Differential expression of KPNA2
We first examined the intersection of 1650 GC-related genes in GeneCards with 1455 RBPs, and we recognized 827 GC-related RBPs (Fig. 2A). Subsequently, 19 differentially expressed RBPs had been recognized from 843 DEGs (Fig. 2B). Varied RBPs, resembling KPNA2, KRT18, PABPC1, FSCN1, CDKN2A and TOP2A, had been discovered to be extremely expressed in GC (Fig. 2C). Utilizing GC and the aforementioned RBPs as key phrases, a PubMed literature search (https://pubmed.ncbi.nlm.nih.gov/) was carried out and revealed that the operate of KPNA2 in GC has not been properly studied. On this research, based on the GEPIA database, KPNA2 expression was elevated in cancers of the digestive system (Fig. 2D). The mRNA expression of KPNA2 was considerably upregulated in GC sufferers based on the TCGA database (Fig. 2E,F). Moreover, the downregulation of KPNA2 was noticed by immunohistochemistry (IHC) of samples that had been acquired from the HPA database (Fig. 2G).
Differential expression of KPNA2. (A,B) Venn diagram. (C) Expression of a number of RBPs in GC. (D) Comparability of KPNA2 expression in digestive system most cancers samples and regular tissue samples from GEPIA. (E,F) KPNA2 was overexpressed in GC in unpaired and paired TCGA samples. (G) Elevated protein expression of KPNA2 in GC from the HPA. *p < 0.05, **p < 0.01, ***p < 0.001.
KPNA2 regulated gene expression in AGS cells
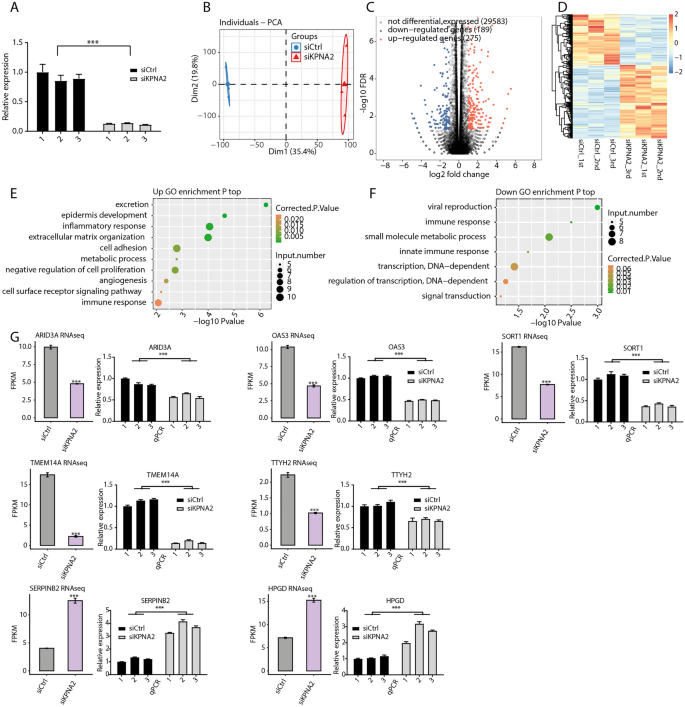
We used RNA-seq to discover the features of latent KPNA2 in AGS cells. We first knocked down KPNA2 in AGS cells by way of siRNA transfection (Fig. 3A). Principal part evaluation (PCA) revealed that the FPKMs of all of the detected genes had been clustered throughout the group, with good intragroup repeatability and related pattern information. There was good differentiation between the siCtrl and siKPNA2 teams (Fig. 3B). Curiously, silencing of KPNA2 resulted within the upregulation of 275 genes and the downregulation of 189 genes (FC ≥ 2 or ≤ 0.5, FDR < 0.5) (Fig. 3C). Particulars about these DEGs might be present in Supplementary Desk 2. Heatmap evaluation of DEG expression profiles based mostly on RNA-seq information revealed a excessive diploma of consistency in KPNA2-regulated transcription among the many three units (Fig. 3D). Subsequent, we recognized a number of pathways that included upregulated genes annotated with classes together with epidermal improvement, inflammatory response, cell adhesion, damaging regulation of cell proliferation, and immune response (Fig. 3E). Moreover, GO evaluation revealed a number of organic processes, together with immune response, small molecule metabolic course of, innate immune response, and transcription, which had been considerably enriched for the downregulated genes (Fig. 3F). In accordance with the gene expression ranges, the considerably upregulated DEGs within the siKPNA2_1st group had been organized in descending order, as had been the considerably downregulated DEGs within the siCtrl_1st group. The highest 30 up- and downregulated DEGs are proven in Supplementary Tables 3 and 4, respectively. We discovered that ARID3A, OAS3, SORT1, TMEM14A, TTYH2, HPGD, and SERPINB2 had been related to cancers35,36,37,38,39,40,41. The hypoexpression of KPNA2 induced downregulation of ARID3A, OAS3, SORT1, TMEM14A, and TTYH2 and the upregulation of HPGD and SERPINB2, and these findings had been additional verified by RT‒qPCR (Fig. 3G).
