Research design and evaluation overview
We carried out single-cell transcriptome evaluation on bone marrow (BM) samples collected from 16 newly identified APL sufferers (Fig. 1a; additionally see “Strategies” part for affected person choice and Supplementary Information 1 for detailed medical traits and pattern data). This cohort included 4 sufferers who skilled early loss of life, which is outlined as loss of life occurring inside 30 days from prognosis. On this endeavor, we generated a complete of 136,497 cells by combining 16 separate scRNA-seq datasets of APL BM samples at illness onset, forming the inspiration for a complete understanding of APL mobile composition. In parallel, we reanalyzed 23 separate scRNA-seq knowledge of regular BM samples from wholesome people (Gene Expression Omnibus with accession IDs of GSE120221 and GSE130116) to assemble regular hematopoietic cell populations (totaling 102,792 cells) to be used as controls. We adjusted for batch results and carried out an built-in evaluation of each APL and regular cell populations to characterize the malignant APL blasts and intratumoral heterogeneity (detailed in “Strategies” part).
Primarily based on the APL blasts characterised on this examine, we designed a collection of analyses and validations as follows: (i) we constructed the mobile structure and differentiation trajectory of APL blasts, with a particular deal with figuring out APL stem-like cells; (ii) we validated the expression of the PML/RARα fusion gene and FLT3-ITD utilizing focused scRNA-seq (scTarget) in two de novo APL sufferers; (iii) we decided the affiliation between mobile compositions and medical displays, together with the incidence of early loss of life, by conducting deconvolution evaluation on a big cohort of 323 APL sufferers (together with 22 sufferers with early loss of life); (iv) we explored the impression of ATRA remedy on adjustments in mobile compositions for 3 sufferers, each at illness onset and on Day 2 after ATRA remedy; and (v) we carried out deconvolution evaluation on RNA-seq knowledge from 10 newly identified APL sufferers earlier than and after ATRA remedy to ascertain the affiliation of mobile compositions, particularly APL stem-like cells, with ATRA responses in vivo.
Single-cell characterization of APL blasts
To unravel the mobile range inside APL, we first carried out evaluation by evaluating BM samples collected from APL sufferers with these from wholesome people. Utilizing 23 regular BM samples, we established the baseline of mobile range, which revealed seven main cell populations, per beforehand printed findings23,24. These populations included hematopoietic stem/progenitor cells (HSPCs), GMPs, monocytes (Mono), dendritic cells (DCs), B cells, T/Pure Killer (NK) cells, and erythroid (Ery) cells (detailed in “Strategies” part and Supplementary Fig. 1).
Subsequent, we employed UMAP to challenge 136,497 cells from 16 newly identified APL sufferers, together with regular hematopoietic cell populations recognized above, onto a two-dimensional house (Fig. 1b). Whereas lymphoid and erythroid populations from APL sufferers fashioned clusters that corresponded to cell varieties additionally annotated by regular hematopoietic cells, APL cells had been predominantly grouped into a definite cluster. The accuracy of APL blast identification was confirmed by the unique presence of PML/RARα expression, as noticed in two APL affected person samples utilizing PML/RARα-targeted scRNA-seq (Fig. 1b, proper panel, and Supplementary Fig. 2). This cluster exhibited a excessive expression stage of the gene MPO, which encodes a broadly used diagnostic marker for APL25, and in addition differed from the conventional spectrum of hematopoietic cell populations (comparable to HSPCs and myeloid cell populations; Fig. 1b, c). The established markers of APL cells had been extremely expressed on this cluster, as highlighted by genes activated by PML/RARα-associated super-enhancers8 (e.g., STAB1, CITED2, CCND2, and GFI1) (Fig. 1d). This discovering additional bolstered our earlier findings, highlighting the position of PML/RARα in figuring out the identification of APL blasts by way of super-enhancer regulation8. Furthermore, this cluster exhibited considerably elevated expression ranges of GMP-specific genes, comparable to azurophilic granule genes (MPO, AZU1, ELANE, and CTSG) (Fig. 1d), supporting the notion that APL blasts could also be blocked on the GMP stage14,23.
