Tissue construction and mobile composition of indolent and aggressive LUADs at early levels
Histological evaluation of the tumors within the cohort recognized 7 strong, 6 acinar, 1 papillary, 1 mucinous acinar, and 1 lepidic histological subtypes (Fig. 1B, Supplementary Desk T1). A part of the samples (6 of 16) had a blended construction with 1 predominant structural sample. In our cohort, tumor samples with predicted aggressive habits had strong (2 of seven), acinar (3 of seven), papillary (1 of seven), and mucinous acinar (1 of seven) patterns (Fig. 1B). Indolent LUADs had strong (5 of 9), acinar (3 of 9) and lepidic (1 of 9) predominant histological patterns (Fig. 1B, Supplementary Desk T1). A broad number of histological subtypes within the LUADs complicates the utilization of histological patterns alone for the prediction of LUAD habits, particularly at early levels. Nevertheless, we noticed elevated collagen deposition in LUADs with predicted aggressive habits (Fig. 1C). It’s well-known that CAFs play a vital position within the processes of ECM deposition and transforming throughout tumor improvement. We determined to research the CAF subpopulations within the tumor microenvironment of indolent and aggressive LUADs utilizing scRNAseq.
A droplet-based scRNAseq methodology (10× Genomics) was used to carry out a complete evaluation of the CAF inhabitants within the tumor microenvironment. This evaluation recognized 43,207 cells with distinct transcriptomes. 21,140 cells (48.93%) have been obtained from indolent LUADs and 22,067 (51.07%) have been from aggressive LUADs. Utilizing graph-based uniform manifold approximation and projection algorithm (UMAP)20 and Leiden clustering algorithm21, 26 cell clusters have been recognized. A set of canonical cell markers have been used to assign all detected cell clusters to 10 cell populations. The next cell populations have been recognized: epithelial/most cancers cells (EPCAM+, CK18+), endothelial cells (PECAM1+, VWF+), stromal cells (LUM+, DCN+), mast cells (CPA3+, TPSAB1+), myeloid cells (LYZ+, CD68+), CD8+ T cells (CD3D+, CD8A+), NK cells (FCGRA1+, NKG7+), B-cells (CD79A+), CD4+ T-cells (CD3D+, CD4+), and plasma cells (CD79A+, MZB1+) (Fig. 1D). The stromal inhabitants was composed of 986 cells from indolent LUADs (57.2%) and 673 from aggressive tumors (42.8%). To determine the CAF inhabitants, an extra clustering of the stromal cell inhabitants was carried out, which recognized 5 further subclusters of stromal cells (Fig. 1E). The comparability of the proportions of stromal cell clusters demonstrated excessive heterogeneity between affected person samples in addition to throughout the indolent and aggressive teams. (Fig. 1F,G).
CAF inhabitants in indolent and aggressive LUADs is comprised of three distinct subpopulations
To categorise the subpopulations of stromal cells and distinguish clusters of CAFs, the expression of LUM, COL1A1, COL3A1, MMP2, ACTA2, FAP, S100A4, PDGFRA, PDPN, COL4A1, DES, CAV1, NDUFA4L2, MYH11, RGS5, MAP1B, EGFL6, and MFGE8 was analyzed. When coupled with hierarchical clustering, the information indicated that clusters 2 and 4 have gene expression patterns related to pericytes and easy muscle cells (Fig. 2A). Cluster 4 is characterised by excessive expression of RGS5 and TINAGL1, that are well-known markers of pericytes. Cluster 2 demonstrated expression of DES and MYH11 and was acknowledged as easy muscle cells (Fig. 2B,C). We observed {that a} small variety of cells from stromal clusters 2 and 4 specific a number of CAF markers (equivalent to LUM and COL1A1). This may be defined by mobile plasticity of stromal cells, purposeful convergence, and comparable microenvironmental influences. Clusters 0, 1, and three have been decided to be CAFs. Additional exploration confirmed that 663 CAFs (62.8%) have been obtained from indolent group and 393 CAFs (37.2%) have been obtained from aggressive group. Cluster 0 was assigned as FAP+ACTA2+ CAFs. Cluster 1 was recognized as ADH1B+ CAFs. Since regular fibroblasts additionally specific ADH1B, we employed one other marker of regular fibroblasts, CD10 (MME), to corroborate that the recognized ADH1B+ inhabitants consists of CAFs. Our findings revealed that the ADH1B+ cluster lacks or demonstrates very low CD10 expression, additional confirming that the cells inside this cluster are certainly CAFs and never regular fibroblasts (Supplementary Fig. S1). This method has additionally been utilized by Grout et al.31. Cluster 3 was assigned as CTHRC1+FAP– CAFs (Fig. 2D). We additionally observed heterogeneity in CAF cluster composition amongst sufferers in each the indolent and aggressive teams. This variability would possibly point out totally different ranges of organic processes associated to CAF capabilities within the tumor microenvironment, such because the deposition and transforming of ECM, modulation of immune processes, and angiogenesis (Fig. 2D,E).
