Elevated GD2 expression upon RB1 or TP53 knockdown in a CRPC cell line mannequin
Given the GD2 overexpression in neuroectodermal lineage tumors3, and prior reviews that CRPC-NE differentiation is related to the acquisition of CSC options31, we surmised that GD2 overexpression may be a characteristic of CRPC upon experimental perturbations proven to advertise neuroendocrine differentiation. RB1 or TP53 depletion in androgen receptor-driven CRPC (CRPC-AR) cell traces32 or mouse fashions33 mixed with PTEN loss is thought to advertise NE trans-differentiation. Therefore, we in contrast the CRPC-AR cell line LNCaP C4-2 (known as C4-2), which has a mutationally-inactivated PTEN34, with its secure TP53, RB1 or TP53/RB1 knockdown (KD) derivatives as earlier research on this mannequin confirmed that TP53/RB1-KD cells had acquired NE differentiation28. qPCR evaluation verified the anticipated TP53 or RB1-KD (Fig. 1B). Western blotting confirmed the knockdown of RB1 and TP53 in particular person KD and DKD cells (Supplementary Fig. S1A). We validated a commercially accessible anti-GD2 antibody (clone 14.G2; Biolegend) by demonstrating its reactivity by FACS and immunohistochemistry (IHC) in opposition to a identified GD2+ neuroblastoma cell line SK-N-BE-235 and the shortage of its reactivity with a GD2– rhabdomyosarcoma cell line A20436 (Supplementary Fig. S1B,C). Utilizing this antibody, FACS staining revealed low cell floor GD2 expression on parental C4-2 cells, however the ranges of GD2 and the scale of the GD2+ subpopulation markedly elevated in RB1-KD, TP53-KD or double-KD (DKD) C4-2 derivatives (Fig. 1C). Notably, the rise in GD2 ranges on DKD was intermediate between that with particular person KD of RB1 or TP53, the latter being increased than with RB1-KD (the MFI and % GD2+ cells are indicated inside FACS plots). IHC analyses confirmed the upregulation of GD2 in RB1-KD and TP53-KD cells in comparison with the parental C4-2 cells (Fig. 1D). Elevated expression of GD2 upon TP53 or RB1-KD of C4-2 cells was related to elevated ranges of the important thing GD2 biosynthesis enzymes GM3 Synthase (GM3S), GD3 synthase (GD3S) and GD2 synthase (GD2S) (Fig. 1E).
Induction of excessive GD2 expression in of GD2 in CRPC cell line fashions with neuroendocrine (NE) differentiation. (A) Schematic of key GD2 biosynthetic steps displaying the enzymes (ST3GAL3, ST8SIA1 and B4GALNT1) and intermediate merchandise (GM3 and GD3) [Adapted from52]. (B and C) C4-2 cells with secure knockdown (KD) of TP53 or RB1, beforehand proven to bear NE differentiation, had been analyzed for GD2. (B) qPCR evaluation for TP53 and RB1 mRNA expression exhibits TP53-KD or RB1-KD cells relative to C4-2 cells. n = 3. (C) Cell floor GD2 expression of C4-2, C4-2 RB1-KD, TP53-KD and RB1/TP53 double knockdown (DKD) cells had been analyzed by dwell cell floor staining with anti-GD2 vs isotype management antibodies adopted by movement cytometry. X-axis, fluorescence depth; Y-axis, Occasions normalized to mode. FACS plots show % cells staining optimistic and imply fluorescence depth (MFI) of staining with anti-GD2 (blue) and isotype management (pink). (D) Immunohistochemistry (IHC) staining affirmation of elevated GD2 expression on TP53-KD or RB1-KD C4-2 cells relative to parental cells. (E) qPCR evaluation exhibits elevated expression of ST3GAL3, ST8SIA1 and B4GALNT1 mRNAs in TP53-KD and RB1-KD cells in comparison with parental C4-2 cell line. n = 3. Information are imply ± SEM with two-tailed unpaired t check. *, P < 0.05; **, P < 0.01.
Elevated GD2 expression in a subset of PC cell traces
Utilizing the C4-2 and its RB1 or TP53-KD derivatives as references, we carried out FACS evaluation of cell floor GD2 expression on a panel of 18 established human and murine PC cell traces, which included variants of generally used PC cell traces and two immortal prostate epithelial cell traces (PHPV18 and PSV40). Amongst these, a majority displayed little or no anti-GD2 staining above that with the management antibody (Desk 1, Supplementary Fig. S2A–C). In distinction, low however clearly detectable ranges of GD2 expression on a small subpopulation of cells had been noticed on 3 PC cell traces (ND1, C4-2 and C4-2B; starting from 0 to 2%) and two immortal prostate epithelial cell traces (PHPV18 and PSV40; 2 to three% and 4 to six%, respectively) (Desk 1, Supplementary Fig. S2A–C). Importantly, a comparatively massive subpopulation of GD2+ cells displaying excessive imply fluorescence intensities (MFI’s) was noticed in human CRPC cell traces 22Rv1 (%+, 20.1; MFI 870), which overexpresses wildtype AR and ARV7 splice variant37, and C4-2B cells rendered enzalutamide-resistant28 (%+, 50.6; MFI 2528), and on murine CRPC cell traces RM-1 (pushed by Ras and Myc)38 (%+, 54.6; MFI 1088) and TRAMP-C1 (transgenic adenocarcinoma mouse prostate [TRAMP]-derived)39 (%+, 28.6; MFI 908) (Desk 1, Supplementary Fig. S2A–C).