We proceeded to check the practical and regulatory traits of the APL blasts with these of the conventional GMP cluster. Firstly, we performed gene ontology (GO) enrichment evaluation on differentially expressed genes (DEGs) calculated utilizing Seurat26. The evaluation revealed that considerably upregulated genes (adjusted P-value < 0.05) in APL blasts had been of practical relevance to a number of key processes, together with HSC self-renewal/differentiation (RUNX1, MYC, and JAG1), histone modification (EP300) and DNA methylation (DNMT3A and MBD1), cell cycle arrest and cell development (CDK6, CCNA1, and WT1), in addition to the response to endoplasmic reticulum stress and unfolded protein (XBP1, ATF6, and USP14) (Fig. 1e and Supplementary Fig. 3a). In distinction, genes that had been considerably downregulated (adjusted P-value < 0.05) in APL blasts had been enriched for immune response-related capabilities, together with antigen processing and presentation (PSMB9, PSMB10, and CTSS), MHC class II protein advanced (CD74, HLA-DPA1, and HLA-DRA), regulation of cytokine manufacturing (CLEC7A, CCR2, and CCL3), and response to interferon-gamma (IRF8, IRF5, and IFI30) (Fig. 1f and Supplementary Fig. 3b). These findings confirmed the disruption of antigen presentation in APL cells27. Secondly, we used VIPER28 to deduce the transcription issue (TF) exercise differentially between the 2 clusters. Our evaluation steered that the hematopoietic TFs and cofactors (comparable to SPI1, ERG, FOS, and RXRA) had been repressed in APL blasts (Fig. 1g), supporting the differentiation blockade noticed in APL blasts. Additionally repressed in APL blasts had been the mediators of interferon (IFN) signaling (e.g., STAT1, IRF8, and IRF1). Conversely, oncogenic TFs (e.g., RB1, HIF1A, and MAX), epigenetic regulators (e.g., SMAD4, MBD1, YY1, and SP3), and cell proliferation-associated TFs (e.g., RNF6, ELF2, and TOP2B) had been activated in APL blasts in comparison with regular GMPs (Fig. 1g). In step with these findings, cell cycle evaluation confirmed that the APL cluster considerably amassed within the S or G2/M phases in contrast with regular myeloid cells, additional indicating the extremely proliferative state of APL blasts (Supplementary Fig. 4).
Characterizing the intratumoral heterogeneity of APL blasts reveals a posh cell-state transition trajectory with leukemic stem-like cells sitting on the prime
Subsequent, we decided the mobile structure and differentiation trajectory inside the characterised APL blasts. Via unsupervised clustering and UMAP evaluation, we recognized 18 clusters, every characterised by distinct expression patterns of recognized marker genes23 (Fig. 2a and Supplementary Information 2). Amongst these clusters, the 12 APL clusters (C1-C12) accounted for 82.9% of the cells and, as anticipated, exhibited excessive expression of GMP-specific genes23, notably these related to azurophilic granules, comparable to ELANE, CTSG, and AZU1 (Fig. 2b, proper panel). Notably, these GMP-like clusters displayed important heterogeneity. For instance, the three main clusters C1-C3 solely expressed GMP-specific genes, whereas clusters C6-C10 confirmed excessive expression of cell proliferation-related genes (TOP2A, MKI67, and PCNA)29, and clusters C11-C12 had been marked by the extremely expressed S100 household genes (S100A8/A9/A10).
a UMAP plot of APL blasts (higher panel; n = 126,802 cells). Eighteen clusters are labeled in several colours and numbers (decrease panel). GMP granulocyte-monocyte progenitors, EOS eosinophils, Prog progenitors, MDP monocyte-DC progenitors, PrecDC pre-conventional dendritic cells, Promono promonocytes. b The left heatmap reveals Spearman’s correlation between the 18 APL clusters, calculated utilizing the common expression profiles of the clusters. The appropriate heatmap illustrates the expression ranges of cell type-specific genes in every cluster. c Visualization of RNA velocity-based cell-state transitions of APL blasts. d UMAP plot of APL blasts with six branches, i.e., stem-like, Prog-like, S100helloGMP-like, GMP-like, biking GMP-like, and MDP-like branches. e Pseudotime-ordered evaluation of 4 main branches in APL blasts, together with stem-like, Prog-like, S100helloGMP-like, and GMP-like branches. f Heatmap displaying the dynamic adjustments in gene expression (n = 116 genes) alongside the pseudotime. Cell branches are labeled by colours (higher panel), together with stem-like cells (middle), S100helloGMP-like cells (left), and GMP-like (proper). Attribute transcription components (TFs) are listed on the correct. g UMAP plots of the focused scRNA-seq (scTarget) knowledge from two APL sufferers, with coloration coding for pattern varieties (left panel) and branches (proper panel). On the correct panel, cells detected greater than three PML/RARα fusion reads had been illustrated. h The expression ranges of PML/RARα in six branches of APL blasts. i Department-specific expression patterns for PML/RARα targets throughout the APL trajectory. The left heatmap visualizes the single-cell expression of PML/RARα-regulated branch-specific marker genes throughout branches, with rows representing genes and columns for cells. To supply a transparent and consultant depiction of the branch-specific expression patterns for PML/RARα targets, we chosen 1000 cells from every department for interpretation. The appropriate heatmap shows the imply gene expression (n = 1758 genes) throughout branches, accompanied by the annotations of consultant marker genes on the correct facet. Cor. correlation, Exp. expression.
Of placing curiosity, clusters C14-C16 exhibited stemness-like traits with excessive expression of marker genes particular to early HSPCs and/or LSCs (comparable to CD20030, CD4431, CD9932, CD233, and FAM30A34) (Fig. 2b), grasp stemness-related TF genes (comparable to SOX435 and MYC36), in addition to APL attribute genes (comparable to MPO) (Supplementary Fig. 5). Comparatively, C15 had the best expression ranges of stemness- and progenitor-specific genes in comparison with C14 and C16. As well as, C14 additionally expressed CD38, whereas C16 extremely expressed marker genes related to monocyte-DC progenitors (MDP)37, comparable to CSF1R/CD115 and FLT3/CD135. Collectively, the cells in C15 appeared to resemble probably the most primitive cells amongst APL blasts, presumably representing leukemic stem-like cells.