KPNA2 regulated gene expression in AGS cells. (A) KPNA2 expression was quantified by RT‒qPCR. Error bars characterize the imply ± SEM. (B) PCA of two teams of samples based mostly on the normalized gene expression stage. (C) Identification of KPNA2-regulated genes. (D) Hierarchical clustering of DEGs in management and KPNA2-knockdown samples. (E,F) The highest ten consultant GO organic processes of up- or downregulated genes. (G) RNA-seq and RT‒qPCR demonstrated that some differentially expressed genes had been regulated by KPNA2. The error bars point out the means ± SEMs. ***p < 0.001.
KPNA2 regulated gene AS in AGS cells
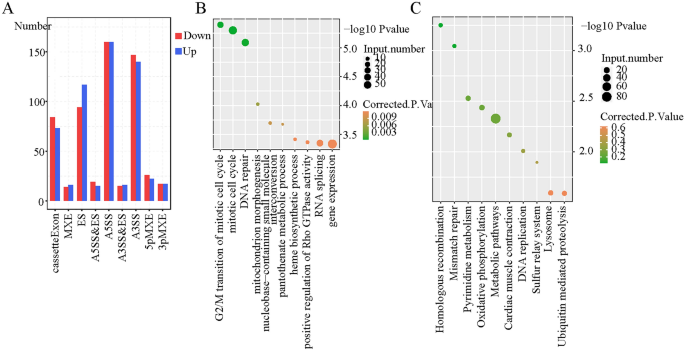
It’s unclear whether or not KPNA2, which is an RBP, impacts gene AS in GC. Due to this fact, we used RNA-seq to find out whether or not KPNA2 regulates AS (RASE) in AGS cells. A complete of 1560 ASEs had been altered after KPNA2 was silenced (Supplementary Desk 5). The principle RASEs had been the choice 5′ splice web site (A5SS), various 3′ splice web site (A3SS), exon skipping (ES), and cassette exon, along with mutually unique 3′UTRs (3pMXE), mutually unique 5′UTRs (5pMXE), A5SS and ES, and mutually unique exons (MXE) (Fig. 4A). Taken collectively, these outcomes indicated that the A5SS, A3SS, ES, and cassette exons had been the primary ASEs that had been regulated by KPNA2 and that KPNA2 had a complete regulatory impact on AGS cells. A complete of 1241 RASGs remained after KPNA2 silencing. GO evaluation was carried out on these RASGs, which had been enriched in organic processes such because the G2/M transition of the mitotic cell cycle, mitotic cell cycle, and DNA restore (Fig. 4B). In accordance with KEGG enrichment evaluation, these RASGs had been enriched within the pathways of homologous recombination, mismatch restore, and pyrimidine metabolism (Fig. 4C).
KPNA2 sure to mRNAs related to GC in AGS cells
The RBPs are assessed based mostly on their capability to bind to RNA and the quantity of those proteins that may be successfully recovered. To achieve perception into the specificity of KPNA2 pull-down experiments, we carried out iRIP-seq to determine the mRNAs that sure to KPNA2. Two replicate experiments had been performed to make sure the reliability of the outcomes. iRIP-seq confirmed good high quality management and reproducibility within the IP group (Desk 1). In abstract, these outcomes demonstrated that KPNA2 can particularly bind to mRNA.