The GD3S enzyme (encoded by the ST8SIA1 gene) is rate-limiting for GD2 synthesis2. Per our staining knowledge, the CCLE-DepMap mRNA expression knowledge accessible for ten PC cell traces confirmed the best worth for the GD2excessive 22Rv1 and the second lowest worth for the GD2– PC3 cell line (Supplementary Fig. S3, higher panel). Evaluation of transcriptomics knowledge of organoids and PDX tumors lately used to categorise CRPC into 4 subtypes, CRPC-AR (AR-driven), CRPC-NE (neuroendocrine), CRPC-WNT (WNT pathway-driven) and CRPC-SCL (stem cell like)26, revealed excessive ST8SIA1 mRNA expression in GD2excessive 22Rv1 and almost undetectable ranges in GD2– LNCaP cell line (Supplementary Fig. S3, higher panel). Utilizing the degrees in these cell traces as reference, > 20% of CRPC organoids and PDX tumors confirmed increased ST8SIA1 expression (these above the pink cutoff line equivalent to TPM values for LNCaP), and this was seen throughout the CRPC subtypes (Supplementary Fig. S3). Curiously, the ST8SIA1 TPM values in a lot of organoid fashions had been considerably increased than these for 22Rv1(e.g., CRPC-AR samples MSK-PCa19, MSK-PCa22; CRPC-NE samples MSK-PCa10, MSK-PCa24; and CRPC-SCL pattern MSK-PCa8). Thus, our outcomes present {that a} subset of established PC cell traces, together with examples of CRPC, categorical low to excessive GD2 ranges on a variable fraction of cells, paying homage to research with breast most cancers cell fashions9,10, and the ST8S1A1 expression in organoid and PDX fashions supported the potential of excessive GD2 expression in patient-derived PCs.
GD2 expression is seen on a small subset of tumor cells in main PC affected person tissues and a better proportion of metastatic lesions
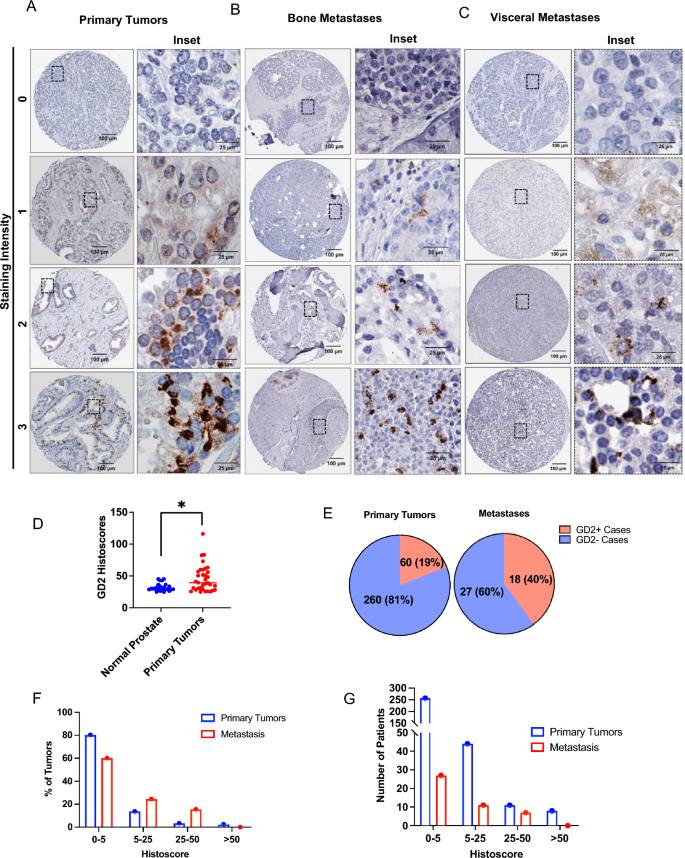
To evaluate if GD2 expression on a subpopulation of cells in PC cell traces could possibly be prolonged to human PC, we used tissue microarrays (TMAs) from two PC affected person cohorts (supplied by the Prostate Most cancers Biorepository Community; PCBN). IHC staining of a 320-sample TMA of paired regular prostate tissues and PC samples (Gleason scores of three–5) demonstrated a statistically vital overexpression of GD2 on main PCs in comparison with paired regular prostate tissue (Fig. 2A–D); not like uniform GD2+ neuroblastomas, however just like triple-negative breast cancers, solely a small subset of tumor cells in PC samples was GD2+. General, 19% of main PCs scored GD2+ primarily based on a histoscore cutoff of < 5 (Fig. 2E). Notably, IHC evaluation of a metastatic PC TMA (the 45-Case Bone and Visceral Metastasis from Fast Post-mortem TMA from PCBN) confirmed GD2+ staining in about twice the share of metastatic lesions in comparison with main tumors (Fig. 2E). The histoscore distribution evaluation confirmed that metastatic tumors exhibited increased GD2 expression (Fig. 2F,G). These outcomes point out that GD2-expression is a characteristic of a subpopulation of tumor cells in a subset of PC sufferers, particularly in metastases.