CAF inhabitants in indolent and aggressive LUADs. (A) Expression of canonical genes related to CAFs, easy muscle cells, and pericytes. Dendrograms are generated in accordance with hierarchical clustering. Heatmap was carried out utilizing Python 3 (https://www.python.org). (B) UMAP illustration of stromal cell subpopulations. Subpopulations of CAFs, pericytes, and easy muscle cells are highlighted by shade. (C) Expression of canonical marker genes of CAFs (FAP+ACTA2+; ADH1B+, CTHRC1+FAP–), pericytes (RGS5+TINAGL+), and easy muscle cells (DES+MYH11+) in stromal cell subpopulations. UMAP representations present expression of canonical marker genes throughout recognized stromal cell clusters. Violin plots present expression of genes throughout recognized stromal clusters. (D) UMAP illustration of CAF populations. Recognized subclusters in CAF inhabitants (prime left). Distribution of cells from sufferers with indolent and aggressive LUADs inside CAF inhabitants (prime proper). Distribution of cells from totally different affected person samples inside stromal cell subpopulation (backside left). Distribution of cells from totally different histological subtypes inside CAF inhabitants (backside proper). (E) Proportion of CAF subpopulations in relation to sufferers with indolent and aggressive LUADs (scRNAseq information).
CAF subpopulations are characterised by totally different operate and position within the tumor development and aggressiveness
The subpopulation of FAP+ACTA2+ CAFs was composed of 276 cells (59.6%) from indolent LUADs and 187 cells (40.4%) from aggressive carcinomas. The cluster of ADH1B+ cells was shaped by 234 (71.8%) and 92 (28.2%) CAFs from the indolent and aggressive tumors, respectively. The group of CTHRC1+FAP− CAFs consists of 153 cells (56.2%) from indolent and 119 cells (43.8%) from aggressive LUADs. The comparability of the common proportions between various kinds of CAFs revealed that aggressive LUADs are characterised by a decreased proportion of cells that kind the ADH1B+ CAF cluster (p = 0.032) (Fig. 3A).
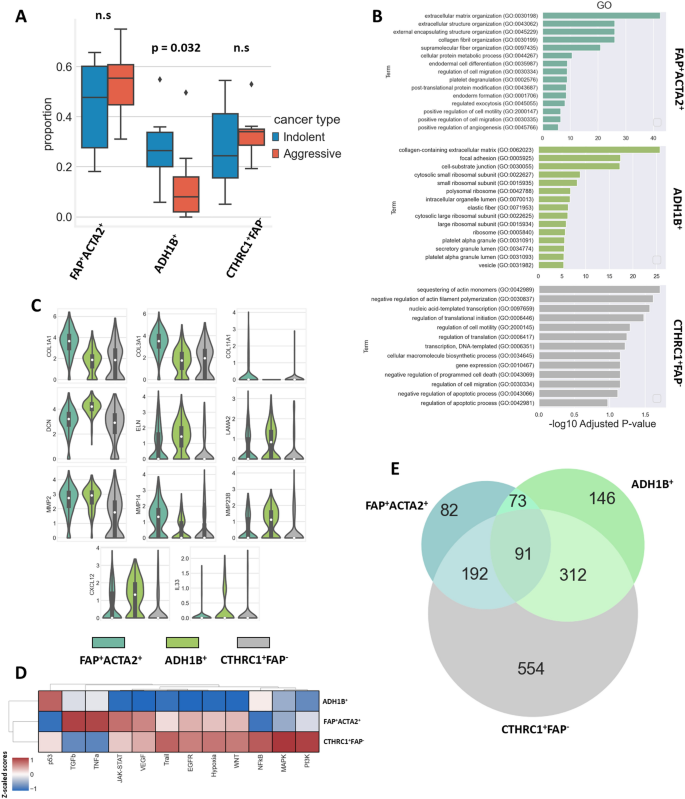
CAF subpopulation proportions and capabilities. (A) Proportion of CAF subpopulations in indolent and aggressive LUADs (indolent n = 6, aggressive n = 7, p = 0.032, outliers are highlighted by diamonds). (B) GO enrichment evaluation of recognized CAF subpopulations. (C) Violin plots reveal expression of genes related to capabilities of recognized CAF subpopulations. (D) Heatmap demonstrates exercise of canonical signaling pathways throughout recognized CAF subpopulations. Dendrograms are generated in accordance with hierarchical clustering. Heatmap was carried out utilizing Python 3 (https://www.python.org). (E) Venn diagram reveals intersection of differentially expressed genes in recognized CAF subpopulations.