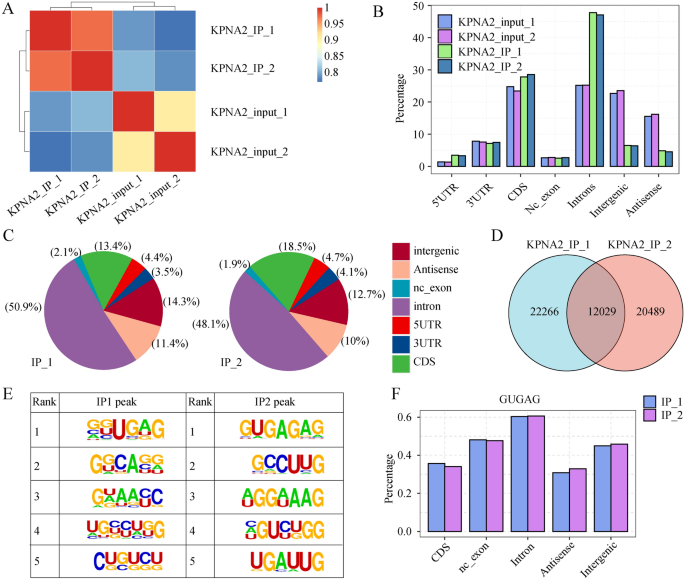
As proven in Fig. 5A and Fig. S1A, the correlations of the samples had been 0.711 and 0.706, respectively, indicating that the reads had been considerably enriched within the IP group relative to the enter group. The genomic distribution of the reads confirmed that the KPNA2-IP reads had been enriched within the 5′UTR, CDS, and intron areas in comparison with these within the enter group (Fig. 5B) and most importantly enriched within the introns (Fig. 5C); this outcome instructed that KPNA2 could bind to those areas to manage transcript stability, AS, and translation processes. A complete of 12,029 KPNA2-binding areas had been recognized within the two replicate trials utilizing the Ablife methodology for peak calling of the binding areas (Fig. 5D). Notably, the binding motifs (GUGAG) of KPNA2 that had been recognized within the two replicate trials had been constant (Fig. 5E). The GUGAG motif was additionally primarily distributed within the intron area (Fig. 5F). The genes related to the KPNA2 binding peak within the KPNA2-IP group are proven in Supplementary Desk 6 and Supplementary Desk 7. GO evaluation revealed that genes that had been related to KPNA2 binding peaks that had been noticed in each trials had been enriched for protein phosphorylation, gene expression, mitotic cell cycle, and different organic processes (Fig. S1B), which supplies a foundation for future research on the built-in regulation of KPNA2-RNA interactions in AGS cells.
KPNA2 sure to mRNAs related to GC in AGS cells. (A) Heatmap exhibiting the correlation between the IP and enter teams. (B) Bar plot exhibiting the learn distribution throughout the reference genome. (C) Pie charts of KPNA2-bound peaks throughout the reference genome. (D) Venn diagram exhibiting the overlap of KPNA2 binding peaks recognized within the two trials. (E) Motif enrichment of KPNA2-bound peaks by HOMER. (F) Bar plot exhibiting the distribution of peaks containing the GUGAG motif.