GD2 expression on a subset of tumor cells in main prostate most cancers tissue with increased expression in metastatic lesions. Tissue microarrays (TMAs) composed of paired regular prostate and prostate most cancers Gleason grades (3, 4 and 5) (n = 320) or 45 circumstances of bone and visceral metastatic samples (each obtained from the Prostate Most cancers Biorepository Community) had been analyzed by IHC staining for GD2. Histoscores had been calculated by multiplying the staining depth (0, destructive; 1, low; 2, average optimistic; 3, sturdy optimistic) by the % tumor cells staining optimistic. (A, B) Consultant examples of destructive, low, average, and excessive GD2 staining in (A) main, (B) bone metastatic and (C) visceral metastatic tumor samples. (D) Scattered plot exhibits histoscore distribution of main tumors and regular prostate. GD2 expression is considerably increased in main tumors than regular prostate. Information represents 34 circumstances of GD2 expression in tumors vs 24 in regular tissue, contemplating histoscore > 25 as optimistic. Mann–Whitney unpaired t-test used as statistical evaluation; *, P < 0.05. (E) Pie charts displaying the % of circumstances that had been GD2+ or GD2– in main and metastatic tumors samples. Histoscores of 0–5 had been thought of destructive whereas scores increased than 5 had been thought of optimistic. (F, G) Frequency distribution plots histoscores plotted in opposition to (F) share of sufferers and (G) variety of sufferers. The indicated histoscore grouping are: 0–5 (destructive); 5–25 (low), 25–50 (average); and > 50 (sturdy).
GD2 overexpression defines a PC cell inhabitants with increased tumorigenic potential in vitro
To evaluate the potential useful function(s) of GD2 overexpression in PC, we first in contrast the parental C4-2 cells (GD2low) with their GD2excessive RB1-KD, TP53-KD or DKD cell traces. Cell Titer-Glo assays confirmed that RB1-KD, TP53-KD or DKD derivatives exhibited increased (fold improve of ~ 6, ~ 20, ~ 31 and ~ 23 for parental, RB1-KD, TP53-KD and DKD, respectively; p values < 0.01 to < 0.001 on the final time level) and sooner cell proliferation, with the TP53-KD cells, which additionally exhibited increased GD2+ subpopulation, exhibiting the best charge and degree of proliferation (Fig. 3A, left panel). Equally, the GD2excessive enzalutamide resistant C4-2BER confirmed elevated proliferation in comparison with GD2low management C4-2B parental cells (fold improve of ~ 60, vs. ~ 11; p worth < 0.001 at varied time factors) (Fig. 3A, proper panel). Transwell migration analyses confirmed a better migratory habits of GD2excessive derivatives (~ twofold for RB1-KD, and ~ 3.5-fold for TP53-KD and DKD respectively in comparison with C4-2; ~ fourfold for C4-2BER vs. C4-2B) in comparison with GD2low parental cell traces (Fig. 3B). Importantly, the GD2excessive derivatives of C4-2 exhibited considerably elevated 3D tumor-sphere progress, an in vitro correlate of CSC exercise40, with RB1-KD yielding ~ 4.5-fold and TP53-KD or DKD cell traces yielding ~ ninefold extra tumorspheres (p-values < 0.001) in comparison with their GD2low parental C4-2 cells (Fig. 3C, higher panel). Equally, GD2excessive C4-2BER cells exhibited a ~ 4.5-fold improve in tumorspheres in comparison with parental C4-2B cells (p-value < 0.001) (Fig. 3C, center panel). Extra tumorsphere assays on ultra-low attachment situations within the absence of added 4% Matrigel, which we utilized in above assays to acquire extra uniform tumorspheres, confirmed the above findings and excluded the likelihood that elevated tumorsphere progress of GD2excessive C4-2 derivatives was as a result of adhesion skills supplied by Matrigel (Fig. 3C, decrease panel).
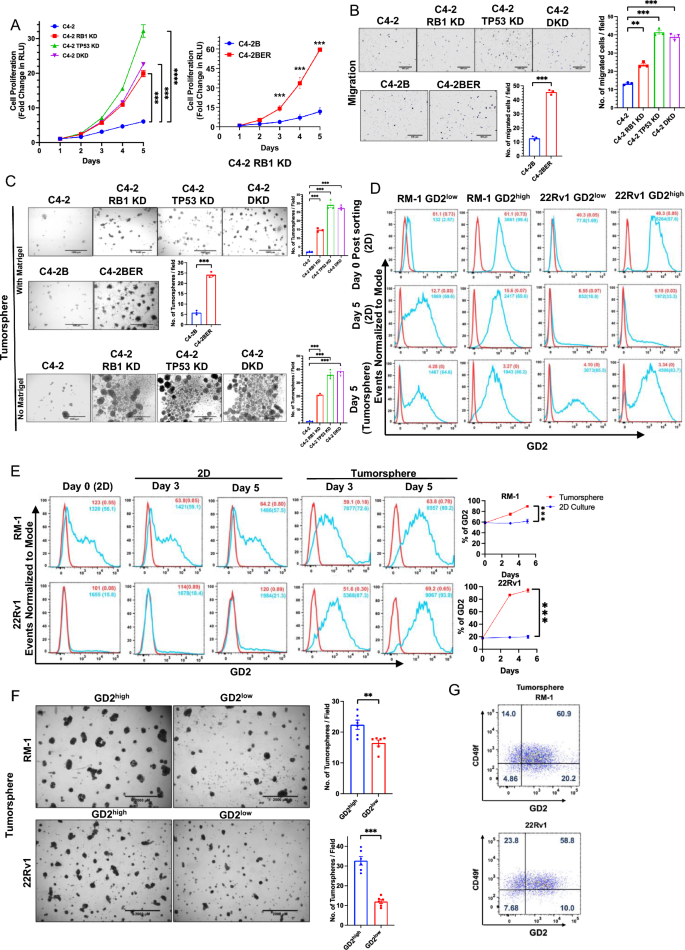
