Expression ranges of HMGNs in LUAD
To research the deregulation of HMGNs in lung adenocarcinoma (LUAD), the RNA-sequencing knowledge of 5 HMGN members in LUAD and regular lung tissues have been extracted from the TCGA database and two GSE knowledge units and analyzed. Volcano plots evaluation revealed an upregulation of HMGN1 (Fig. 1A). The up-regulation of HMGN1 was confirmed within the GSE18842 and different knowledge set (Fig. 1B, Supplementary Fig. 1A and B), whereas HMGN3 and HMGN5 didn’t exhibit the same development. Evaluation of gene expression of LUAD and matched adjoining non-tumor tissues from three cohorts confirmed constant up-regulation of HMGN1 in LUAD throughout all cohorts, with HMGN5 down-regulated in two of them (Fig. 1C–E). Lastly, pan-cancer evaluation throughout numerous cancers, together with breast, colon, liver, and abdomen cancers, additionally indicated excessive expression of HMGN1 (Supplementary Fig. 1C).
Aberrant expression of HMGN household in lung adenocarcinoma. (A) Volcano plots illustrating differential HMGN expression in lung adenocarcinoma versus regular tissues from the TCGA database (A) and GSE18842 (B). Expression values have been introduced as log2 (FPKM + 1). Fold modifications have been introduced as log2 (fold change). (C–E) Transcriptional expression of HMGNs in 57 paired lung adenocarcinoma and adjoining regular tissues from TCGA (C); 60 pairs from GSE19804 (D) and 33 pairs from GSE10072 (E). *p < 0.05, **p < 0.01, ***p < 0.001. ns: not statistically important. (F) Protein expression ranges of HMGNs in lung most cancers versus regular lung tissues from the CPTAC dataset.
Protein expression of HMGNs in LUAD was then assessed utilizing the Scientific Proteomic Tumor Evaluation Consortium (CPTAC) mass-spectrometry-based proteomics dataset. The outcomes have been consistent with mRNA expression knowledge, displaying elevated protein ranges of HMGN1 and decreased ranges of HMGN4/5 in LUAD (Fig. 1F). Moreover, the Human Protein Atlas was used to corroborate these findings on the protein degree (Supplementary Fig. 1B). In brief, these findings point out that irregular expression of HMGN members of the family is related to sufferers with LUAD.
Genetic and epigenetic alterations of HMGNs in LUAD
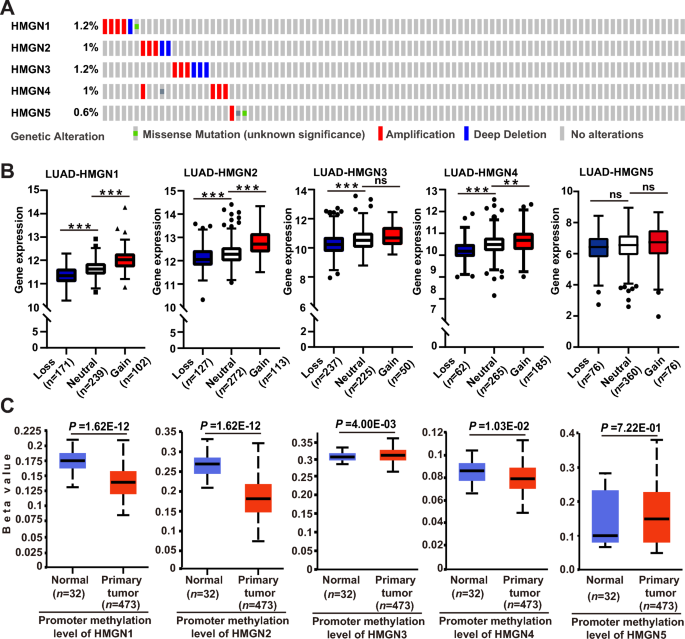
The aberrant expression of HMGNs could be because of genetic alterations that happen in coding areas and dysregulation of epigenetics on the promoter area. Subsequently, we characterised the mutation patterns of HMGNs in LUAD utilizing the cBioPortal on-line instrument. On the whole, the mutation charges of HMGNs are comparatively low. HMGN genes have been altered in 25 of 507 specimens (5%). Two alterations have been concurrently detected in virtually 1/20 of the samples (Fig. 2A). Since genomic structural variations are probably the most genetic alterations in LUAD, the MEXPRESS on-line instruments have been utilized to evaluate the transcriptional penalties of HMGN copy quantity variation (CNV). As proven in Fig. 2B, a considerably higher proportion of copy quantity acquire and losses of HMGN1/2/3/4/5 in LUAD was noticed. Growing proof has proven that DNA methylation degree at promoter area is a vital issue figuring out transcription. Thereby the promoter methylation of HMGNs in LUAD was analyzed utilizing the UALCAN portal. As proven in Fig. 2C, diminished methylation ranges of HMGN1/2/4 have been present in major lung tumors. And promoter methylation of HMGN3/5 have been up-regulated in tumor. These outcomes implied that HMGNs gene have been mutated sometimes, and the deregulation of HMGNs could be not less than partly because of copy quantity variation and irregular promoter methylation.