Gene ontology (GO) enrichment evaluation signifies that CAFs from the FAP+ACTA2+ cluster have been principally concerned in processes related to the deposition of ECM proteins and matrix transforming (Fig. 3B). Certainly, that subpopulation of CAFs demonstrated elevated expression of COL1A1, COL3A1, COL11A1, and FN1. Additionally, this cluster was characterised by excessive expression of MMP2 and MMP14 (Fig. 3C). As well as, the evaluation of genes related to totally different signaling pathways confirmed that the FAP+ACTA2+ cluster was characterised by elevated exercise of TGFβ pathway (Fig. 3D).
GO enrichment evaluation signifies that the ADH1B+ subpopulation additionally has been concerned within the deposition and transforming of ECM buildings (Fig. 3B). Nevertheless, compared to the FAP+ACTA2+ cluster, it had decrease expression of COL1A1, COL3A1, and MMP14. However, we recognized elevated expression of ELN, LAMA2, and DCN in ADH1B+ CAFs. Additional evaluation revealed that the ADH1B+ cluster was additionally characterised by enhanced expression of cytokines equivalent to CXCL12 and IL33 (Fig. 3C). The evaluation of genes related to signaling pathways confirmed that ADH1B+ cluster was characterised by decreased exercise of JAK-STAT, VEGF, Path, EGFR, hypoxia, and WNT pathways compared to different CAF clusters (Fig. 3D).
Elevated exercise of the JAK-STAT, VEGF, Path, EGFR, hypoxia, and WNT signaling pathways was noticed in each the CTHRC1+FAP– and FAP+ACTA2+ CAF subpopulations. However, the CTHRC1+FAP– subpopulation had decreased exercise of the TGFβ and TNFa pathways, resembling the exercise of the ADH1B+ CAFs (Fig. 3D). The Venn diagram reveals that CTHRC1+FAP– CAFs share 283 and 403 comparable differentially expressed genes with FAP+ACTA2+ and ADH1B+ CAFs respectively, whereas FAP+ACTA2+ and ADH1B+ subpopulations solely share 164 overlapping genes (Fig. 3E). The expression of COL1A1 and COL3A1 within the ADH1B+ cluster was near that of the CTHRC1+FAP– cluster. The expression of DCN, ELN, and LAMA2 in FAP+ACTA2+ CAFs was just like that of CTHRC1+FAP– CAFs. The expression of CXCL12 and IL33 cytokines within the CTHRC1+FAP– subpopulation was decreased compared to that of each the ADH1B+ and FAP+ACTA2+ clusters (Fig. 3C). Genes related to iCAF, myCAF, and apCAF profiles revealed that each one 3 recognized CAF subpopulations exhibit similarity to myCAF cells (Supplementary Fig. S2). Altogether, the evaluation of the gene expression profile, the exercise of signaling pathways, and the Venn diagram of overlapping differentially expressed genes throughout subtypes might point out that the three noticed subpopulations of CAFs may very well be transient types of one another. However, this query requires additional investigation.