GD2 overexpression promotes the in vitro oncogenic traits: (A) Evaluation of cell proliferation. The parental C4-2 and its TP53-KD or RB1-KD or TP53/RB-KD (DKD) cells and C4-2B and its enzalutamide resistant variant C4-2BER had been plated in 96-well plates and cell proliferation was assessed on the indicated days utilizing the CellTiter-Glo luminescent cell viability assay. The Y-axis represents the fold improve in relative luminescence models (RLU) relative to Day 1. n = 3 with six replicates every (B) Evaluation of cell migration in indicated cell traces utilizing 20,000 cells as enter. Left, consultant photos. Proper, quantification. n = 3 with three replicates every. (C) Evaluation of tumorsphere forming capacity. The indicated cell traces had been plated in 24-well ultra-low attachment plates in presence of 4% Matrigel (higher and center panels) and in absence of Matrigel.(decrease panel) and pictures had been obtained on Day 7.Left, consultant photos; scale bar, 1000 µm. Proper, quantified tumorsphere numbers per 4X microscopic discipline. n = 3 with three replicates every and 6 photos per properly (D) GD2excessive and GD2low inhabitants had been FACS sorted in RM-1 and 22Rv1 cells and grown as 2D and tumorspheres. After Day 5, the cells had been reanalyzed for GD2 fractions in each GD2excessive and GD2low submit sorted cells. In each tumorspheres and 2D tradition the GD2low cells re-expressed GD2. (E) Enrichment of GD2excessive cells in tumorspheres of CRPC cell traces. The indicated cell traces had been cultured in 2D or had been seeded in tumorsphere cultures. Cells harvested on the indicated instances had been analyzed for cell floor GD2 expression (vs. isotype management) on the indicated instances. % GD2+ cells and MFI are indicated. Left, consultant FACS photos; Proper, quantitation of information. two-way ANOVA check. (F) GD2excessive fraction of CRPC cell traces is enriched for tumorsphere-forming capacity. Constitutively GD2-expressing RM-1 and 22Rv1 CRPC cell traces had been FACS sorted for GD2excessive and GD2low populations, grown on 2D in a single day and plated at 20,000 cells per properly of 24-well ultra-low attachment plates in tumorsphere media containing 4% Matrigel. Tumorspheres had been imaged on Day 5. Left, consultant photos; scale bar, 2000 µm. Proper, Quantification of tumorspheres (> 250 µm) per 2× magnification discipline. n = 3 with three replicates every and 6 photos per properly. (G) FACS evaluation of tumorsphere-grown RM-1 and 22Rv1 cell traces after anti-GD2 and anti-CD49f double-staining exhibits a majority of GD2+ cells to be CD49fhello % GD2 and/or CD49f +/− populations are proven within the respective quadrants. Information represented in all experiments are imply ± SEM, unpaired t-test; ns, not vital; *, P < 0.05; **, P < 0.01; ***, P < 0.001.
Earlier research have proven that GD2-expressing cell fraction is enriched in tumorigenic behaviors9. We subsequently used FACS-sorting to purify the GD2excessive and GD2low populations of 22Rv1 and RM-1 cell traces as these cell traces harbor each GD2excessive and GD2lowcell subpopulations (Desk 1). Per earlier research demonstrating that GD2 expression on cell traces is dynamic41, the FACS-sorted populations drifted in the direction of a mixture of GD2excessive and GD2low populations upon continued 2D or 3D tradition (Fig. 3D). Notably, FACS evaluation revealed a big enrichment for the GD2excessive inhabitants throughout tumorsphere progress vs 2-D tradition in each RM-1 (72% and 89% at days 3 and 5 in 3D, and 56% and 59% at 3 and 5 days in 2D, vs. 56% on Day 0; p < 0.001) and 22Rv1 (87% and 94% at days 3 and 5 in 3D, and 18% and 21% at 3 and 5 days in 2D, vs. 16% on Day 0; p < 0.001) cell traces (Fig. 3E). An identical, time-dependent, improve within the GD2excessive inhabitants was noticed upon tumorsphere vs. 2D progress of C4-2 (GD2low) and its RB1-KD (GD2excessive) spinoff (Supplementary Fig. S4). Regardless of the dynamic nature of GD2 expression, the GD2excessive fractions of each 22Rv1 and RM-1 cell traces yielded considerably extra tumor-spheres (~ 22/discipline vs. 16/discipline for GD2excessive and GD2low RM-1, and 32/discipline vs. 11/discipline for GD2excessive and GD2low 22Rv1; p < 0.01 and < 0.001, respectively) (Fig. 3F) indicating a better tumorsphere-forming capacity of GD2excessive subpopulation in PC cell traces. Twin FACS staining of RM-1 and 22Rv1 cell traces grown as tumorspheres for CD49f. and GD2 demonstrated {that a} majority (60.9% for RM-1 and 58.8% for 22Rv1) of GD2excessive cells had been throughout the CD49fexcessive fraction (Fig. 3G). These outcomes instructed that GD2excessive fraction defines a cell inhabitants with increased oncogenic potential and doubtlessly the CSC-like habits.