Genetic and epigenetic alterations of HMGN household in lung adenocarcinoma. (A) OncoPrint from the cBioPortal database displaying the distribution and proportion of samples with genetic alterations in HMGN genes in LUAD. (B) Correlation between copy quantity variation and gene expression variation for HMGNs in LUAD. (C) The contribution of promoter methylation degree of HMGNs to the expression of corresponding genes in LUAD. *p < 0.05, **p < 0.01,***p < 0.001, ns: not statistically important.
Potential of HMGN1 as a biomarker and prognosis in LUAD
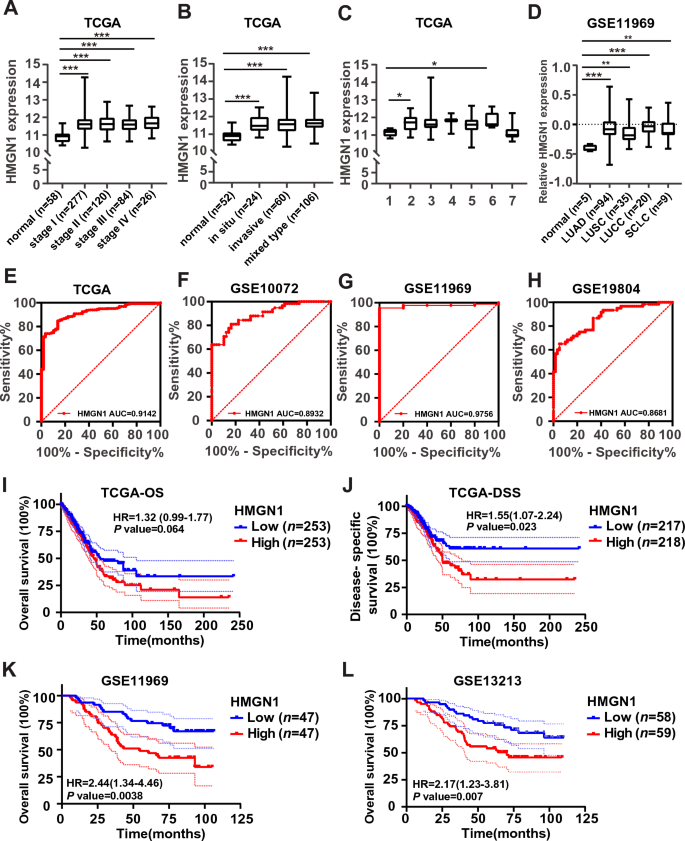
We chosen HMGN1 for subsequent evaluation of prognostic values and performance enrichment evaluation in LUAD since HMGN1 shows most distinguished modifications in expression, copy quantity variation promoter methylation. First, the connection between HMGN1 mRNA ranges and clinicopathological subgroups of LUAD sufferers was evaluated within the TCGA lung most cancers and GSE11969 cohorts. As proven in Fig. 3A, the expression of HMGN1 in all tumor levels (I–IV) was considerably larger than that in regular group. As well as, the expression of HMGN1 in several subtypes of LUAD was considerably larger than that in regular group (Fig. 3B and C). Moreover, HMGN1 expression in 4 main subtypes of lung most cancers was considerably larger than regular management (Fig. 3D). Nonetheless, no distinction was seen between completely different tumor levels/subtypes (Fig. 3B–D).
