We investigated 230 grownup T-ALL sufferers enrolled in GMALL registries and trials with a median age of 32 years (vary 17-83 years); of those 11 sufferers (5%) had been older than 55 years. 166 (72%) sufferers had been male: 107 (46.5%) sufferers confirmed an immunophenotype of a thymic T-ALL, 43 (18.7%) of a mature T-ALL, and 60 (26.1%) of an early T-ALL, for 20 sufferers immunophenotype was lacking (Desk 1).
Transcriptome sequencing reveals 9 molecular subgroups in grownup T-ALL
For the molecular subgroup task, we constructed a category prediction mannequin primarily based on two cohorts. One a part of the investigated cohort served as an inside reference cohort, consisting of 169 of the 230 grownup T-ALL GMALL samples. This cohort was beforehand assigned primarily based on oncogene expression and served as a coaching cohort. As well as, an exterior cohort of pediatric T-ALL circumstances (n = 241) printed by Liu and colleagues was used as a second reference cohort [5]. We had been capable of assigned the expected molecular subgroup in 175 samples (76%) primarily based on overlapping predictions utilizing the inner reference cohort and the exterior reference cohort. Of the remaining 55 samples, we assigned extra 11 samples primarily based on their particular fusion genes and 23 samples manually categorized by a transparent oncogene overexpression and similarity of gene expression profiles to these of already outlined samples (Supplementary Fig. S1). Twenty-one of 230 T-ALL samples remained unclassified, representing 9.1% of the full cohort (Fig. 1).
The most important molecular subgroup in grownup T-ALL was the HOXA subgroup outlined by overexpression of HOXA genes in 57 samples (24.7%) (Fig. 1, Supplementary Desk S4). Within the majority of those circumstances early stem cell markers like MEF2C and BAALC had been co-overexpressed in comparison with the expression in subgroups TLX1, TAL1/LMO, TLX3 and LMO1 (p < 0.001). The expression of stem cell genes can also be an indicator of the immature subgroup LYL1/LMO2, protecting 32 samples (13.9%) and missing expression of HOXA cluster genes. Samples with TLX1 overexpression comprised 44 samples (19.1%) harboring the defining TLX1 overexpression. Eleven of those TLX1 samples expressed NKX2, much like the NKX2 subgroup comprising 4 samples (1.7%) missing TLX1 overexpression. Of the TAL1/LMO subgroup (in complete 45 samples), 15 samples (6.5%) had an LMO1 overexpression, whereas the remaining 30 samples (13%) had been missing LMO1 expression. Consecutively, we outlined these two teams LMO1 and TAL1/LMO as separate entities. Whereas samples of the TLX1 and TLX3 subgroup confirmed some HOXA expression, TAL1/LMO and LMO1 circumstances lacked overexpression of HOXA genes. TLX3 overexpression was present in 26 samples (11.3%) defining the TLX3 subgroup with absence of expression of TLX3 in all different subgroups. We recognized a single pattern with TAL2 overexpression. Notably, no fusion or overexpression of SPI1 was detected in our cohort.
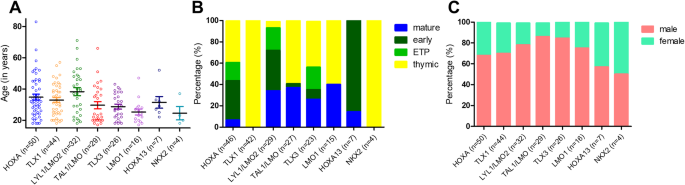
The typical age ranged from 24 years to 38 years throughout the molecular cohort and displayed highest in LYL1/LMO2 with a median of 39 years and lowest in NKX2 with a median of 21.5 years (Fig. 2A). In our cohort, age distribution revealed youthful sufferers with TAL1/LMO overexpression (16-25 years: 35% versus >35 years: 3%; p = 0.001) and extra LYL1/LMO2 and HOXA overexpression amongst older sufferers (16–25 years: 23% vs. >35 years: 40%; n.s.).