Subsequent, we carried out RNA velocity evaluation to confirm the differentiation trajectory from the leukemic stem-like cell cluster (C15) to the GMP-like cell clusters. The 18 APL clusters had been reorganized into six branches (Supplementary Information 3), with the stem-like cell cluster (C15) sitting on the root of differentiation trajectories, supported by each the velocity-based cell-state transition possibilities and the similarity of expression patterns (Fig. 2c, d). Possible going by way of the Prog-like department (C14), the stem-like cell cluster (C15) gave rise to the three fundamental branches of APL blasts: the GMP-like department (C1-C5 and C13), the biking GMP-like department (C6-C10), and the S100helloGMP-like department (C11 and C12) (Fig. 2c, d and Supplementary Fig. 6). One other trajectory ranging from C15 led to the MDP-like cell cluster (C16), which subsequently differentiated into cells within the CD1C+ PrecDC-like cell cluster (C17) and the CD14+ Promonocyte-like cell cluster (C18). This trajectory helps the present view that, as an alternative of being derived from GMPs37, MDPs may characterize an earlier stage, presumably even sooner than the CMP stage. Moreover, Monocle2 pseudotime ordering trajectory evaluation additionally confirmed that the stem-like cell cluster was the start line of APL blasts, giving rise to the S100helloGMP-like and GMP-like branches (Fig. 2e). This evaluation additionally revealed that RUNX1, RUNX2, and interferon-related components (i.e., STAT1 and FOS) is perhaps concerned within the GMP-like differentiation trajectory, whereas CEBP members of the family (CEBPA, CEBPB, and CEBPE), MAFB, JUNB, and JUND had been possible related to the lineage resolution in direction of the S100helloGMP-like APL blasts (Fig. 2f).
Moreover, we investigated the position of PML/RARα within the recognized APL trajectory. PML/RARα-targeted scRNA-seq confirmed that PML/RARα was uniformly expressed throughout all six branches of APL blasts, with notable expression within the stem-like cell cluster (Fig. 2g, h and Supplementary Fig. 7a). Moreover, we built-in PML/RARα chromatin occupancy knowledge31 obtained from CUT&Tag-seq (Cleavage Beneath Targets and Tagmentation sequencing) in an APL patient-derived cell line, NB4 (Supplementary Information 4 and Supplementary Fig. 7b). Notably, every department possessed a substantial variety of distinct PML/RARα targets (Fig. 2i and Supplementary Information 5), suggesting the presence of branch-specific expression patterns for PML/RARα targets throughout the APL trajectory. GO evaluation offered insights into their practical significance, revealing that these targets had been related to distinct practical pathways (Supplementary Fig. 7c). For example, the PML/RARα targets situated inside the APL stem-like cells had been discovered to be predominantly concerned in stem cell upkeep. These inside biking GMP-like cells had been primarily linked to E2F targets concerned within the G2-M checkpoint. These inside GMP-like cells exhibited marked enrichment in ribosomal capabilities. These outcomes collectively illuminate the impression of PML/RARα in shaping the intratumoral heterogeneity of APL cells.
The traits of APL stem-like cells had been decided by PML/RARα goal genes and additional enhanced by FLT3-ITD
To delve into the traits of the APL stem-like cells outlined in our examine, we recognized APL-specific leukemic stemness genes by evaluating the transcriptome knowledge between the APL stem-like cell cluster (C15) in Fig. 2a and the HSPC cluster in Fig. 1b (Supplementary Information 6). We obtained the next findings.
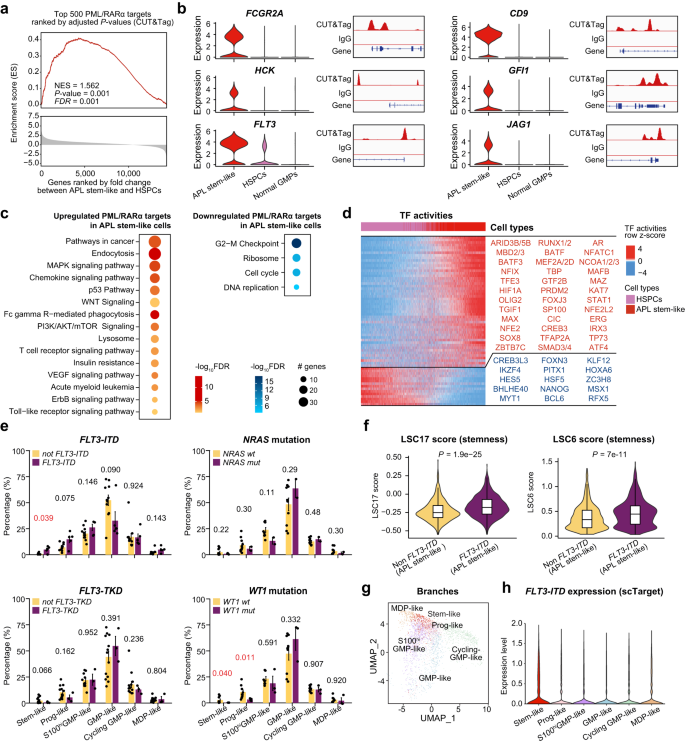
First, PML/RARα targets had been considerably enriched in these APL-specific leukemic stemness genes (Fig. 3a and Supplementary Fig. 8). Notably, well-known LSC marker genes16,38,39,40, comparable to FCGR2A, CD9, ITGA5, IL1RAP, and CD82, had been solely expressed in APL stem-like cells however not in HSPCs. They had been additionally direct targets of PML/RARα (Fig. 3b and Supplementary Fig. 9). Furthermore, well-known stemness/self-renewal-related PML/RARα targets8, comparable to HCK and GFI1, had been extremely expressed in APL stem-like cells. Genes carefully associated to APL leukemogenesis, comparable to FLT3 and JAG141,42, weren’t solely targets of PML/RARα but additionally exhibited elevated expression in APL stem-like cells (Fig. 3b).