Scientific significance of HMGN1 expression in lung adenocarcinoma. (A) Boxplot depicting HMGN1 expression affiliation with tumor levels within the TCGA dataset. (B) Boxplot illustrating the affiliation between HMGN1 expression and tumor clinicopathological classification within the TCGA dataset. (C) Boxplot displaying HMGN1 expression in several invasive tumor subtypes from the TCGA database, with varieties 1–7 representing numerous subtypes: 1, lung bronchioloalveolar carcinoma mucinous; 2, lung bronchioloalveolar carcinoma nonmucinous; 3, lung acinar adenocarcinoma; 4, lung micropapillary adenocarcinoma; 5, lung papillary adenocarcinoma; 6, lung stable sample predominant adenocarcinoma; 7, mucinous (colloid) carcinoma. (D) Boxplot demonstrating HMGN1 expression in several lung most cancers subtypes from the GSE11969 dataset. *p < 0.05, **p < 0.01, ***p < 0.001, ns: not statistically important. (E–H) Receiver working attribute curve (ROC) evaluation of HMGN1 expression in LUAD from TCGA (E) and three GEO datasets (F–H). (I–J) Kaplan–Meier curves for general survival and disease-specific survival in LUAD sufferers, related to HMGN1 expression. (Ok–L) Total survival evaluation of HMGN1 in LUAD from two GEO datasets.
To additional search the opportunity of HMGN1 as a biomarker, we analyzed the diagnostic effectivity of HMGN1 expression in discriminating LUAD sufferers from wholesome people utilizing receiver working attribute (ROC) curves from TCGA and GEO datasets. The info confirmed that HMGN1 had a excessive space beneath the curve (AUC = 0.91) worth (Fig. 3E). Comparable excessive AUC values have been noticed in different three lung most cancers cohorts (AUC = 0.89 in GSE10072, AUC = 0.98 in GSE11969, and AUC = 0.87 in GSE19804) (Fig. 3F–H). And we analyzed the diagnostic effectivity of HMGN1 expression in discriminating LUAD sufferers from different lung most cancers utilizing ROC curves from GEO dataset. The info confirmed that HMGN1 had no excessive space beneath the curve worth (Supplementary Fig. 2A). To this point, the potential prognostic worth of HMGN members of the family stays unclear. Subsequently, the prognostic significance of HMGN1 mRNA expression have been evaluated through the use of three public out there datasets. Kaplan–Meier (KM) curve evaluation confirmed that HMGN1 expression was related to general survival (OS, Fig. 3I and Supplementary Fig. 2C) and disease-specific survival (DSS, Fig. 3J) in LUAD sufferers within the TCGA cohort. Nonetheless, the affiliation of HMGN1 mRNA expression with development free survival and illness free survival in LUAD was additionally seen not considerably in TCGA cohort (Supplementary Fig. 2B).Furthermore the affiliation of excessive HMGN1 mRNA expression with worse survival in LUAD was additionally seen in two different cohorts (Fig. 3Ok and L). Moreover, the univariate and multivariate Cox analyses have been carried out to analyze the connection between HMGN1 expression and OS or DSS. As indicated in Tables 1 and 2, the evaluation proven that T stage, N stage, HMGN1 expression have been related to OS and DSS in LUAD sufferers. These outcomes indicated that the expression of HMGN1 could be useful within the prognosis of LUAD sufferers.