The subgroups indicated a transparent correlation with immunophenotype reflecting the T-cell growth stage (Fig. 2B). All sufferers within the TLX1 and NKX2 subgroup revealed a thymic immunophenotype (p < 0.001). LYL1/LMO2 (p = 0.017) and HOXA (p = 0.05) had been characterised by a extra immature phenotype, exhibiting floor expression of antigens typical for an early T-ALL in 50% of samples. TLX3 was not related to a selected immunophenotype on this cohort with 23 samples (early: n = 7, thymic: n = 10, mature: n = 6, n.s.). Alternatively, TAL1/LMO was related to extra mature T-cell growth levels, mirrored by a thymic and mature T-ALL immunophenotype (p = 0.002). Solely a single TAL1/LMO circumstances had an early T-ALL immunophenotype. Twenty-three (34%) of the 67 early T-ALL samples confirmed an ETP-ALL immunophenotype. By be aware, of 23 samples with an ETP-ALL immunophenotype 11 belong to the HOXA molecular subgroups, six had been in LYL1/LMO2, 5 in TLX3, and one couldn’t be categorized. Concerning intercourse distribution, no important variations had been noticed between molecular subgroups (Fig. 2C).
HOXA13 as distinct T-ALL subgroup
Seven sufferers (3% of the full cohort) confirmed a definite overexpression of HOXA13 with related overexpression of the lncRNA HOTTIP and EVX1 (Figs. 1 and 3A). Notably, in these samples the anterior HOXA genes weren’t expressed on transcript degree. In 5 out of the seven samples, fusion genes focusing on the posterior HOXA advanced had been recognized. Three circumstances with recurrent MIR181A1HG::HOTTIP had been brought on by a chromosomal t(1;7) translocation with subsequent overexpression of lncRNA HOTTIP (Fig. 3B). In two distinct detected fusion genes in HOXA13 cluster, HOXA13 and EVX1 had been concerned (Supplementary Desk S5). All 7 samples depicted an early immunophenotype mirrored by lacking sCD3 expression. 4 extra samples confirmed an unique HOXA13 overexpression among the many HOXA genes, however had been categorized within the LYL1/LMO2 subgroup. All 4 samples had been missing EVX1 overexpression.
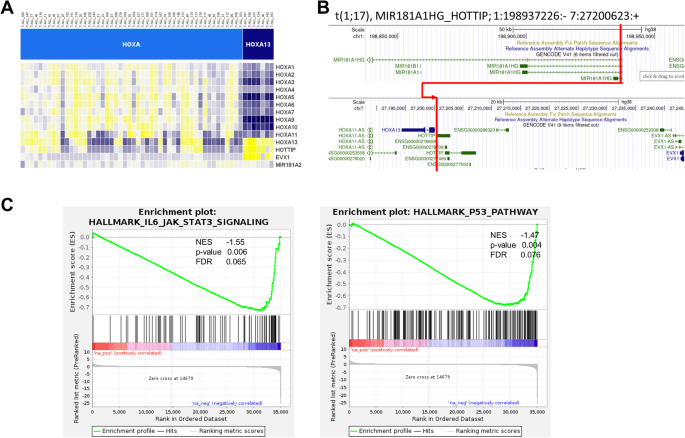
A Expression of posterior HOXA genes in HOXA13 cluster. B Fusion gene mir181A1HG_HOTTIP with the 5′ fusion companion breakpoint after exon companion and full utilization of three′ fusion companion HOTTIP. C Differentially expressed genes between HOXA13 and HOXA cluster. GSEA enrichment evaluation for JAK/STAT and TP53 pathway in contrast between HOXA13 and HOXA.
HOXA13 circumstances confirmed a definite gene expression profile in comparison with the HOXA samples with nearly all of differentially expressed genes being downregulated in HOXA13 cluster, together with anticipated downregulation of anterior HOXA genes (Supplementary Desk S6). In gene set enrichment analyses of KEGG and Hallmark of most cancers gene lists, we recognized 88 differentially regulated gene units (FDR < 0.1, Supplementary Desk S7). Amongst these, JAK-STAT-signaling and TP53 pathway had been altered in HOXA13 in comparison with HOXA (Fig. 3C), which have been related to poor prognosis (TP53) and affiliation to the unfavorable immunophenotype of early T-ALL (JAK/STAT mutations).
