-Seq experiments carried out after treating cells with doxorubicin or 5-FU to raise the p53 protein ranges and induce DNA binding66,67. Thus, p53 DNA occupancy ranges in our setting have been anticipated to replicate the decrease tonic situations of the early-stage TP53-mutated tumor cells within the absence of extreme DNA harm or mobile stresses.
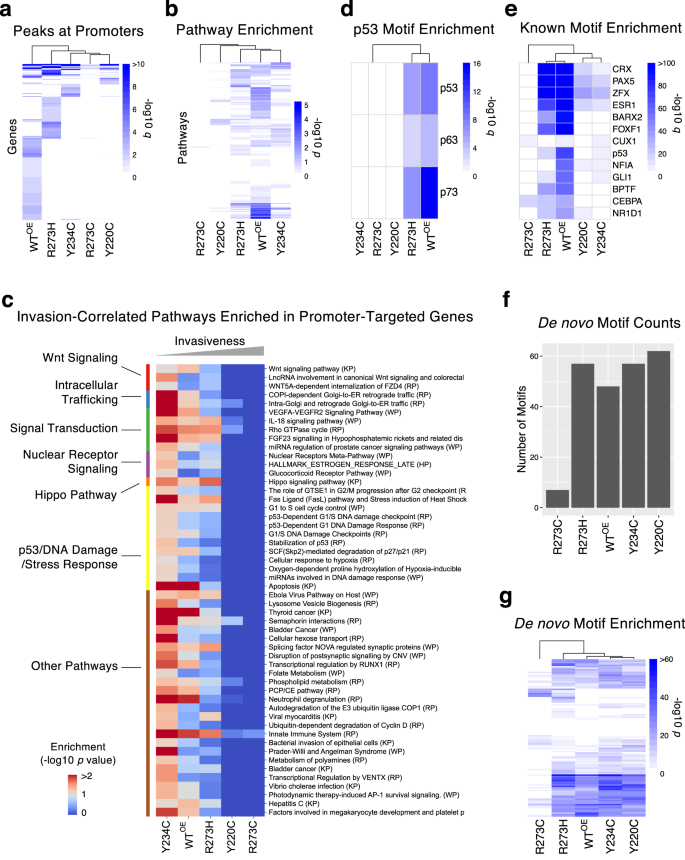
When in comparison with the WTOE cells with 11,277 peaks, fewer peaks have been recognized for R273C, Y220C, and Y234C cells (733, 1517, and 2145 peaks, respectively), suggesting considerably lowered DNA binding, whereas R273H confirmed elevated DNA binding capability (25,309 peaks) (Supplementary Fig. 16a). It was notably attention-grabbing that two mutations on the identical DNA-contacting residue, R273C and R273H, resulted in contrasting modifications in DNA binding capability, which can underlie their reverse phenotypic traits. Solely 6.7% and 4.7% of peaks for R273C and R273H, respectively, have been recognized throughout the promoter areas (outlined as −2500 to +100 bps of the transcription begin websites) (Supplementary Fig. 16b), markedly lower than the fractions noticed in Y220C (17.3%), Y234C (21.9%), WTOE (18.3%) in addition to the complete human genome (8.7%). These mutations at R273 apparently distinguish the DNA binding capability from the sequence-specific recognition inside promoters, particularly R273H, which binds to extra websites however not particularly. All 4 mutant p53 proteins, notably R273C and Y220C, misplaced promoter binding to native transcriptional goal genes of WT p53 and certain to various genes (Fig. 6a and Supplementary Desk 3a) with a restricted overlap between them (Supplementary Fig. 16c). Pathway enrichment evaluation (hypergeometric check) on the promoter-bound genes demonstrated that the goal genes of p53 mutants shared solely a small fraction of enriched pathways for the WT p53 (Fig. 6b), implying a serious alteration in downstream mobile occasions. General, the weakly aggressive mutants, Y234C and R273H, shared considerably extra frequent pathways (Chi-squared check p < 1.0e−10) with WT than the aggressive Y220C and R273C mutants. After we examined correlation between the pathway enrichments and invasiveness, a complete of 52 pathways confirmed important inverse correlation with invasiveness (Pearson’s correlation p worth < 0.05, Fig. 6c and Supplementary Desk 3b). Not surprisingly, given the lack of DNA binding within the extra invasive cells, no pathways have been extra enriched in invasive cells. The recognized pathways included the Hippo Pathway, implying transcriptional dysregulation by extra invasive p53 mutants. Demonstrating lack of binding to identified p53-targeted genes, p53/DNA harm/stress response-related pathways have been depleted in additional invasive Y22C and R273C cells. Different invasion-associated pathways have been associated to Wnt signaling, nuclear receptor signaling, and intracellular trafficking. Collectively, these outcomes point out that lack of binding to canonical p53 targets by mutant p53 proteins and the concomitant neomorphic acquire of DNA binding choice could also be contributing mechanisms to heterogeneity in invasiveness induced by totally different missense mutant p53 proteins.