KPNA2 regulated pre-RNA AS by binding to pre-mRNAs
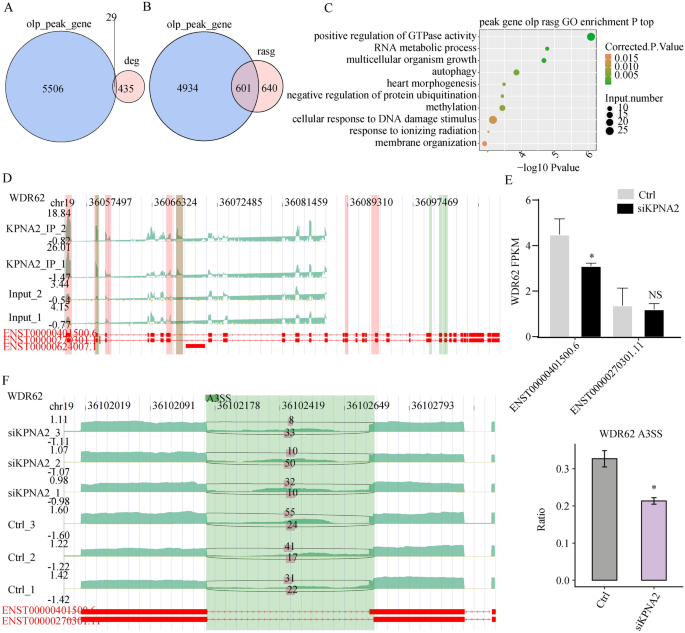
Twenty-nine genes had been discovered to have mRNAs or antisense RNAs with KPNA2 binding websites (Fig. 6A). As a result of KPNA2, which is a splicing issue, binds to RNA and impacts AS, our purpose was to research the impact of KPNA2 binding to pre-mRNAs on AS. Overlay evaluation was carried out utilizing the 1241 genes that corresponded to 1560 considerably totally different AS occasions that had been recognized within the RNA-seq information and the 5535 genes the place the binding peaks appeared in each trials of iRIP-seq. The outcomes revealed that KPNA2 could bind 601 pre-mRNAs to manage AS (Fig. 6B), and detailed info on the 601 genes is supplied in Supplementary Desk 8. A complete of 601 genes had been enriched for organic processes, resembling constructive regulation of GTPase exercise, RNA metabolic processes, methylation, and autophagy (Fig. 6C).
Built-in evaluation of KPNA2-bound genes and RASE in response to KPNA2 knockdown. (A) Venn diagram exhibiting the overlap of KPNA2-bound peaks and KPNA2-regulated DEGs. (B) The overlap of KPNA2-binding genes with KPNA2-regulated RASGs. (C) The highest ten consultant GO organic processes of the overlapping genes. (D) KPNA2 binds to WDR62 mRNA. (E) Modifications within the expression ranges of the WDR62 transcript. (F) KPNA2 regulated the AS of WDR62. *p < 0.05, **p < 0.01, ***p < 0.001.
We analyzed ASEs that considerably modified based on their ratios between the experimental and management teams. The iRIP-seq outcomes confirmed that KPNA2 may bind to WDR62 pre-mRNA (binding motif: GUGAG) (Fig. 6D), and the RNA-seq outcomes confirmed that after KPNA2 was downregulated, the expression of WDR62 (ENST00000401500.6) decreased (p = 0.027), whereas the expression of the ENST00000270301.11 transcript didn’t considerably change (p = 0.726) (Fig. 6E). Moreover, the A3SS AS mode of the WDR62 pre-RNA was decreased (p < 0.05, Fig. 6F).
Correlation evaluation of KPNA2 expression and immune infiltration
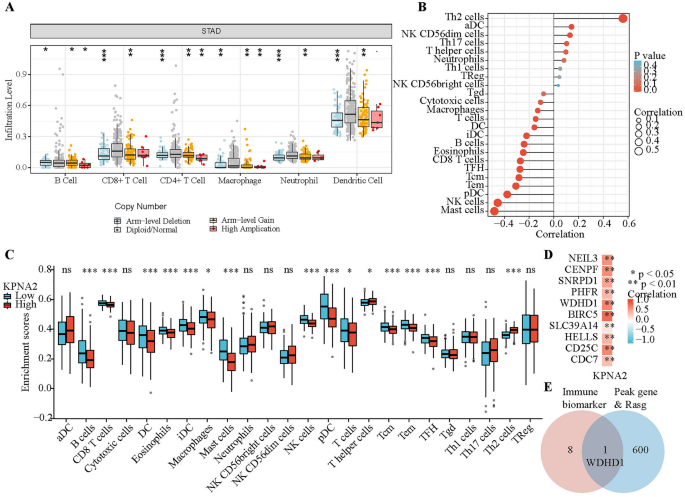
Enrichment evaluation revealed that the DEGs which can be regulated by KPNA2 had been intently associated to immunity. As immune cells within the tumor microenvironment (TME) affect affected person prognosis, exploring the correlation between KPNA2 expression and immune infiltration is a helpful space of analysis. First, KPNA2 gene copy numbers gave the impression to be associated to the infiltration of a number of