Underlying fusion genes drive T-ALL leukemogenesis
In 106 of the 230 T-ALL samples we detected underlying fusion genes in T-ALL (Fig. 1, Supplementary Desk S5). Essentially the most frequent rearrangements in our cohort had been STIL::TAL1 (n = 14) and TLX1-rearrangements (n = 15). SET::NUP214 occurred in 12 samples and was related to the HOXA subgroup. Fusions of NKX2 had been a uncommon occasions in grownup T-ALL sufferers with a price of 1% within the general cohort. Additional recurrent gene fusions affected amongst others KMT2A (n = 6), MLLT10 (n = 5) or LMO2 (n = 2) (Supplementary Desk S5). In LYL1/LMO2 solely few rearrangements had been discovered, underlying the stem cell character of LYL1/LMO2 with similarities to acute myeloid leukemia (AML). Notably, all detected fusions confirmed subgroup task primarily based on gene expression.
Mutational spectrum of T-ALL in keeping with molecular subgroups
The completely different composition of molecular subgroups was mirrored by the mutational spectrum of grownup T-ALL. We investigated full protein coding sequence of 206 leukemia-associated genes (Supplementary Desk S3). Twenty-two of those genes had been affected a minimum of in 3% of all samples. Essentially the most regularly altered gene was NOTCH1 with 42 out of 83 (51%) mutated samples adopted by PHF6 (31/83 samples; 37%) and PTEN (15/83 samples; 18%). The small pattern sizes within the molecular subgroups made agency comparisons between them troublesome (see Supplementary Tables S8 and S9). Nevertheless, a pattern indicated that the TLX1 subgroup has the next price of NOTCH1 mutations, with 15 out of 21 samples (71%) being mutated, in comparison with the extra immature molecular subgroups equivalent to LYL1/LMO2 (5/12; 42% NOTCH1 mutated) or HOXA (4/12; 33% NOTCH1 mutated; not important). Alternatively, genes affecting the JAK/STAT pathway (JAK1, JAK3, STAT5B, SH2B3, IL7R), had been extra regularly affected within the HOXA (5/12, 42%) and LYL1/LMO (7/12, 58%) subgroups in comparison with the TLX1 subgroup (6/21, 29%). Nevertheless, these variations remained descriptive and didn’t attain statistical significance.
As well as, for 78 samples (all TAL1 and LMO1 samples and unclassified samples), we investigated the mutational standing of the enhancer area of TAL1 with a beforehand reported mutation within the non-protein-coding area [31]. We solely discovered two mutations (2/78, 3%) within the investigated cohort, each in samples of the molecular subgroup TAL1. As it’s unlikely to seek out enhancer mutations in non-TAL1 overexpressing samples, the speed of 1% referring to the full grownup T-ALL cohort appears decrease than reported for pediatric sufferers.
Methylation patterns revealed hypomethylation in STIL::TAL1 subgroup
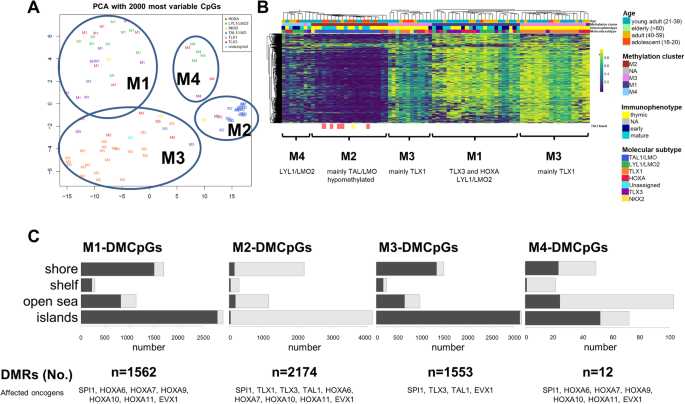
In 84 investigated samples, 4 clusters had been recognized primarily based on unsupervised clustering, reflecting molecular subgroups on DNA methylation degree (Fig. 4A, Supplementary Fig. S2). Cluster M1 consisted of 25 samples, comprising 8 out of 10 TLX3 samples, 8 LYL1/LMO2 samples and eight HOXA samples. Just one NKX-1 pattern was included. When it comes to immunophenotype, cluster M1 mirrored a extra heterogeneous image comprising high-risk options in keeping with GMALL protocols (10 mature T-ALL samples and eight out of 12 samples with an early immunophenotype; Fig. 4B). Cluster M2 included 20 samples, practically solely TAL1/LMO samples (n = 18). Vice versa 18 of the 20 TAL1/LMO2 samples had been assigned into the M2 cluster, together with all samples with a TAL1 fusion (STIL::TAL1 fusion n = 5; TCF7::TAL1 fusion n = 