Useful enrichment evaluation of the HMGN1 in LUAD
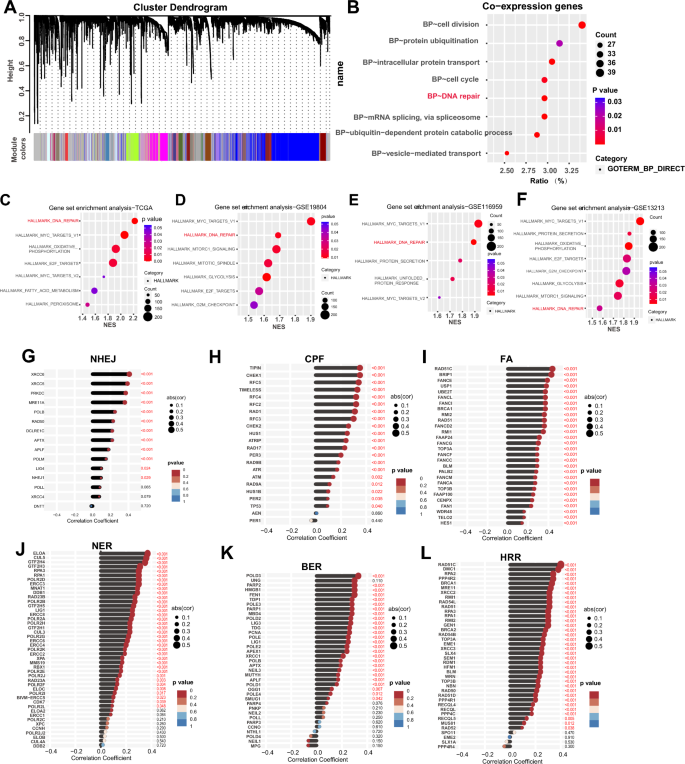
To research the perform of HMGN1 in lung adenocarcinoma, we first explored the potential co-expression genes of HMGN1 utilizing the weighted gene co-expression community evaluation (WGCNA) adopted by GO evaluation. The community constructed on 1157 genes led to the identification of 100 modules comprising genes with related expression patterns. GO evaluation additional identified that cell division, protein ubiquitination, intracellular protein, cell cycle, and DNA restore are the highest 5 modules which have been associated to the expression of HMGN1 in LUAD (Fig. 4A and B). To additional confirm the physiological capabilities of HMGN1, Gene Set Enrichment Evaluation (GSEA) evaluation was then undertaken to analyze hallmarks related to HMGN1 expression. The outcomes confirmed the affiliation of HMGN1 with DNA restore pathways (Fig. 4C and Supplementary 3F). Moreover, DNA restore have been the highest two gene units with important affiliation with HMGN1 in a number of LUAD cohorts (Fig. 4D–F and Supplementary 3A). Notably, GSEA analyses of HMGN1 in a number of different most cancers varieties have additionally emphasised the important roles of HMGN1 in DNA restore (Supplementary 3B–E and G). DNA restore pathways primarily include six modalities: base excision restore (BER), nucleotide excision restore (NER), crypto-chrome/photolyase household (CPF), Fanconi Anemia DNA Restore Pathway (FA), non-homologous finish becoming a member of (NHEJ), and homologous recombination restore (HRR). Correlation evaluation indicated that HMGN1 expression was linked to all of those six DNA injury restore pathways, particularly HRR (Fig. 4G–L).
Useful enrichment evaluation of HMGN1 in LUAD. (A) WGCNA cluster dendrogram displaying co-expression gene modules in LUAD; the ‘brown’ module incorporates genes co-expressed with HMGN1. (B) Bubble diagram of GO enrichment evaluation for HMGN1 co-expressed genes. (C–F) Bubble map of GSEA for HMGN1 from TCGA (C) and three GEO datasets (D–F). (G–Ok) Lollipop charts correlating HMGN1 with genes in several DNA restore pathways; CPF: verify level components; FA: fanconi anemia; NER: nucleotide excision restore; BER: base excision restore; NHEJ: non-homologous finish becoming a member of; HRR: homologous recombination restore.