ADH1B+ CAF markers are prognostic for sufferers with lung adenocarcinoma at early levels
Because the proportion of ADH1B+ subpopulation within the indolent group was considerably elevated, we determined to focus our consideration on this subtype of CAFs. A rating of the differentially expressed genes was carried out and, along with ADH1B, this rating recognized CFD, DCN, SEPP1 (SELENOP), A2M and MFAP4 as prime marker genes of this subpopulation of cells (Fig. 4A). As well as, the expression of those genes amongst different populations of cells and different CAF subpopulations was assessed. This expression evaluation confirmed that myeloid cells are characterised by comparatively excessive expression of CFD. Furthermore, excessive expression of SEPP1 (SELENOP) was additionally noticed in endothelial and epithelial/most cancers populations. All CAF subpopulations expressed ADH1B, CFD, DCN, SEPP1 (SELENOP), A2M, and MFAP4, however ADH1B+ CAFs demonstrated dramatically increased expression of those genes compared to different CAFs subtypes (Fig. 4A,B, Supplementary Fig. S3A).
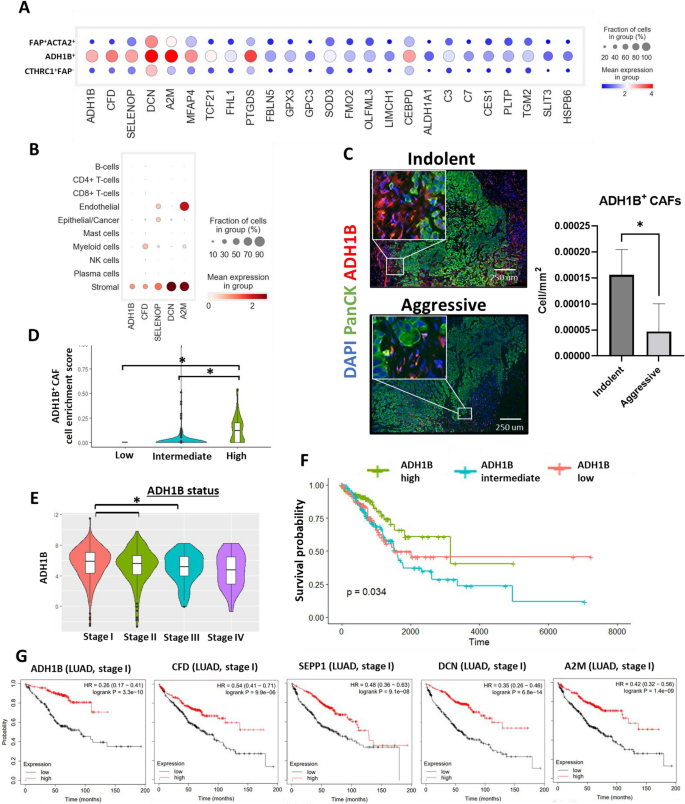
ADH1B+ CAF markers are prognostic for sufferers with lung adenocarcinoma at early levels. (A) Expression evaluation of the highest 25 differentially expressed genes associated to the ADH1B+ CAF cluster throughout all CAF clusters. (B) Gene expression of ADH1B+ CAF markers throughout all recognized subpopulations of cells. (C) Consultant photos of MxIF staining displaying expression of PanCK and ADH1B in LUAD tissue remoted from sufferers with predicted indolent and aggressive tumor habits. Corresponding mobile density of ADH1B+ CAFs (cell/mm2) within the LUAD tissue extracted from the sufferers with predicted indolent or aggressive tumor habits (indolent n = 7, aggressive n = 6, *—p < 0.05, information confirmed as imply ± SEM). (D) Cell enrichment evaluation (CIBERSORT) of LUAD affected person samples (stage I and II, TCGA dataset). Violin plots reveal ADH1B+ CAF enrichment scores in TCGA samples with low (n = 105), intermediate (n = 209), and excessive (n = 105) expression of ADH1B (*p < 0.05). (E) Gene expression stage of ADH1B in LUAD samples (TCGA dataset) at totally different levels of improvement (unbiased samples, stage I n = 294, stage II n = 125, stage III n = 84, stage IV n = 26, *p < 0.05). (F) Kaplan–Meier evaluation of the general survival of LUAD sufferers with low (n = 105), intermediate (n = 209), and excessive (n = 105) expression of ADH1B (stage I and II, TCGA dataset). (G) Kaplan–Meier evaluation (KM-plotter information) of the general survival of sufferers with LUADs at stage I and characterised by high and low expression of prime marker genes related to ADH1B+ CAF subpopulation (ADH1B (low n = 185, excessive n = 185), CFD (low n = 288, excessive n = 289), SEPP1 ((low n = 288, excessive n = 289)), DCN (low n = 288, excessive n = 289), and A2M (low n = 288, excessive n = 289).