a Gene set enrichment evaluation (GSEA) plot of prime 500 PML/RARα targets. The gene set for GSEA evaluation was outlined primarily based on the highest 500 PML/RARα targets in accordance with adjusted P-values derived from CUT&Tag. Genes had been ranked by the fold change between APL stem-like cells and HSPCs on the mRNA stage. NES normalized enrichment rating. The P-value was calculated utilizing GSEA. A two-sided P-value was calculated. b Violin plots illustrating consultant genes extremely expressed in APL stem-like cells in contrast with HSPCs. c GO enrichment evaluation displaying the KEGG pathways enriched in upregulated (left panel) and downregulated (proper panel) PML/RARα targets in APL stem-like cells in contrast with HSPCs. d Grasp regulator evaluation to discover activated (pink) and repressed (blue) transcription components (TFs) in APL stem-like cells in contrast with HSPCs. e Comparability of the odds of every department with and with out indicated mutations. n = 16 sufferers with FLT3-ITD/TKD mutation data and n = 12 sufferers with NRAS/WT1 mutation data. Error bars in bar plots characterize the means ± SE. The P-values had been calculated utilizing Pupil’s t-test and labeled in pink when P-values < 0.05. Two-sided P-values had been calculated. f Comparability of the LSC17 rating (left panel) and the LSC6 rating (proper panel) of the stem-like cells in APL sufferers with or with out FLT3-ITD. n = 2344 stem-like cells had been used for visualization, excluding these with a rating of 0 because of the absence of detected gene expression. Within the boxplot, a black line inside the field marks the median. The underside and prime of the field are situated on the 25th and 75th percentiles, respectively. The bars characterize values greater than 1.5 occasions the interquartile vary from the border of every field. The P-values had been calculated utilizing the Wilcoxon rank-sum take a look at. Two-sided P-values had been calculated. g Visualization of FLT3 expression by way of projection onto the UMAP of APL blasts utilizing the scTarget knowledge from two sufferers. Cells detected with greater than three FLT3-ITD mutated reads had been color-coded in accordance with the totally different branches. h The expression ranges of FLT3-ITD within the six branches of APL blasts, analyzed utilizing knowledge from FLT3-ITD-specific focused scRNA-seq (scTarget).
Second, we carried out pathway enrichment evaluation to disclose the involvement of the PML/RARα-dysregulated signaling pathways in APL stem-like cells. This evaluation recognized quite a few LSC-associated pathways, together with classical WNT, MAPK, VEGF, P53, and mTOR signalings43,44 (Fig. 3c), all essential for sustaining the LSC inhabitants. Moreover, pathway crosstalk evaluation primarily based on these LSC-associated PML/RARα goal genes indicated that they coordinated the regulation of APL stem-like cells (Supplementary Fig. 10 and Supplementary Information 7).
Third, we carried out VIPER evaluation to elucidate potential TFs concerned within the PML/RARα-induced transcriptional community in APL stem-like cells (Fig. 3d). By evaluating with regular HSPCs, we discovered that stemness-associated TFs, comparable to TGIF1 and HIF1A45,46 had been activated, suggesting their potential roles in regulating the stemness of APL cells. Moreover, we noticed the activation of regulators related to histone modification, together with MBD2/3 (methylated-DNA binding proteins) and PRDM2 (H3K9 methyltransferase), in APL stem-like cells. These findings steered their potential involvement in epigenetic management of self-renewal and quiescence of APL stem-like cells47. TFs linked to malignant transformation and stemness properties, comparable to OLIG2 and ARID3B48,49, had been additionally discovered to be energetic in APL stem-like cells. These findings highlighted the essential roles of PML/RARα in regulating APL stem-like cells on the single-cell stage.