HMGN1 loss reduces the DNA restore response
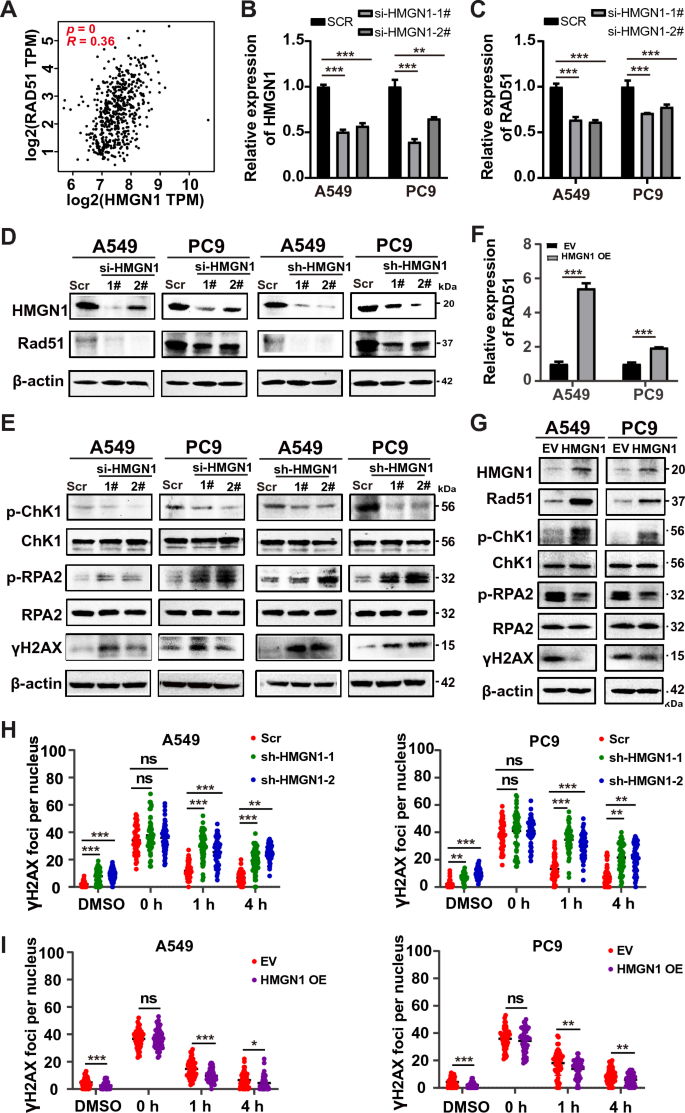
Provided that double-strand break (DSB) is probably the most deadly type of DNA injury and HMGN1 perform in DSB restore stays largely unknown, above evaluation prompted us to check whether or not HMGN1 is concerned in DSB restore in lung adenocarcinoma cells. First, the mRNA expression of HMGN1 confirmed optimistic correlation with RAD51, a key recombinase in DSB restore, within the TCGA LUAD cohorts (Fig. 5A). Subsequent, environment friendly depletion HMGN1 expression utilizing siRNA was confirmed on the RNA and protein degree in A549 and PC9 cells (Fig. 5B). We then assessed the impact of HMGN1 knockdown on Rad51 expression. The outcomes confirmed that RAD51 expression was remarkably decreased on the mRNA and protein degree (Fig. 5C). Importantly, p-CHK1, an integral part of the ATR-ChK1 pathway and DSB restore, was markedly diminished. On the identical time, p-RPA2, a readout of DSB finish resection in response to DNA injury, was remarkably collected after pulling down HMGN1 (Fig. 5D). In settlement with the western outcomes for Rad51, p-CHK1, and p-RPA2, HMGN1 depletion led to strong accumulation of γH2AX, a traditional DSB marker. Comparable phenomenon was additionally seen in HMGN1 stably knockdown A549 and PC9 cells (Fig. 5D and E). Quite the opposite, HMGN1 overexpression promoted DNA injury response and diminished γH2AX expression (Fig. 5F and G). Moreover, we’ve got carried out immunofluorescence staining for gamma-H2AX over time after the cells have been handled with etoposide. The immunostaining outcomes demonstrated that extra γH2AX foci remained in HMGN1-depleted cells than HMGN1-proficient cells (Fig. 5H and I, Supplementary Fig. 4), suggesting diminished DSB restore capability within the absence of HMGN1. In sum, these knowledge counsel that HMGN1 inhibition attenuates the ATR-ChK1 pathway and subsequent Rad51-mediated recombination, thereby impairing DSB restore.
Influence of HMGN1 knockdown on DNA injury response in lung adenocarcinoma. (A) Pearson correlation evaluation displaying the correlation between HMGN1 expression and RAD51 expression in LUAD. (B, C) RT–qPCR evaluation of HMGN1 silencing in A549 and PC9 cells and its impact on RAD51 expression. (D, E) Western blot evaluation of Rad51, p-RPA2, γH2AX and p-Chk1 proteins following HMGN1 depletion. (F) RT–qPCR evaluation of RAD51 expression upon ectopic expression of HMGN1. (G) Western blot evaluation of Rad51, p-RPA2, γH2AX and p-Chk1 proteins following HMGN1 overexpression. (G) Quantification of Gamma-H2AX fluorescence foci from three experiments proven as imply ± SD. *p < 0.05, **p < 0.01,***p < 0.001, ns: not statistically important.