1) exhibiting a big international hypomethylation in CpG islands (Fig. 4C). The most important cluster, M3, included 30 samples. Remarkably, 29 samples on this cluster confirmed a thymic immunophenotype and all TLX1 samples (n = 20) demonstrated a DNA methylation sample consultant of cluster M3. Within the remaining samples of cluster M3 we discovered 6 HOXA samples, and one TLX3, NKX2-1, and TAL1/LMO pattern every. The fourth cluster M4 revealed to be the smallest with solely 7 samples (4 LYL1/LMO2, 2 HOXA and 1 TLX3 samples). Taken collectively, DNA methylation profiling enabled a strong subclassification of grownup T-ALL samples. TLX1 and TAL1/LMO subgroups had been characterised by a homogenous frequent DNA methylation sample mirrored by the task in the identical methylation cluster (TLX1: 20/20 in M3, TAL1/LMO: 18/20 in M2), separating each subgroups from different molecular subgroups. Alternatively, TLX3, HOXA and LYL1/LMO2 had frequent DNA methylation patterns in cluster M1 and M4, related to an early immunophenotype (early T-ALL: 17/19 in M1 and M4 vs. 2/19 in M2 and M3, p < 0.0001). Among the many differentially methylated areas (DMRs) for every of the 4 clusters, defining oncogenes of the corresponding molecular subgroups had been discovered (Fig. 4C, Supplementary Tables S10 and S11). The affiliation of mutational occasions with the methylation clusters revealed a excessive price of NOTCH1 mutations in cluster M3 and an elevated price of mutations within the JAK/STAT pathway and epigenetic regulators in M1 and M4 clusters (knowledge not proven).
A Principal element evaluation of 84 T-ALL samples primarily based on their methylation standing utilizing the 2000 most variant CpGs leading to 4 secure clusters (M1–M4). B Methylation clusters confirmed a excessive concordance with molecular subgroups and immunophenotype. Methylation degree of the 2000 most variant CpGs are offered with a transparent hypomethylation for cluster M2. Under the graph most distinguished molecular subgroups within the methylation clusters are depicted. Samples with an underlying TAL1 fusion are marked. C Variety of differentially methylated areas and their methylation standing in keeping with the 4 methylation subgroups. Genes within the differentially methylated areas are reflecting driving oncogenes of molecular T-ALL subgroups.
MRD response in keeping with molecular subgroup in grownup T-ALL
With respect to response, 95 (68%) sufferers with a molecular marker achieved a molecular CR (MolCR) after consolidation I, 25 (18%) sufferers confirmed an intermediate molecular response (IntMR) and 19 (14%) failed molecular remission (MolFail) (Desk 2). Within the subgroup of thymic T-ALL, 65 sufferers (79%) achieved MolCR, 9 (11%) sufferers IntMR and eight (10%) sufferers failed. Between the molecular subgroups, MRD after consolidation I differed considerably. Noteworthy, 37/39 (95%) of TLX1 sufferers had a MolCR with solely two sufferers in TLX1 subgroup with a MolFail. On the opposite facet, in LYL1/LMO2 solely 3/9 (33%) received into MolCR. Equally, TLX3 and TAL1/LMO subgroup confirmed a considerably worse response with a price of solely 50% of MolCR (Desk 1). HOXA subgroups confirmed an intermediate response with a price of 65% (20/37 sufferers) of MolCR. These response charges of the molecular subgroups are comparable in sufferers with a thymic immunophenotype regardless of their favorable CD1a expression: MolCR price in HOXA 73%, in TLX3 60% and in TAL1/LMO 40% with solely restricted variety of sufferers. Thymic T-ALL sufferers with a TLX1 or NKX2 profile confirmed glorious MRD response (95% resp. 100% of MolCR; Desk 2).