Fourth, we proceeded to affiliate the mobile structure of APL blasts with the cooperating genetic alterations generally present in APL, together with FLT3 (FLT3-ITD and FLT3-TKD), WT1, and NRAS mutations. As illustrated in Fig. 3e, FLT3-ITD was considerably related to a extra primitive illness phenotype than different investigated mutations, suggesting that the presence of FLT3-ITD may play a big position in enhancing the leukemic stemness. We additional used two established leukemic stemness scorings34,50, LSC17 and LSC6, to check the stemness traits between APL stem-like cells with and with out FLT3-ITD. The evaluation revealed that the stem-like cells with FLT3-ITD obtained considerably larger scores than these with out FLT3-ITD (Fig. 3f), supporting the notion that FLT3-ITD may improve the stemness traits. Furthermore, we carried out FLT3-ITD-targeted scRNA-seq in two APL sufferers to confirm its existence and, extra importantly, to substantiate its larger expression in APL stem-like cells (Fig. 3g, h).
The predictive energy of the APL stemness rating in early loss of life and remedy consequence in APL
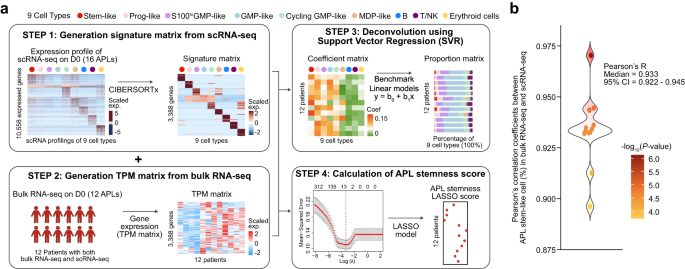
On this part, we explored how APL stem-like cells outlined by scRNA-seq might be utilized to foretell the medical obstacles in APL, extra exactly, the incidence of early loss of life, danger stratification, and remedy consequence. We first established deconvolution-based prediction procedures, as graphically illustrated in Fig. 4a and detailed within the “Strategies” part, and demonstrated their robustness and efficiency (Fig. 4b). At Step 1, we utilized the CIBERSORTx algorithm51 to scRNA-seq knowledge, producing an APL-specific signature matrix that concerned six APL blast cell populations (i.e., stem-like, Prog-like, GMP-like, biking GMP-like, S100helloGMP-like, and MDP-like) and three non-leukemic cell varieties (i.e., T/NK, B, and erythroid cells). At Step 2, we ready the transcripts per kilobase of the exon mannequin per million mapped reads (TPM) matrix from bulk RNA-seq knowledge. At Step 3, we employed assist vector regression (SVR)52 to deconvolute each the signature and TPM matrices, leading to a coefficient matrix. The proportion of every cell sort within the coefficient matrix was used to construct a linear regression mannequin for benchmarking. At Step 4, we constructed an 11-gene scoring mannequin by way of LASSO to judge the stemness of APL blast cells from bulk RNA-seq knowledge, the place the next rating signifies the next stemness. Moreover, we designed a leave-one-out take a look at to show the robustness of the inferred APL stem-like cell proportions (median R = 0.933, 95% CI = 0.922–0.945, Fig. 4b). In different phrases, our deconvolution method may precisely predict APL stem-like cells from bulk APL transcriptomes.
a Schematic depicting the APL deconvolution method and the technology of the APL stemness rating utilizing the 16 APL scRNA-seq knowledge because the reference. This improved deconvolution method relies on the assist vector regression (SVR) algorithm: (1) technology of signatures from scRNA-seq populations, together with the six APL blasts branches and T/NK cells, B cells, and erythroid cells; (2) calculating the TPM matrix of bulk RNA-seq of the 12 sufferers with matched scRNA-seq (detailed in “Strategies” part); (3) utilizing SVR to calculate the coefficients of every scRNA-seq inhabitants from bulk RNA-seq and performing linear regression to benchmark the share and the coefficient of every inhabitants; (4) technology of APL stemness signature genes, which had been PML/RARα targets extremely expressed in APL stem-like cells; and (5) the APL stemness rating was calculated by the imply expression stage of APL stemness signature genes. b Mannequin efficiency of the deconvolution method to foretell the APL stem-like cell proportion from bulk RNA-seq of the 12 sufferers. The efficiency of the mannequin is evaluated by Pearson’s correlation coefficients between the noticed APL stem-like cell proportion from scRNA-seq and the expected APL stem-like cell proportion from the deconvolution method. Depart-one-out (LOO) is used to judge the robustness of the mannequin. The P-values had been calculated utilizing the Pearson’s correlation. Two-sided P-values had been calculated.