HMGN1 depleted cells are extra delicate to HU therapy
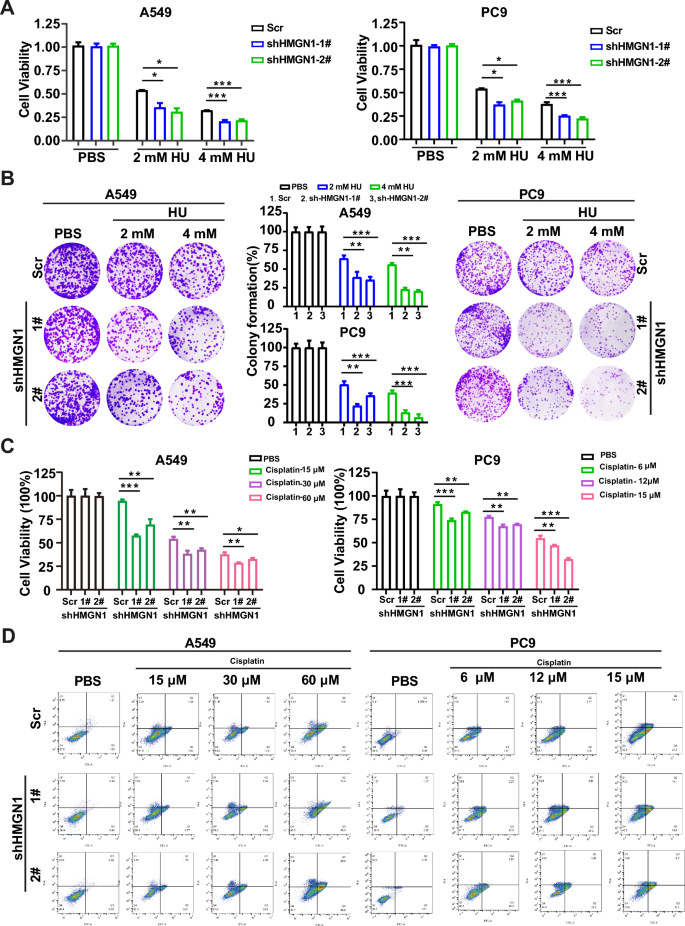
The indispensable function of HMGN1 in HRR prompted us to check the potential of HMGN1 inhibition to sensitize lung adenocarcinoma cells to DNA injury brokers, which is ceaselessly utilized in scientific setting. First, we investigated the impact of HMGN1 knockdown on the sensitivity of hydroxyurea (HU). As proven in Fig. 6A and Supplementary fig 5, HMGN1-deficent A549 and PC9 cells have been extra rather more delicate to HU therapy, which was in keeping with the function of HMGN1 within the ATR-ChK1 pathway. Furthermore, long-term colony formation assays demonstrated that HMGN1 might defend the cells from HU-induced toxicity (Fig. 6B). To discover the chemosensitizing potential of focusing on HMGN1, we handled A549 and PC9 cells with a spread of concentrations of cisplatin to evaluate cell survival within the absence of HMGN1. As displayed in Fig. 6C, HMGN1 silencing might improve the cytotoxic exercise of cisplatin, a generally used chemotherapeutic drug within the clinic. Concomitantly, a markedly enhance was seen in HMGN1-deficient cells in contrast with HMGN1-proficient cells (Fig. 6D).
The impact of HMGN1 knockdown on HU or cisplatin sensitivity in lung adenocarcinoma. (A, C) Cell viability assays displaying the consequences of cisplatin and hydroxycarbamide (HU) on A549 and PC9 cells with HMGN1 knockdown. (B) Clonogenic assays of A549 and PC9 cells handled with various concentrations of HU after HMGN1 knockdown. Surviving fraction percentages calculated relative to untreated cells. (D) Impact of HMGN1 depletion on cisplatin-induced cell apoptosis. *p < 0.05, **p < 0.01, ***p < 0.001, ns: not statistically important (Pupil’s t check).