Favorable end result in grownup T-ALL is related to TLX1, NKX2, and LMO1 subgroups
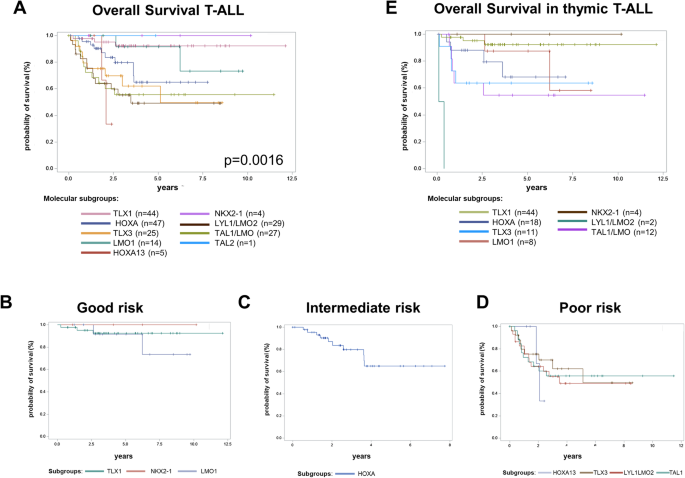
MRD response translated in OS with variations among the many molecular subgroups. OS after three years differed considerably throughout the molecular subgroups in grownup T-ALL (p = 0.0016, Fig. 5A). Among the many bigger T-ALL molecular subgroups, TLX1 sufferers (n = 44) confirmed an distinctive favorable 3y-OS of 92%. Though the pattern dimension is simply too small to attract agency conclusions, within the small teams of NKX2-1 (n = 4) and TAL1 (n = 1), all sufferers had been alive after three years. As well as, sufferers of the LMO1 subgroup (n = 14) had a really favorable 3y-OS of 92%. Sufferers of the HOXA (n = 47) and of the TLX3 (n = 25) subgroups with a 62% 3y-OS confirmed a barely inferior prognosis (70% 3y-OS and 62% 3y-OS, respectively). In distinction, the subgroups LYL1/LMO2 (55% 3y-OS, n = 29), HOXA13 (33% 2y-OS, n = 5) and TAL1 (with out LMO1 overexpression, 56% 3y-OS, n = 27) had a poorer prognosis.
(A) Total survival for the molecular subgroups in 196 grownup T-ALL sufferers impartial of their immunophenotype. Unassigned samples usually are not displayed. Colours for the completely different subgroups are proven under the determine. B–D Total survival of grownup T-ALL sufferers in keeping with their molecular threat classification. Good threat group includes 62 sufferers: TLX1 (n = 44), NKX2-1 (n = 4) and LMO1 (n = 14); intermediate threat group 47 sufferers: HOXA (n = 47); poor threat subgroup 76 sufferers: HOXA13 (n = 5), TLX3 (n = 25), LYL1/LMO2 (n = 29), TAL1/LMO2 (n = 27). E Total survival for the molecular subgroups in 99 grownup T-ALL sufferers with thymic immunophenotype. Unassigned samples usually are not displayed. Colours for the completely different subgroups are proven under the determine.
Primarily based on our cohort, we had been capable of classify T-ALL sufferers primarily based on their molecular subgroups into good threat (TLX1, NKX2-1, LMO1), intermediate threat (HOXA), and poor threat (LYL1/LMO2, HOXA13, TAL1/LMO, TLX3). (Fig. 5 B-D). The favorable influence of TLX1 (92% 3y-OS, n = 44), LMO1 (89% 3y-OS, n = 8) and NKX2-1 (100% 3y-OS, n = 4) was not solely discovered within the general cohort but in addition throughout the already good threat subgroup of thymic T-ALL sufferers (Fig. 5E). In thymic T-ALL, fewer sufferers had been assigned to the unfavorable subgroups of LYL1/LMO2 (0% 3y-OS, n = 2), TAL1 (55% 3y-OS, n = 12), TLX3 (64% 3y-OS, n = 11) and HOXA (79% 3y-OS, n = 13) in comparison with mature and early T-ALL. Though the relative incidences of those subgroups had been decrease, 45% of thymic T-ALL belonged to the HOXA, TLX3, TAL1 or LYL1/LMO2 subgroup. Due to this fact, a related share lacked the great prognosis. The above established molecular threat classification (Fig. 5 B–D) additionally allowed to stratify till now customary threat thymic T-ALL sufferers in keeping with their molecular subgroup into distinct threat teams (good threat: 92% 3y-OS, intermediate threat 79% 3y-OS, poor threat 58% 3y-OS, p = 0.0022, Supplementary Fig. S3).