GD3 synthase, the rate-limiting enzyme in GD2 biosynthesis, is required for in vitro oncogenic attributes of GD2excessive CRPC cell traces
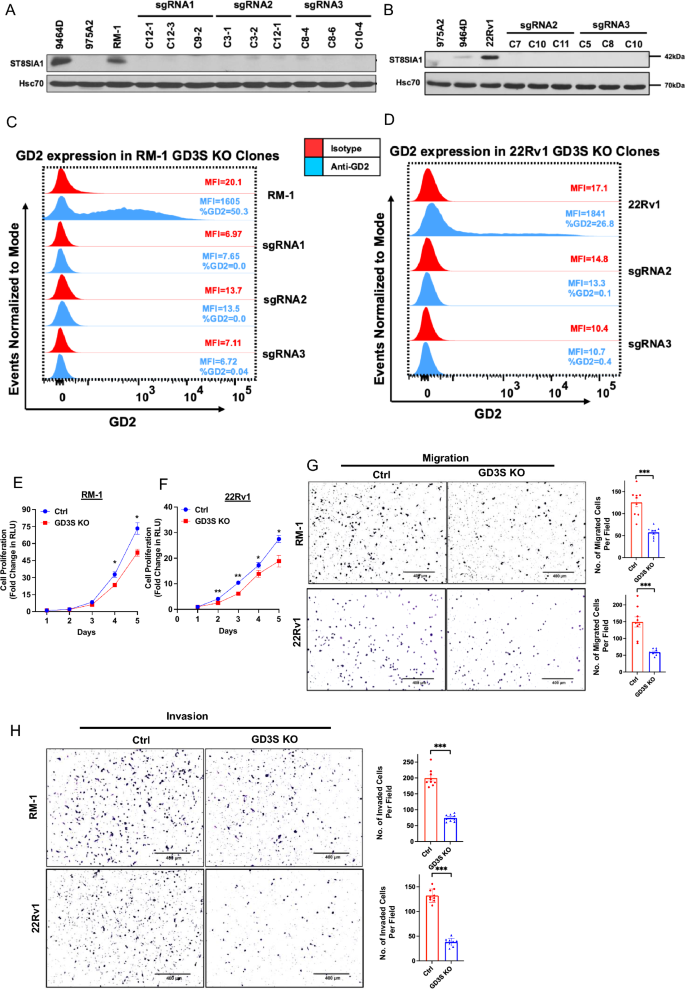
In view of our outcomes above, we hypothesized that the GD2+ cell subpopulation is the most important driver of the oncogenic attributes of PCs. To check this speculation, we focused the rate-limiting GD2 biosynthesis enzyme GD3S2 for CRISPR-Cas9 knockout (KO), utilizing a single-vector Cas9/sgRNA expression system, in GD2-overexpressing mouse (RM-1) and human (22Rv1) CRPC cell traces and obtained clonal traces by serial limiting dilution cloning. Western blotting confirmed the lack of GD3S expression in GD3S sgRNA-targeted derivatives (Fig. 4A, B). These GD3S-KO clones certainly lacked the cell floor GD2 expression by FACS evaluation (Fig. 4C, D). We additionally analyzed the parental vs. GD3S-KO RM-1 and 22Rv1 cell traces along with GD2+ 9464D and GD2– 975A2 mouse neuroblastoma cell traces41 by skinny layer chromatography (TLC) and TLC/immunoblotting, wherein lipid extracts of cells had been resolved by TLC subsequent to the purified GD2 commonplace, adopted by both chemical staining to visualise the separated sphingolipid species or immunoblotted with the 14G2a antibody, respectively. Based mostly on pilot research utilizing varied ratios of the chloroform/methanol/0.2% CaCl2 solvent system utilized in prior research42,43,44, we resolved the sphingolipid extracts of cell traces utilizing the 50:40:10 ratio of chloroform/methanol/0.2% CaCl2 as these had been optimum to resolve GD2 and GD3 species from different lipids. Phosphomolybdic acid (PMA) staining of the resolved species on the TLC plate confirmed lipid species comigrating with commonplace GD2 in RM-1, 22Rv1 and the optimistic management 9464D cells, whereas these species had been absent within the GD3S-KO cell traces and the GD2 destructive 975A2 cells (Supplementary Fig. S5A). Notably, 14G2a immunoblotting of the TLC-resolved sphingolipid species confirmed sturdy reactivity with the GD2 commonplace, as anticipated; moreover, the 14G2a antibody detected the lipid species comigrating with commonplace GD2 in parental RM-1 or 22Rv1 and within the optimistic management cell line 9464D (equivalent to the comigrating species detected by chemical staining) however not in GD3S-KO RM-1 or 22Rv1 cell extracts or within the destructive management cell line 975A2 (Supplementary Fig. S5B). These outcomes additional verify the reactivity of 14G2a with GD2 and the absence of its expression upon GD3S-KO. As GD3 is the instant product of GD3S exercise and a precursor of GD2, we additionally assessed the degrees of GD3 in panels of PC cell traces (Supplementary Fig. S6). As well as, parental vs. GD3S-KO CRPC cell traces RM-1 and 22Rv1 had been assessed for expression of GD3 utilizing FACS. Comparatively low to modest ranges of GD3 had been seen on parental cells and this expression was eradicated in KO cells (Supplementary Fig. S7). Thus, GD3S-KO successfully eradicated the expression of GD2 (and GD3) on CRPC cell traces.