immune cell varieties, together with neutrophils, B cells, CD4 + T cells, CD8 + T cells, macrophages, and dendritic cells (DCs), in GC based on the “SNCA” plate of TIMER (Fig. 7A). Moreover, we analyzed the correlation between the extent of infiltration of every immune cell subset and expression of KPNA2 in sufferers with GC from the TCGA database (Fig. 7B and Desk 2). To find out whether or not KPNA2 expression correlated with the extent of immune infiltration in GCs, we decided the immune cell enrichment scores of high- and low-KPNA2 expression teams (Fig. 7C). The intersection of those two analyses revealed that top KPNA2 expression was correlated with elevated infiltration of Th2 cells and decreased infiltration of B cells, CD8 + T cells, DCs, eosinophils, immature DCs, macrophages, mast cells, plasmacytoid DCs, T cells, T central reminiscence cells, T effector reminiscence cells, and T follicular helper cells in GC. The correlation coefficient of the Th2 cell infiltration stage with KPNA2 expression was the very best amongst all of the immune cells. Thus, we additional explored the affiliation between KPNA2 and markers of Th2 cells34. Curiously, KPNA2 expression was distinctively and positively correlated with the expression of the Th2 cell markers NEIL3, CENPF, SNRPD1, PHFR, WDHD1, BIRC5, SLC39A14, HELLS, CD25C and CDC7 in GC (Fig. 7D). We then screened WDHD1 from among the many KPNA2-regulated genes and located that KPNA2 can regulate WDHD1 to mediate the A3SS type of AS (Fig. 7E and Supplementary Desk 9).
KPNA2 was related to immune infiltration in GC. (A) Relationship between KPNA2 gene copy quantity and the extent of immune cell infiltration within the GC cohort. (B) Correlation evaluation of KPNA2 expression and immune infiltration in GC. (C) Enrichment scores of immune cells within the excessive and low KPNA2 expression teams. (D) Affiliation between KPNA2 and gene markers of Th2 cells. (E) Venn diagram. *p < 0.05, **p < 0.01, ***p < 0.001.
KPNA2 could regulate the AS of IRGs
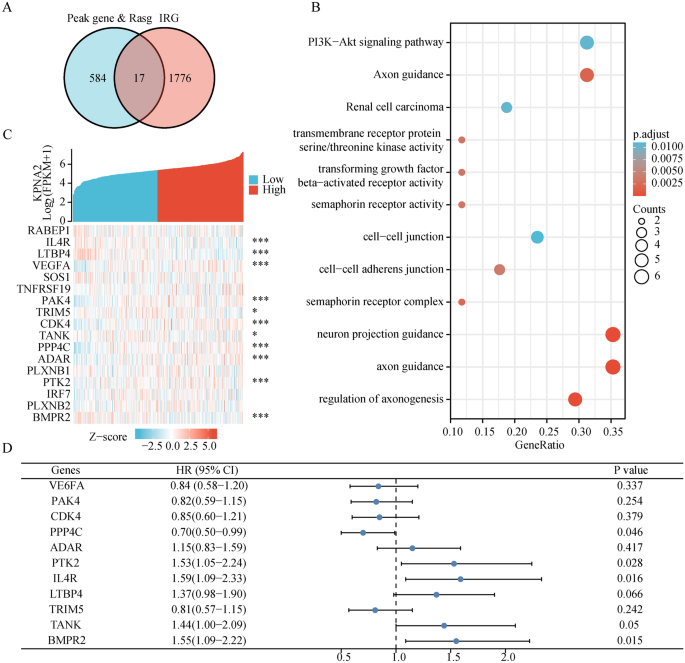
For the reason that first immune checkpoint inhibitor was permitted by the US FDA in 2011, now we have witnessed a rise in advances within the discipline of most cancers immunotherapy42. Therefore, we aimed to additional discover the relevance of AS to immunity. We extracted 17 KPNA2-binding IRGs from the height gene and Rasg (IPRG) (Fig. 8A). KPNA2 may regulate these 17 genes to endure varied types of AS (RABEP1: ES; PAK4: A5SS; TRIM5: A3SS, and so on.), and