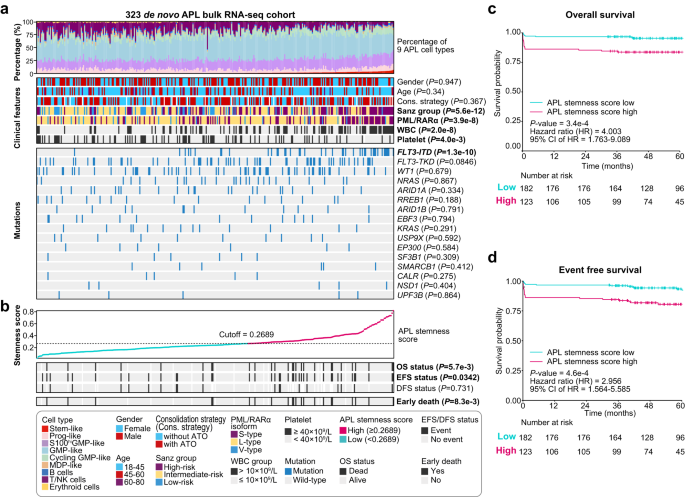
Subsequent, using our established deconvolution prediction procedures, we examined a big cohort comprising 323 APL sufferers53 to discover the correlation between APL stem-like cells and the medical traits of APL sufferers (Supplementary Information 8). Firstly, the next proportion of APL stem-like cells was considerably related to an elevated white blood cell (WBC) depend (P < 0.0001) and a decrease platelet depend (P = 4.0e-3) (Fig. 5a). Notably, the APL stem-like cell sort confirmed the strongest correlation with the WBC depend, adopted by GMP-like and biking GMP-like cell varieties (Supplementary Figs. 11, 12 and Supplementary Information 9). This discovering additionally emphasised the intertumoral heterogeneity amongst APL sufferers. Additional evaluation revealed {that a} larger proportion of stem-like cells was considerably related to the next proportion of APL blasts in BM cells (R = 0.53, P < 0.0001; Supplementary Fig. 13a) and the next blast depend in peripheral blood (R = 0.35, P < 0.0001; Supplementary Fig. 13b). This was additionally notably correlated with an elevated WBC (P < 0.0001; Supplementary Fig. 14). These outcomes indicated that APL sufferers with the next proportion of APL stem-like cells in APL blasts may need an elevated tendency for blasts to flow into in peripheral blood. Secondly, we examined the connection between the share of APL stem-like cells and recurrent mutations in APL sufferers, together with three widespread isoforms of PML/RARα (L-type, S-type, and V-type)54, FLT3 mutations (ITD and TKD), and mutations involving WT1, NRAS, and ARID1A. Remarkably, the next proportion of APL stem-like cells was considerably related to the S-type PML/RARα transcript (S-type vs. L-type: P < 0.0001; S-type vs. V-type: P = 7.3e-3; Fig. 5a and Supplementary Fig. 15) and FLT3-ITD (P < 0.0001). As well as, FLT3-ITD was recognized as probably the most important co-occurrence occasion, supporting the significance of FLT3-ITD in enhancing the stemness exercise of APL stem-like cells (Fig. 3f).
a Profiling of the expected APL cell sort proportion, medical options, and gene mutations within the 323 APL sufferers. Columns characterize particular person APL sufferers, and the P-values had been calculated utilizing the Wilcoxon rank-sum take a look at (statistical exams with two teams) or evaluation of variance (ANOVA, with greater than two teams) for example the correlation between the APL stem-like cell proportion and medical options/genetic alternations. Two-sided P-values had been calculated. b Examination of the APL stemness rating and its relationship to the prognosis of the 323 APL sufferers. The P-values had been calculated utilizing the Wilcoxon rank-sum take a look at to disclose the correlation between the APL stemness rating and the prognosis of the 323 APL sufferers. * PML/RARα isoforms primarily embody three typical varieties: lengthy (L), brief (S), and variant (V) varieties, respectively outlined by the breakpoint of PML on intron 6, intron 3, and exon 6. OS total survival, EFS event-free survival, DFS disease-free survival. c, d Kaplan–Meier estimates of total survival (OS) (c) and event-free survival (EFS) (d) of APL sufferers (n = 305 sufferers with prognostic data) in all the cohort stratified by the APL stemness rating. The P-values had been calculated utilizing the log-rank take a look at. Two-sided P-values had been calculated.
Given the shut affiliation of APL stem-like cells with potential unfavorable prognostic components (together with the excessive WBC depend, S-type PML/RARα, and FLT3-ITD), we sought to develop a stemness scoring system tailor-made for APL blast cells. Using the LASSO algorithm, we established an APL-specific stemness rating primarily based on the estimated cell proportions of APL stem-like cells, which was then utilized to quantify the stemness of leukemic cells in every affected person (detailed in “Strategies” part; Supplementary Fig. 16). We recognized eleven genes (SKAP2, IL1RAP, PLD1, HOPX, TRIM47, MAP2K1, TNFSF4, OLFML2A, P2RY14, NPTX2, and RALA) to assemble the APL stemness rating, which confirmed a big correlation with the proportion of APL stem-like cells (Pearson’s correlation = 0.802; P < 0.0001). We then explored the connection between the APL stemness rating and prognosis, together with total survival (OS), event-free survival (EFS), and disease-free survival (DFS). Remarkably, a excessive APL stemness rating was considerably related to a poorer OS (P = 5.7e-3) and EFS (P = 0.0342), however not DFS (P = 0.731; see Fig. 5b, with the cutoff optimized utilizing the R ‘maxstat’ algorithm). Univariate Cox evaluation additionally revealed that sufferers with the next APL stemness rating had a poor prognosis, as mirrored by OS (P = 3.4e-4; Fig. 5c) and EFS (P = 4.6e-4; Fig. 5d) utilizing the optimized cutoff. Multivariate Cox evaluation confirmed the APL stemness rating as an impartial prognostic issue for OS and EFS (Supplementary Fig. 17).