a Genes that have been focused (peak detection q < 0.05) by WT and mutant p53 proteins on the promoter area are proven as a warmth map. The colours characterize the −log10 q values. b Enriched pathways (hypergeometric check p < 0.05) within the recognized targets of WT and mutant p53 proteins are proven as a warmth map of −log10 p values. Consultant outcomes primarily based on the WikiPathways pathway set are proven. c Enrichment of the canonical DNA binding motif of the p53 protein household in promoter areas for WT and mutant p53 proteins. The colour represents −log10 (hypergeometric check q). d Invasion-correlated (Pearson’s correlation p worth < 0.05) pathways that have been enriched in goal genes (hypergeometric check p worth < 0.1) of WT or mutant p53 proteins. The phrases have been sorted by the pathway teams (proven in left). e Enrichment of high identified transcription issue binding motifs inside recognized peaks for WT and mutant p53 proteins. The colour represents −log10 (hypergeometric check q). f The bar plot exhibits the variety of de novo motifs (p < 1.0e-10) discovered for WT and mutant p53 proteins. g Enrichment profile of de novo binding motifs for WT and mutant p53 proteins are proven. The colour represents −log10 (hypergeometric check p). The detailed ChIP-Seq evaluation outcomes will be present in Supplementary Desk 3.
In step with the absence of p53-binding inducers, we noticed the canonical p53 binding motif of RRRCWWGYYY68 in solely 15.5% of all recognized peaks even for the WTOE cells. This agrees with a earlier research of WT p53 within the absence of any stress stimuli69 however was lower than 30% for different research that used p53-inducing situations67. Nonetheless, the p53-binding motif sequence was considerably enriched for the WT p53 (hypergeometric check p = 1.0e-9) adopted by R273H (hypergeometric check p = 1.0e-7), however not for different mutants (Fig. 6d). Equally, well-known targets of WT p53 akin to CDKN1A have been certain by solely p53 WTOE and R273H however not the opposite p53 mutants (Supplementary Desk 3a). When enrichment of all identified transcription issue binding motifs have been in contrast, R273H and WTOE cells confirmed the same profile, whereas different mutants had lowered ranges of motif enrichment (Fig. 6e, Supplementary Fig. 16e, and Supplementary Desk 3c). Notably, the R273C mutant virtually misplaced general DNA binding capability (Fig. 6a) however appeared to realize binding to a brand new motif, such because the CUX1 motif (Fig. 6e). Additional, a de novo motif discovery evaluation confirmed {that a} whole of 101 motifs (FDR < 0.05) have been recognized throughout all cell strains, and R273C and R273H certain to fewer motifs than WT, Y220C, and Y234C (Fig. 6f and Supplementary Desk 3d). When the cell strains have been clustered primarily based on the enrichment ranges of de novo motifs, R273C confirmed essentially the most distinct profile from that of WT p53 (Fig. 6f, g). General, our ChIP-Seq evaluation demonstrated that R273H retained related DNA binding functionality and choice because the WT p53, whereas the R273C was dissimilar.