the main points had been introduced within the Supplementary Desk 10. Subsequent GO-KEGG enrichment revealed enrichment of the PI3K-Akt signaling pathway and transmembrane receptor protein serine/threonine kinase exercise (Fig. 8B). We examined the sample of KPNA2 co-expression with these 17 genes. The pink dots point out genes with a constructive affiliation with KPNA2, whereas the inexperienced dots point out genes with a damaging affiliation with KPNA2 (Fig. 8C). Furthermore, we discovered that GC sufferers with excessive expression of PPP4C (hazard ratio [HR] 0.70, 95% confidence interval [CI] 0.50–0.99, p = 0.046) had superior general survival (OS), whereas GC sufferers with excessive expression of IL4R (HR 1.59, 95% CI 1.09–2.33, p = 0.016) and BMPR2 (HR 1.55, 95% CI 1.09–2.22, p = 0.015) had inferior OS (Fig. 8D). Consequently, we speculated that KPNA2 may regulate AS, which in flip impacts the prognosis of sufferers with GC.
Experimental verification
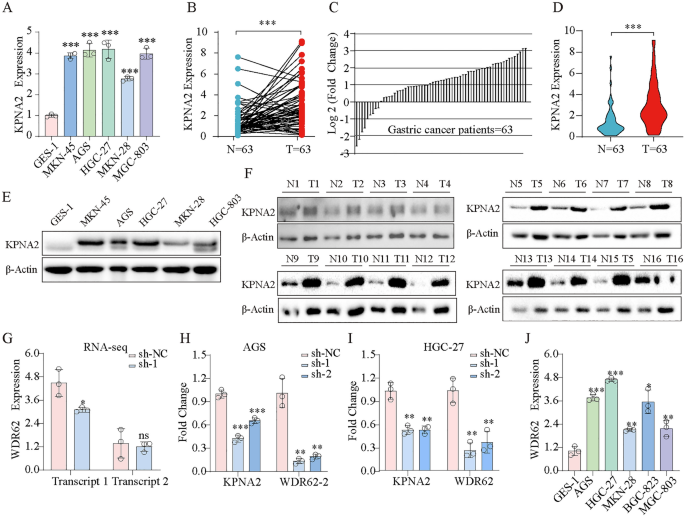
Initially, we explored KPNA2 expression in GES-1, MKN-45, AGS, HGC-27, MKN-28, and MGC-803 cells and revealed that KPNA2 was dramatically upregulated within the GC cell traces (Fig. 9A). We then in contrast the expression of KPNA2 in GC tissues with that in corresponding adjoining GC tissues and evaluated the scientific significance of KPNA2 expression in 63 sufferers with GC. Forty-five sufferers with GC exhibited elevated KPNA2 expression in most cancers tissues (p < 0.0001, Fig. 9B), and 40 (63.5%) sufferers with GC exhibited KPNA2 mRNA ranges that had been at the least twofold larger than these within the corresponding adjoining tissues (Fig. 9C). These outcomes revealed that KPNA2 expression was considerably larger in GC tissues than in corresponding adjoining tissue (p < 0.001, Fig. 9D). And the protein stage of KPNA2 was extremely expressed in gastric most cancers cells and gastric most cancers tissues (Fig. 9E,F). Moreover, we carried out a Chi-fang evaluation, which revealed that KPNA2 mRNA ranges had been correlated with lymph node metastasis (p = 0.001) and the Lauren classification (p = 0.000) however not with age, intercourse, tumor web site, or pathological stage of sufferers with GC (p > 0.05, Desk 3). RNA-seq revealed that the downregulation of KPNA2 decreased WDR62 transcript 1 (ENST00000401500.7) expression (Fig. 9G), and we verified this outcome by qRT‒PCR in AGS and HGC-27 cells (Fig. 9H,I). Furthermore, WDR62 transcript 1 was extremely expressed in GC cells (Fig. 9J).