To check the variety of ADH1B+ CAFs in indolent and aggressive LUADs MxIF staining was carried out. The outcomes obtained by scRNAseq steered that ADH1B by itself can act as a marker for the identification of this subpopulation of CAFs as its expression could be very low in different populations of cells. As we anticipated, the variety of ADH1B+ fibroblasts within the tumor tissue of indolent LUADs was considerably increased (FC = 4.02, p < 0.05) compared to the aggressive group, supporting the outcomes obtained utilizing scRNAseq (Fig. 4C). Subsequent, we now have used bulk RNAseq dataset obtained from TCGA and carried out cell enrichment evaluation utilizing the CIBERSORT algorithm to estimate cell enrichment rating of ADH1B CAFs. We discovered that the expression of ADH1B positively correlates with ADH1B+ CAF enrichment rating (Fig. 4D). Moreover, we noticed a optimistic correlation between the expression of ADH1B and different recognized ADH1B+ CAF markers, together with CFD (DF), A2M, MFAP4, SEPP1 (SELENOP), and DCN (Supplementary Fig. S3B).
ADH1B expression could be very particular for fibroblast-like cells within the LUAD tumor microenvironment that makes it some of the necessary markers for the detection of ADH1B+ CAFs (Fig. 4B; Supplementary Fig. S3A). Gene expression of ADH1B in LUADs is related to tumor improvement and reduces as tumor progresses (Fig. 4E). We additionally recognized a lower within the variety of ADH1B + CAFs in stage II LUADs in comparison with stage I LUADs (Supplementary Fig. S3C). Survival evaluation demonstrated that top expression of ADH1B is related to higher survival of LUAD sufferers (stage I and II, p = 0.034) (Fig. 4F).
Additional evaluation recognized that different prime marker genes of ADH1B+ CAFs are related to higher survival of LUAD sufferers at early stage. We utilized KM-plotter software (https://kmplot.com) for this objective. Sufferers have been stratified into two teams based mostly on the expression of a number of marker genes (excessive and low): ADH1B, CFD (DF), A2M, SEPP1 (SELENOP), and DCN. The median expression was used as separation level. Sufferers with excessive expression of ADH1B (logrank p < 0.01, HR = 0.26), CFD (DF) (logrank p < 0.01, HR = 0.54), SEPP1 (SELENOP) (logrank p < 0.01, HR = 0.48), DCN (logrank p < 0.01, HR = 0.35), and A2M (logrank p < 0.01, HR = 0.42) demonstrated higher survival at stage I of LUAD (Fig. 4G). Altogether these outcomes point out that the presence of ADH1B+ CAFs within the LUAD tissue is related to the indolent habits of the tumor and higher general survival of sufferers.
Organic processes, immune cell signature, and ECM deposition processes related to ADH1B+ CAF tissue enrichment
Since we recognized that ADH1B expression positively correlates with ADH1B+ CAF enrichment (Fig. 4D), and it is rather particular marker for ADH1B+ CAFs within the tumor microenvironment (Fig. 4A, Supplementary Fig. S3A), we determined to stratify TCGA samples into ADH1B low, intermediate and excessive teams. To research gene expression profile of LUAD samples with high and low expression of ADH1B, we carried out differential gene expression evaluation. We discovered ADH1B+ CAF markers equivalent to A2M and MFAP4 among the many prime upregulated genes in ADH1B excessive group. Furthermore, this group of LUAD samples demonstrated the improved expression of genes related to higher prognosis equivalent to SFTPC, SCGB3A2, GRIA1, CYP4B1, RSPO2, CD300LG, ADAMTS8, and TMEM132C (Fig. 5A, Supplementary Desk T2). However, the group of sufferers with low expression of ADH1B was characterised by upregulation of pro-tumorigenic genes equivalent to MYBL2, UBE2C, CENPA, TROAP, KIF18B, TPX2, HJURP, KIF2C, NEK2, and COL11A1 (Fig. 5A, Supplementary Desk T2). We then carried out GO enrichment evaluation for the highest genes upregulated within the group of samples characterised by excessive ADH1B expression. GO—organic course of evaluation revealed enrichment within the mechanisms associated to cell–cell sign transduction, tissue improvement, and group of extracellular construction together with extracellular matrix. GO—cell composition confirmed enrichment primarily in collagen-containing extracellular matrix and cell membrane construction. GO—molecular operate was related primarily with binding to sulfur compounds, heparin, and glycosaminoglycans (Fig. 5B).