GD3S knockout in CRPC cell traces impairs the in vitro tumorigenic and pro-metastatic traits. (A-D) Technology of GD3S knockout of RM-1 and 22Rv1 CRPC cell traces. Cells had been transduced with lentiviral All-in-One CRISPR/Cas9 constructs and secure clones analyzed by Western blotting with anti-GD3S antibody; Hsc70, loading management. Recognized GD2+ (9464D) and GD2– (975A2) mouse neuroblastoma cell traces had been used as controls. (C, D) Full lack of cell floor GD2 expression in GD3S-KO RM-1 and 22Rv1 clones. The parental cell traces and their GD3S-KO clones had been live-cell stained with anti-GD2 (blue) or isotype management (pink) antibody adopted by FACS evaluation. FACS plots point out imply fluorescence depth (MFI) on X-axis vs. Occasions Normalized to Mode on Y-axis. % optimistic cells and MFIs are indicated in FACS plots. (E, F) Decreased CRPC cell proliferation upon GD3S-KO. Three clones of parental and GD3S-KO cell traces maintained individually had been blended plated at 1,000 cells per properly in 96-well plates and dwell cells quantified on the indicated instances utilizing the CellTiter-Glo cell viability assay. The Y-axis represents the fold improve in relative luminescence models (RLU) relative to Day 1. n = 3 with six replicates every. (G, H) Decreased CRPC cell migration (G) and invasion (H) upon GD3S-KO. 20,000 parental or GD3S-KO RM-1 or 22Rv1 cells had been plated in low serum medium in high chambers of transwell chambers with out (migration) or with (invasion) Matrigel coating and allowed emigrate or invade for 16 h in the direction of serum-containing medium within the backside chambers. Left, consultant photos. Proper, quantification. n = 3 with three replicates every and 6 photos per properly. All Information represents imply ± SEM with unpaired t check; **, P < 0.01; ***, P < 0.001, ***p < 0.001.
We additionally generated GD3S-KO spinoff of the enzalutamide-resistant C4-2BER cell line, which has been reported to precise NE markers28 and expresses excessive GD2 ranges in comparison with its parental cell line (Supplementary Fig. S2C and S8A). GD3S-KO eradicated the expression of GD2 on this mannequin as properly (Supplementary Fig. S8B). Moreover, GD3S-KO led to discount within the expression of NE differentiation markers, neuron particular enolase (NSE), Enhancer of Zeste homolog 2 (EZH2) and chromogranin A (CHGA) in comparison with C4-2BER cells, with ranges in GD3S-KO cells akin to these in GD2low C4-2B cells (Supplementary Fig. S8C). These outcomes recommend GD2 expression might regulate NE differentiation.
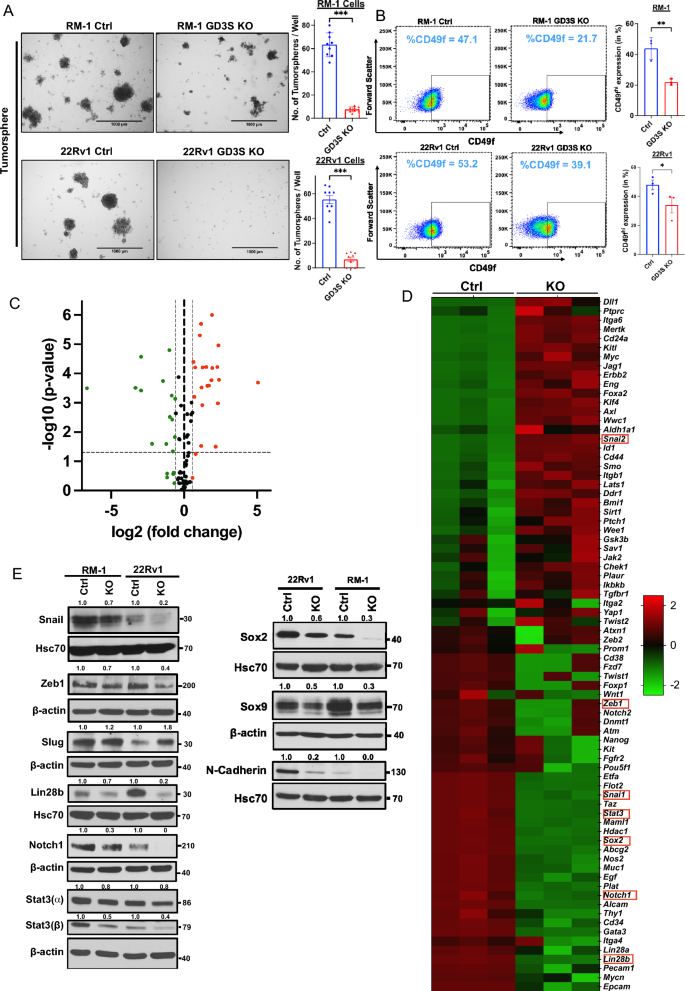
Practical analyses of varied oncogenic traits confirmed a big influence of GD3S-KO. Utilizing CellTiterGlo assays, the GD3S-KO RM-1 cells exhibited a fold-increase of ~ 51-fold in comparison with ~ 73-fold for management cells, whereas GD3S-KO 22Rv1 cells confirmed a ~ 18-fold-increase in comparison with a ~ 28-fold improve for parental cells, with the variations being statistically vital (p < 0.05) (Fig. 4E,F). In cell migration assays, GD3S-KO RM1 and 22Rv1 cell traces exhibited a 55% or 65% decrease migration, respectively, relative to their controls (p < 0.001) (Fig. 4G). An identical impairment of transwell invasion capacity was seen upon GD3S-KO, with the GD3S-KO RM-1 and 22Rv1 cell exhibiting 67.5% and 73% decrease invasion, respectively, relative to their parental cell traces (p < 0.001) (Fig. 4H). Tumorsphere assays confirmed that GD3S-KO RM-1 and 22Rv1 cell traces exhibited 88% or 90% lowered tumorsphere forming skills in comparison with their parental non-targeted cells (p < 0.001) (Fig. 5A), supporting