Notably, the next APL stemness rating was considerably related to an elevated danger of early loss of life (P = 8.3e-3; Fig. 5b). This discovering was per our scRNA-seq knowledge, revealing that in comparison with sufferers with out early loss of life, these sufferers with early loss of life had a comparatively larger proportion of stem-like cells, in addition to elevated expression of stemness-associated genes, comparable to FCGR2A, IL1RAP, MAP2K1, and KLF9 (Supplementary Fig. 18). Furthermore, additional evaluation confirmed that our APL stemness rating was an impartial danger issue (HR = 5.627; 95% CI, 1.981–15.980; P = 0.001) with a superior predictive worth for early loss of life (Supplementary Fig. 19). These outcomes collectively underscore the utility of the APL stemness rating in assessing APL danger, together with the chance of early loss of life in APL.
In vivo impact of ATRA on differentiation of primitive APL blasts and its affect on early loss of life danger
Administrating ATRA as early as doable has been confirmed important in decreasing the early loss of life charge in APL48,49. We then delved additional into exploring the in vivo impression of ATRA remedy on APL mobile hierarchies, with a specific deal with the primitive blasts, as their stemness may affect remedy response. We carried out scRNA-seq on BM samples collected from three sufferers (APL03, APL04, and APL05) after two days of ATRA remedy. At the moment level, a notable improve in CD11b expression was noticed (Supplementary Fig. 20a). Utilizing the pre-defined six branches of APL blasts at prognosis served because the reference, the cell kinds of APL cells on Day 2 after ATRA remedy had been decided by using the KNN algorithm in a merged dataset that included cells from each Day 0 and Day 2. We then investigated the in vivo results of ATRA by way of adjustments within the abundance of cell teams and differential transcriptional regulation. We additional utilized the deconvolution methodology on bulk transcriptomes from RNA-seq carried out on 10 APL sufferers (together with three who suffered from early loss of life) earlier than and after ATRA remedy to discover the potential affect of leukemic stemness on the differential in vivo response to ATRA, which is perhaps a contributing issue to early loss of life. The stemness scores had been certainly larger in sufferers who skilled early loss of life than those that achieved full remission in bulk RNA-seq knowledge of those 10 sufferers, per our findings from the big cohort (Supplementary Fig. 20b).
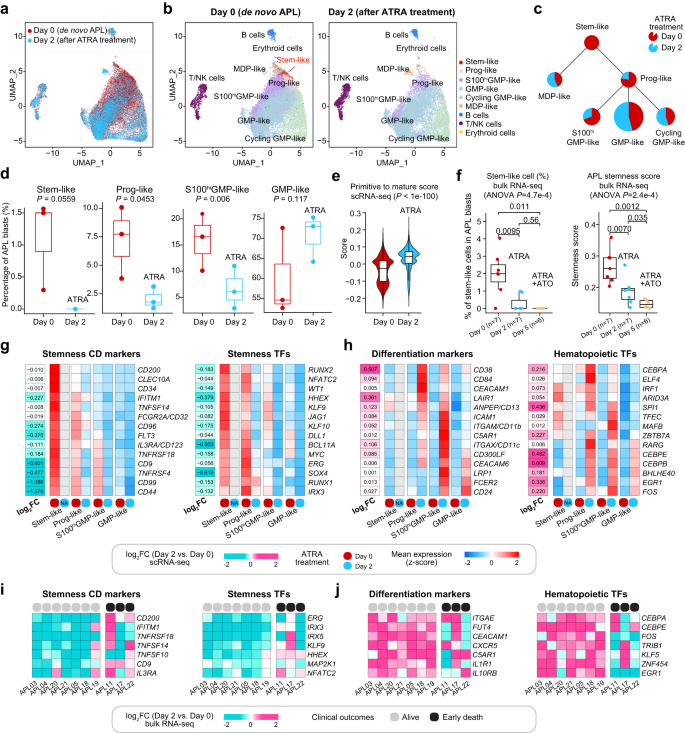
First, comparability evaluation of scRNA-seq knowledge revealed that diagnostic APL samples had been enriched in additional primitive cells than post-treatment samples, which had been comparatively enriched in additional mature progenitor cells (Fig. 6a). Particularly on Day 2 after remedy, the stem-like cells had been nearly undetectable within the post-treatment samples (Fig. 6b). Extra exactly, ATRA remedy considerably elevated the odds of the extra mature GMP-like cell sort and decreased the odds of three primitive cell varieties (stem-like, Prog-like, and S100helloGMP-like), with stem-like cells nearly undetectable after the remedy, suggesting that ATRA had a notable impression on primitive APL cells, particularly the stem-like cells (Fig. 6c, d). To substantiate the flexibility of ATRA to induce the differentiation of APL primitive blasts in direction of extra mature progenitor cells, we additionally employed the established rating to quantify the matureness of leukemic cells14, and noticed a big improve on this rating within the 2-day post-treatment samples in comparison with samples at prognosis (Fig. 6e), indicating that APL blasts turned extra mature upon ATRA remedy. Comparable findings had been additionally obtained by analyzing bulk transcriptomes of seven APL sufferers who achieved full remission earlier than and after ATRA remedy. As illustrated in Fig. 6f, the proportion of APL stem-like cells and the stemness of leukemic cells had been notably decreased following ATRA remedy, particularly on Day 2.