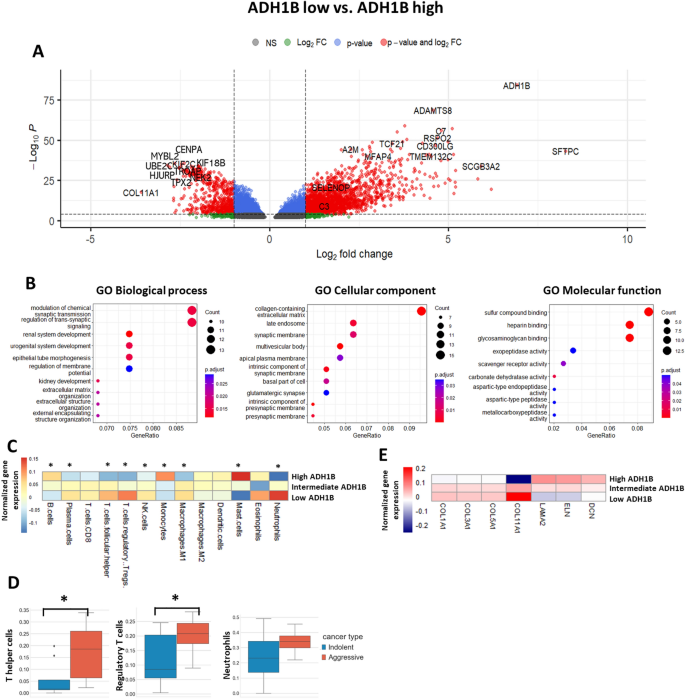
Organic processes, immune cell signature, and ECM deposition processes related to presence of ADH1B+ CAF within the LUAD tissue. (A) Volcano plot reveals the differentially expressed genes amongst teams of LUAD sufferers (levels I and II, TCGA dataset) with low (n = 105) and excessive (n = 105) expression of ADH1B. (B) Gene ontology enrichment evaluation of differentially expressed genes in relation to organic processes, mobile element, and molecular operate (TCGA dataset). (C) Heatmap demonstrates immune cell enrichment scores (CIBERSORT) within the samples with low (n = 105), intermediate (n = 209), and excessive (n = 105) expression of ADH1B (*—p < 0.05, low vs. excessive, TCGA dataset). Heatmap was carried out utilizing R 4.2.1 (https://www.r-project.org). (D) Proportions of regulatory T cells, T helper cells, and neutrophils (scRNAseq information) in LUAD affected person samples with predicted indolent (n = 6) and aggressive (n = 7) tumor habits (*—p < 0.05). (E) Expression of ECM proteins within the samples with low (n = 105), intermediate (n = 209), and excessive (n = 105) expression of ADH1B (TCGA dataset). Heatmap was carried out utilizing R 4.2.1 (https://www.r-project.org).
The following query to be analyzed was whether or not ADH1B expression in LUADs correlates with anti-cancer immune response. To estimate immune cell signature in TCGA samples, we now have utilized the CIBERSORT algorithm. We discovered that cell enrichment rating of plasma cells, regulatory T cells, T follicular helper cells, NK cells, and neutrophils was considerably increased in ADH1B low group. However, the rating of B cells, monocytes, and mast cells was increased in ADH1B excessive group (Fig. 5C). These outcomes point out that ADH1B low samples are characterised by considerably extra lively inflammatory processes. Curiously, scRNAseq information revealed that aggressive LUADs are characterised by elevated proportion of T helper cells, regulatory T cells, and neutrophils, which is in line with the information obtained utilizing the CIBERSORT algorithm (Fig. 5D).
As well as, we estimated gene expression of various ECM proteins in TCGA samples characterised by low, intermediate, and excessive expression of ADH1B. We found that LUADs with excessive expression of ADH1B have been characterised by decreased expression of COL1A, COL3A1, COL5A1, and COL11A1. However, the expression of ELN, LAMA2, and DCN was elevated (Fig. 5E). These outcomes help and supply further validation of our scRNAseq findings. Altogether, it highlights the potential position of ADH1B+ CAFs within the dedication of LUAD habits at early levels by regulation of ECM transforming and modulation of immune response.