the significance of GD3S and its ganglioside merchandise in selling the CSC habits of CRPCs. Additional supporting this notion, FACS evaluation for the proportion of CD49fhello cells, a marker related to PC CSCs45, confirmed a discount from 47 to 22% for GD3S-KO RM-1 (p < 0.01) and 53% to 39% for GD3S-KO 22Rv1(p < 0.05), relative to their parental traces (Fig. 5B). To additional consider the potential linkage of GD3S and GD2 expression with CSC traits, we analyzed the mRNA expression ranges of 84 mouse genes included in a industrial CSC signature panel (RT2 profiler array) utilizing quantitative PCR (qPCR) evaluation of management vs. GD3S-KO RM-1 cells. The represented genes correspond to these related to CSCs in addition to Epithelial Mesenchymal Transition (EMT), given the sturdy mechanistic linkage of EMT and CSC behaviors46. Volcano plot evaluation revealed a number of genes within the array to be both down-regulated or upregulated within the GD3S-KO spinoff vs the management RM-1 cells (Fig. 5C). As offered in a heatmap (Fig. 5D), we noticed a big downregulation of 17 genes together with Notch1, Stat3, Sox2, Pou5f1(Oct4), Mycn, Lin28b, Snai1, Zeb1, Epcam, Pecam1 and upregulation of twenty-two genes reminiscent of CD24a, Mertk, Slug, Foxa2, CD44, Aldha1a, Klf4 and Myc in GD3S-KO cells (Fig. 5D). Western blotting evaluation validated the qPCR-observed downregulation of Snail, ZEB1, Lin28b, Notch1, Stat3 (⍺, β), SOX2 and the noticed upregulation of Slug in GD3S-KO RM-1 and 22Rv1 cells vs. their management cells (Fig. 5E). Extra Western blot evaluation of CSC/EMT markers SOX9 and N-cadherin revealed their downregulation in GD3S-KO RM-1 and 22Rv1 cells vs. their management cells (Fig. 5E). Collectively, these outcomes are in keeping with a conclusion that the GD2excessive fraction in CRPC fashions defines a CSC-like inhabitants.
Impairment of most cancers stem cell (CSC)-associated traits in CRPC cell traces upon GD3S-KO. (A) Decreased CRPC tumorsphere forming capacity upon GD3S-KO. The indicated cell traces had been plated in 24-well ultra-low attachment plates in 4% Matrigel and pictures had been obtained on Day 7. Left, consultant photos; scale bar, 1000 µm. Proper, quantified tumorsphere numbers per 4X microscopic discipline. n = 3 with three replicates every and 6 photos per properly (B) Decreased expression of CSCs markers in GD3S-KO CRPC cell traces. The indicated cell traces had been subjected to dwell cell staining adopted by Left, consultant FACS plots show CD49f. staining (MFI’s) on X-axis and ahead scatter on Y-axis. Proper, quantitation of % of CD49f.-high inhabitants. n = 3. (C) Volcano plot of the mouse most cancers stem cell RT2 profiler quantitative PCR array evaluation of RM-1 controls vs. GD3S-KO cells. X-axis, log fold change; Y-axis, -log p-value. Inexperienced dots characterize the downregulated genes and pink dots show the upregulated one with a log1.5-fold change cut-off. (D) The heatmap exhibits the considerably upregulated (pink) and downregulated (inexperienced) genes in RM-1 GD3S-KO cells in comparison with management cells.; n = 3 unbiased experiments, p < 0.05 is deemed vital. (E) Decreased expression of most cancers stem cells and epithelial-mesenchymal transition (EMT) markers in GD3S-KO CRPC cell traces. Lysates of the parental vs. GD3S-KO RM-1 or 22Rv1 cell traces had been subjected to western blotting with antibodies in opposition to the indicated proteins, with Hsc70 or β-actin used as loading controls. Densitometric quantification is proven on high of every blot. Information represents imply ± SEM, unpaired t check; *p < 0.05, **p < 0.01, ***p < 0.001.