a UMAP plot displaying the mixing of APL BM cells (n = 39,415 cells) collected on Day 0 (pink) and Day 2 after ATRA remedy (blue). b UMAP plots displaying all outlined cell populations of APL BM cells (n = 39,415 cells) collected on Day 0 and Day 2 after ATRA remedy. c Comparability of the odds of six branches earlier than (pink) and after two days of ATRA remedy (blue). d Comparability of the odds of the stem-like, Prog-like, S100helloGMP-like, and GMP-like cells on Day 0 (pink, n = 3 samples) and Day 2 after ATRA remedy (blue, n = 3 samples). The P-values had been calculated utilizing Pupil’s t-test. A one-sided P-value was calculated. Within the boxplot, a black line inside the field marks the median. The underside and prime of the field are situated on the 25th and 75th percentiles, respectively. e Comparability of the Primitive-to-Mature scores of APL blasts on Day 0 (pink, n = 13,966 cells) and Day 2 after ATRA remedy (blue, n = 10,852 cells). f Comparability of APL stem-like cell percentages (left panel) and stemness scores (proper panel) on Day 0 (in pink, n = 7 samples), Day 2 (in blue, n = 7 samples handled with ATRA alone), and Day 5 (in orange, n = 5 samples handled with ATRA + ATO). Notably, inside the boxplot proven in (e and f), a black line marks the median, and the underside and prime of the field are situated on the 25th and 75th percentiles, respectively. The bars characterize values greater than 1.5 occasions the interquartile vary from the border of every field. The P-values had been calculated utilizing the Wilcoxon rank-sum take a look at. Two-sided P-values had been calculated. g Heatmap displaying the normalized expression of stemness CD markers and TFs in APL stem-like cells, Prog-like, S100helloGMP-like, and GMP-like clusters on Day 0 (pink) and Day 2 after ATRA remedy (blue). h Heatmap displaying the normalized expression of differentiation markers and hematopoietic TFs in APL stem-like cells, Prog-like, S100helloGMP-like, and GMP-like clusters on Day 0 (pink) and Day 2 (blue) after ATRA remedy. i Heatmap illustrating the log2(fold adjustments) (log2FC) of gene expression ranges for stemness-associated CD markers and TFs between APL samples on Day 0 and Day 2 (Day 2 vs. Day 0) in several APL sufferers. j Heatmap illustrating the log2FC of gene expression ranges for differentiation markers and hematopoietic TFs between APL samples on Day 0 and Day 2 (Day 2 vs. Day 0) in several APL sufferers. Grey represents sufferers alive after induction remedy, and black for sufferers with early loss of life.
Subsequent, we regarded on the expression adjustments of hematopoietic differentiation-related CD markers and TFs to indicate that ATRA may induce a stepwise differentiation, ranging from APL stem-like cells in direction of extra mature cells. Notably, we noticed a big downregulation of APL stemness CD markers (comparable to CD200, CD34, FCGR2A/CD32, CD9, and IL3RA/CD123) and TFs (comparable to HHEX, MYC, JAG1, and ERG) in primitive cell varieties (Fig. 6g and Supplementary Information 10). Conversely, markers and TFs related to mature hematopoietic cell lineages had been upregulated following ATRA remedy. For instance, CD38, CD84, CEBPA, and ELF4 had been upregulated within the Prog-like cells; ITGAM/CD11b, C5AR1, TFEC, and MAFB had been upregulated within the S100helloGMP-like cells; and CD24, CEACAM6, CEBPE, and CEBPB upregulated within the GMP-like cells (Fig. 6h and Supplementary Information 10).
Of explicit curiosity, we recognized distinct transcriptional responses induced by ATRA in APL sufferers with and with out early loss of life. As illustrated in Fig. 6i, the constant downregulation of a number of stemness-associated CD markers (CD200 and CD9) and TFs (HHEX and NFATC2) was not noticed in APL sufferers with early loss of life (Supplementary Information 11). Equally, the constant upregulation of differentiation-related CD markers (FUT4 and IL1R1) and TFs (FOS and EGR1) was additionally not noticed in sufferers with early loss of life (Fig. 6j and Supplementary Information 11). GO evaluation revealed related outcomes: along with the induction of differentiation, the repression of stemness-associated pathways, such because the MAPK cascade, was additionally not noticed in sufferers with early loss of life after ATRA remedy (Supplementary Fig. 20c, d). Moreover, a big lower of the APL stemness scores after ATRA remedy was noticed in sufferers who achieved full remission, however not in sufferers who skilled early loss of life (Supplementary Fig. 20b). Our findings recommend that ATRA remedy had a lesser impression on the stemness program in APL sufferers with early loss of life, doubtlessly explaining the numerous affiliation we noticed between the stemness exercise of leukemic cells and early loss of life, as revealed by our deconvolution evaluation of transcriptomes from 323 APL sufferers (Fig. 5b).