(A) Expression of KPNA2 mRNA within the regular GC cell line (GSE-1) and GC cell traces (MKN-45, AGS, HGC-27, MKN-28 and MGC-803). (B) Comparability of KPNA2 expression ranges between 63 GC tissues and corresponding adjoining regular tissues. (C) Relative expression of KPNA2 within the 63 GC tissues and corresponding adjoining tissues. (D) Expression of KPNA2 in 63 pairs of GC tissues and corresponding adjoining tissues. (E) Expression of KPNA2 protein within the regular GC cell line (GSE-1) and GC cell traces (MKN-45, AGS, HGC-27, MKN-28 and MGC-803). (F) KPNA2 protein expression ranges between GC tissues and corresponding adjoining regular tissues. (G-I) Downregulation of KPNA2 decreased WDR62 transcript 1 expression. (G) RNA-seq. (H) RT‒qPCR in AGS cell. (I) RT‒qPCR in HGC-27 cell. (J) Expression of WDR62 transcript 1 within the regular GC cell line (GSE-1) and GC cell traces (MKN-45, AGS, HGC-27, GCB-823, and MGC-803). *p < 0.05, **p < 0.01, ***p < 0.001.
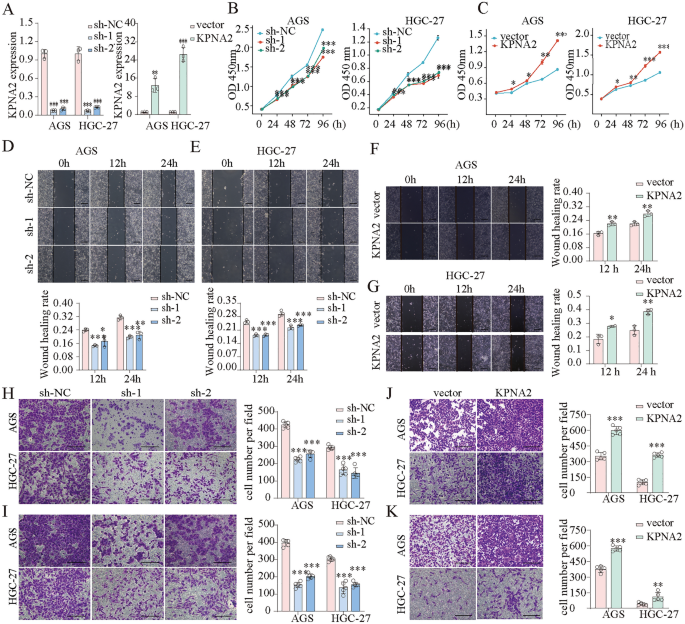
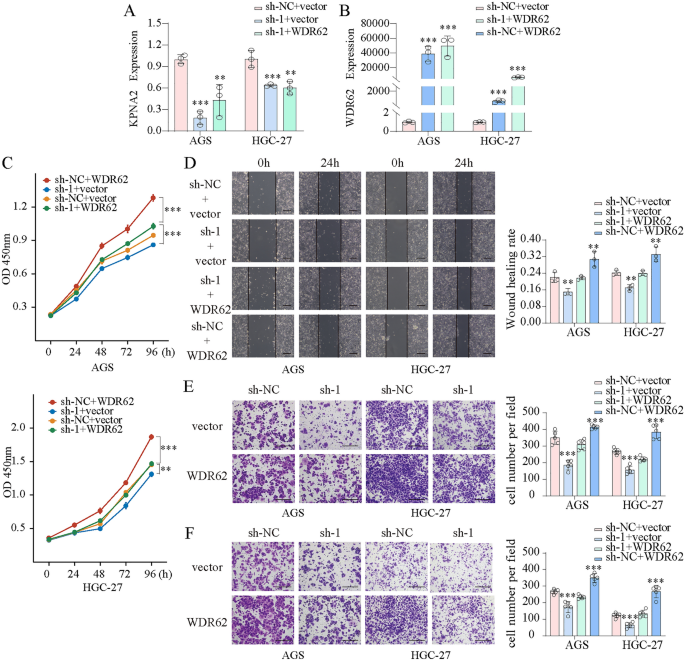
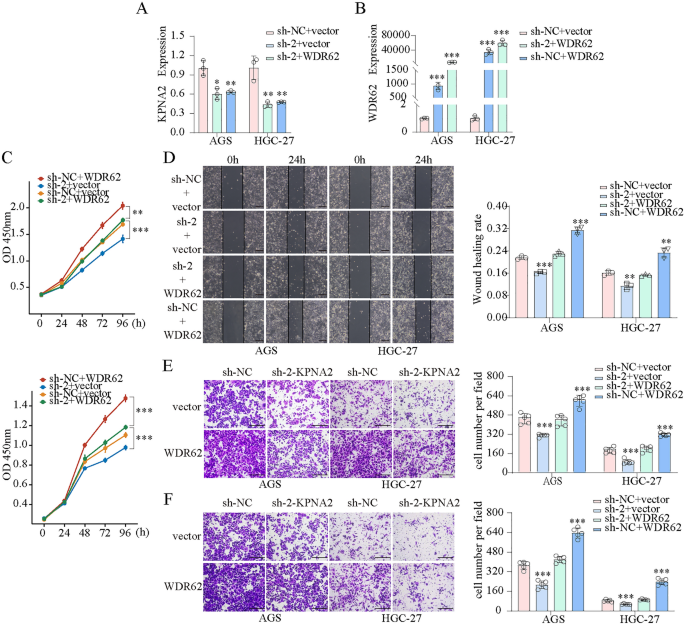
We transfected AGS and HGC-27 cells with KPNA2-shRNA and WDR62 plasmids and confirmed the transfection effectivity utilizing qPCR (Fig. 10A). CCK8 assays confirmed that KPNA2 downregulation considerably inhibited the proliferation, whereas KPNA2 overexpression enhanced proliferation in AGS and HGC-27 cells (Fig. 10B,C). Wound therapeutic and Transwell assays indicated that migration and invasion had been considerably suppressed by KPNA2 downregulation or enhanced by KPNA2 overexpression in AGS and HGC-27 cells (Fig. 10D–Okay). Furthermore, we concurrently down-regulated KPNA2 (sh-1 or sh-2) and up-regulated WDR62 in gastric most cancers cells (Figs. 11A,B, 12A,B), and located that overexpression of WDR62 reversed the KPNA2 knockdown-induced inhibition of proliferation (Figs. 11C, 12C), migration (Figs. 11D,E, 12D,E) and invasion (Figs. 11F, 12F). These outcomes demonstrated that KPNA2 could play a pro-oncogenic function by regulating WDR62.
KPNA2 promoted the proliferation, migration and invasion of AGS and HGC-27 cells. (A) KPNA2 was downregulated or upregulated. (B,C) Cell proliferation was measured utilizing CCK8 assay. (D–G) Cell migration was measured utilizing a wound therapeutic assay. (H–Okay) Cell migration and invasion had been measured utilizing Transwell assays. (H,J) Migration. (I,Okay) Invasion. *p < 0.05, **p < 0.01, ***p < 0.001.
The oncogenic roles of KPNA2 in GC partially depend on WDR62 expression. (A,B) Downregulation of KPNA2 (sh-1) and upregulation of WDR62 in AGS and HGC-27 cells. (C) The potential of WDR62 to reverse the KPNA2-induced adjustments in proliferation (CCK8 assay). (D–F) The potential of WDR62 to reverse the KPNA2-induced adjustments in migration and invasion. (D) Wound therapeutic assay, (E) Transwell migration assay, (F) Transwell invasion assay.
The oncogenic roles of KPNA2 in GC partially depend on WDR62 expression. (A,B) Downregulation of KPNA2 (sh-2) and upregulation of WDR62 in AGS and HGC-27 cells. (C) The potential of WDR62 to reverse the KPNA2-induced adjustments in proliferation (CCK8 assay). (D–F) The potential of WDR62 to reverse the KPNA2-induced adjustments in migration and invasion. (D) Wound therapeutic assay, (E) Transwell migration assay, (F) Transwell invasion assay.