GD3S-KO impairs the tumorigenic capacity of GD2excessive CRPC cell traces
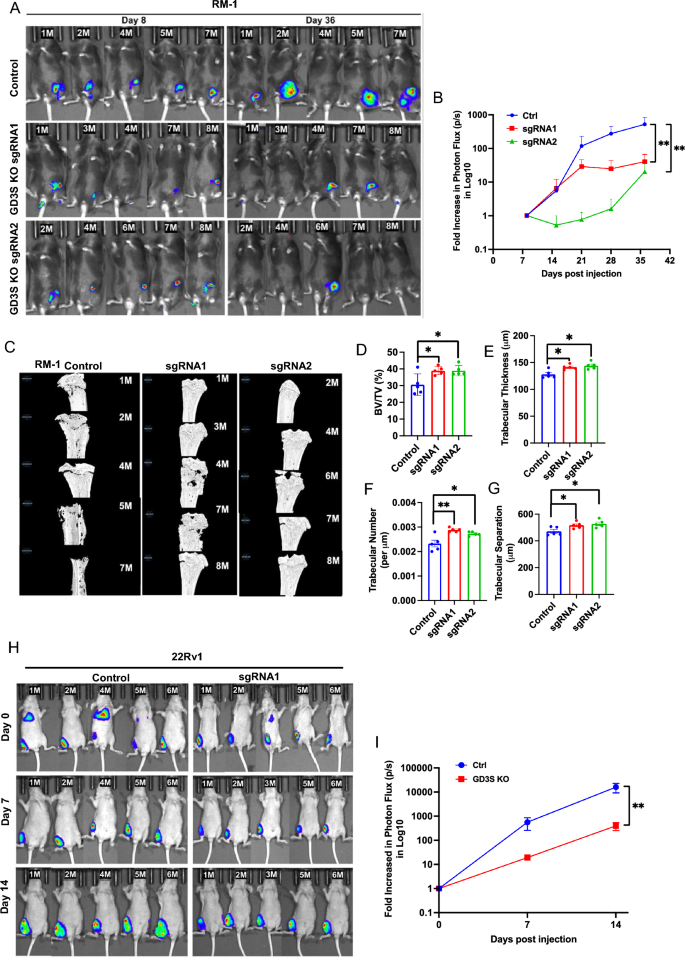
To evaluate if the marked discount within the in vitro pro-oncogenic traits of PC cell traces upon GD3S-KO interprets into impaired tumorigenesis in vivo, we used lentiviral an infection to introduce a tdTomato-luciferase twin reporter into management and GD3S-KO RM-1 cell traces (the latter with two distinct sgRNAs). The tdTomato-high fractions had been enriched by FACS-sorting and secure cell traces chosen. We implanted the reporter-bearing management and GD3S-KO RM-1 cells within the tibias of castrated syngeneic C57BL/6 mice and monitored tumor progress (5 mice/group) over time by bioluminescence imaging. Whereas management RM-1 cell implants generated tumors that confirmed a time-dependent improve in log10 photon flux in IVIS imaging that peaked at 512-fold improve on the endpoint in comparison with day 7, each of the GD3S-KO RM-1 cell implanted teams generated considerably smaller tumors whose common peak photon flux was 40-fold (sgRNA1) and 20-fold (sgRNA2) decrease in comparison with the management group (p < 0.01 for each KO teams relative to manage) (Fig. 6A,B). Morphometric evaluation of tibial bone by micro-CT scanning confirmed lowered bone destruction by implants of GD3S-KO RM-1 cells relative to manage as demonstrated by quantification of the bone quantity (38.85% and 38.86% for sgRNA1 and sgRNA2 teams, respectively, relative to 30.5% in management; p values for each < 0.05), trabecular thickness (thickness of 141mm and 144 mm for sgRNA1 and sgRNA2 teams, respectively, relative to 127 mm in management; p values for each < 0.05), trabecular quantity (trabecular quantity per mm of 0.00288 and 0.00278 for sgRNA1 and sgRNA2 teams, respectively, relative to 0.0023 in management; p values of < 0.01 and < 0.05 for sgRNA1 and sgRNA2 teams, respectively) and separation (516mm and 527mm for sgRNA1 and sgRNA2 teams, respectively, relative to 470 mm in management; p values for each < 0.05) (Fig. 6C–G). To substantiate the influence of GD3S-KO to impair the flexibility of implanted PC cells to kind tumors in vivo, we additional implanted the mCherry-luciferase expressing parental or GD3S-KO 22Rv1 cells intra-tibially in nude mice and analyzed these by IVIS imaging. In comparison with a 15,984-fold improve in Log10 photon flux relative to time 0 for the management group, the GD3S-KO cell implants confirmed a rise of solely 397.5-fold in peak Log10 photon flux (p < 0.001) (Fig. 6H,I). Utilizing IVIS, we didn’t observe distal metastases throughout the time of statement utilized in our research (though some animals confirmed bioluminescent alerts on the preliminary statement time level) (Fig. 6H). Collectively, these analyses show that GD3S-KO in CRPC cell line fashions considerably impairs their capacity to kind tumors when implanted in bone and to induce bone destruction.
Impairment of in vivo bone-implanted CRPC tumorigenesis by GD3S-KO. (A) Parental (Management) RM-1 cells or their edited variations carrying sgRNA1 or sgRNA2 (GD3S-KO) had been engineered with tdTomato-luciferase and 4 × 104 cells injected in tibias of castrated male C57/BL6 mice (8/group) and first tumor progress was monitored by bioluminescence imaging on the indicated time factors in mice (5/group) with detectable bioluminescent alerts on day 8. (B) Bioluminescence alerts of RM-1 tumors over time are proven as log fold-change in photon flux over the time. Information are imply ± SEM, n = 5; Two-way ANOVA (blended mannequin); *p < 0.05. (C) Micro-CT scanned photos of tibias remoted from mice implanted with the indicated RM-1 cell traces. μCT analyses of the trabecular bone was used to calculate the: (D) bone quantity fraction (BV/TV%); (E) trabecular thickness (µm); (F) trabecular quantity (per µm); and (G) trabecular separation (µm). Information characterize imply ± SEM, with unpaired t-test used for statistical evaluation; *p < 0.05 and ** p < 0.01. (H) Impairment of tumorigenesis of 22Rv1 CRPC cells by GD3S-KO. 1 × 105 management or GD3S-KO (sgRNA1) 22Rv1 cells engineered with mCherry-luciferase had been injected in tibias of castrated male athymic nude mice (6/group) and tumor progress was monitored by bioluminescence imaging in mice (5/group) with detectable bioluminescent alerts on day. (I) Bioluminescence alerts of 22Rv1 xenografts over time are proven as log fold-change in photon flux over time. Information are imply ± SEM, n = 5, with evaluation by Two-way ANOVA (blended mannequin); **p < 0.01.