Transcriptional regulatory operate of various p53 missense mutants inferred by built-in evaluation of RNA-Seq and ChIP-Seq information
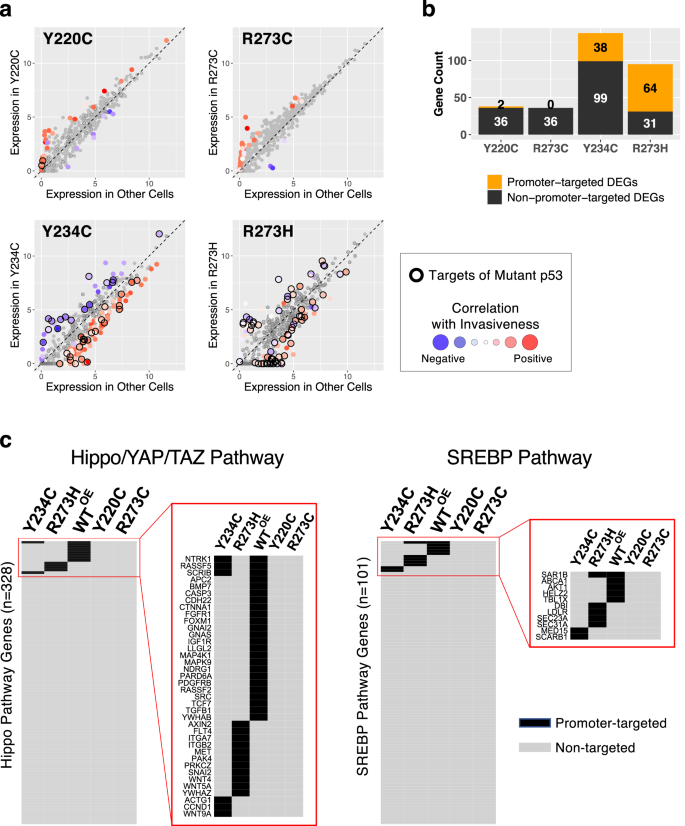
To estimate the contribution of p53 mutant protein to altered gene expression, we built-in the RNA-Seq and ChIP-Seq information and quantified the fraction of the differentially expressed genes (DEGs) that have been promoter-bound by the p53 mutant protein (Fig. 7a). As p53 features as each a direct transcriptional activator and an oblique transcriptional repressor70, the path of differential expression was additionally examined. The mutants R273H and Y234C induced each constructive and adverse gene expression modifications, of which a big fraction (64/95 and 38/137 genes, respectively) corresponded to direct promoter-binding targets of the mutant p53 types (indicated by a black ring). In distinction, the invasive p53 mutants, R273C and Y220C, have been biased in the direction of upregulated genes, and only a few of the mutant DEGs have been promoter binding targets, indicating an oblique mode of transcriptional regulation (Fig. 7b).
a Gene expression ranges, proven as log2(TPM), in every p53 mutant cell line (y axis) have been in comparison with the imply expression ranges within the different 9 mutant p53 expressing cell strains (x axis), and the differentially expressed genes (DEG, Z score-converted q < 0.05) have been color-coded in keeping with the correlation between expression ranges and invasiveness throughout all cell strains. To look at the impression of p53 binding on the path and the diploma of expressional modifications, the promoter-targeted genes recognized by ChIP-Seq for every p53 mutant are highlighted with black border. b The numbers of promoter-targeted (in orange) and non-targeted (black) DEGs by every p53 mutant are proven. c Promoter-targeted genes (peak detection q < 0.05, in black) by WT and mutant p53 proteins within the Hippo/YAP/TAZ and the SREBP pathways are displayed as warmth maps.
Because the Hippo/YAP/TAZ and the SREBP pathways have been recognized as the highest invasion-associated pathways, we examined whether or not the genes in these pathways have been instantly focused by mutant p53 proteins. Apparently, all 4 p53 mutants misplaced promoter binding to all or most genes within the pathways that have been usually certain by WT p53 akin to PDGFRB and AKT1, however much less invasive mutants Y234C and R273H gained binding to a couple genes that weren’t certain by WT p53 (Fig. 7c), implying pathway dysregulation and consequent phenotypic alteration.
Examination of particular person goal genes of mutant p53 proteins supplied insights on the potential regulatory motion of every mutant. For instance, TFAP2A was a binding goal of the least invasive Y234C mutant and displayed a twofold enhance in mRNA expression in Y234C cells, in settlement with a report that overexpression of this gene associates with much less cell migration and invasion71. CASZ1 gene promoter was certain solely by R273H, and the expression was particularly abolished in R273H cells. CASZ1 encodes a zinc finger transcription issue and a tumor suppressor72,73, and the somatic mutations in breast most cancers have been related to poor survival (Supplementary Fig. 17).
In abstract, our most cancers phenotype characterization and built-in evaluation on the molecular degree reveal heterogeneous mobile phenotypes induced by totally different p53 missense mutant proteins that may be attributed partly to the mutant-specific modifications in downstream goal gene expression through each direct and oblique regulatory mechanisms by altered DNA binding functionality and choice.