Acute myeloid leukemia (AML) is a heterogeneous blood-borne malignancy with an total 5-year survival price of 25 % [1]. Regardless of substantial genetic characterization within the final many years, just a few focused therapies, resembling FLT3 inhibitors, have entered scientific observe. Remedy nonetheless largely depends on chemotherapy and hematopoietic stem cell transplantation in youthful sufferers, and epigenetic therapies as azacytidine together with the Bcl-2 inhibitor venetoclax in aged sufferers, with the primary inflicting comparatively excessive charges of therapy-associated morbidity and mortality, and the latter displaying restricted responses [2]. Due to this fact, the identification of recent actionable vulnerabilities, past genetics and epigenetics, is extremely demanded. Approaches investigating aberrant mechanisms on protein stage, resembling disease-specific post-translational modifications (PTM), maintain nice promise to supply new therapeutic targets. Ubiquitylation marks the key PTM that regulates the abundance of proteins, thus orchestrating key processes as proliferation, survival, and differentiation [3, 4]. SCF (SKP1/CUL1/F-box) E3 ubiquitin ligases are characterised by modular complexes containing one of many 72 particular person F-box proteins as substrate recruiting receptors, the F-box adapter SKP1, the scaffold protein CUL1, and the RING protein RBX1 [5]. Whereas most SCF complexes haven’t but been linked to substrates or organic actions, particular person members have been discovered to play key roles in numerous tumor entities [6, 7]. Current reviews have now supplied first insights into the roles of SCF E3 ligases in AML. Subsequent to figuring out the F-box protein FBXL2 as a tumor suppressor [8], the NEDD8 inhibitor pevonedistat confirmed some further profit when mixed with azacytidine in AML sufferers in scientific trials [9], and is present process additional investigation in AML and additional hematological malignancies. Notably, NEDD8 inhibitors forestall the neddylation-dependent activation of all cullin-based RING E3 ubiquitin ligases (CRLs), which embody SCF complexes. We subsequently hypothesized that some SCF ubiquitin ligases, or their substrate-recruiting F-box proteins, are particularly deregulated in AML, thus representing potential therapeutic targets.
With a purpose to display screen for AML-specific vulnerabilities throughout the household of SCF complexes, we first generated a targeted CRISPR/Cas9-based knockout library protecting all 72 human genes coding for F-box proteins (sgRNA sequences derived from the genome-scale CRISPR Knock-Out library GeCKOv2 [10]), along with constructive and detrimental controls. Utilizing this tradition library, we carried out pooled CRISPR/Cas9 drop-out screens in two unbiased AML cell strains and recognized the poorly studied F-box protein FBXL6 as essentially the most outstanding widespread dependency between OCI-AML3 (DNMT3A and NMP1 mutated) and MOLM-13 (FLT3-ITD) cells (Fig. 1A, Supplementary Desk 1). Confirmatory assays certainly demonstrated a big lower in proliferation upon knockout of FBXL6 in AML cell strains as decided by aggressive progress experiments, trypan blue exclusion cell counting and MTS assays (Fig. 1B–E, Supplementary Fig. 1A). Notably, the results of FBXL6 loss had been stronger within the two FLT3-ITD mutated AML cell strains MOLM-13 and MV4-11 as in comparison with the DNMT3A and NMP1 mutated OCI-AML3 line (Fig. 1B–E, Supplementary Fig. 1A), doubtlessly pointing in direction of a extra particular dependency.
A Outcomes from CRISPR/Cas9 drop-out screens utilizing a custom-designed sgRNA knockout library concentrating on all 72 human F-box proteins in 2 AML cell strains. The delta of normalized and log remodeled sgRNA learn‐counts of day 14 and day 0 was calculated and scores of all sgRNAs concentrating on the identical gene or controls had been averaged and are proven as one knowledge level. B Aggressive progress assay of Cas9-expressing AML cell strains transduced with GFP-expressing sgRNA constructs concentrating on FBXL6 (sgFBXL6), POLII (sgPOLII) as constructive management or non-targeting management (sgCtrl) at 30–50% effectivity. The ratio of GFP constructive to non-transduced cells was measured by move cytometry on the indicated days and normalized to day 2. ***P < 0.001; **P < 0.01; *P < 0.05, by One pattern t-test. C Immunoblot evaluation of AML Cas9 cell strains transduced with sgRNA constructs concentrating on FBXL6 or sgCtrl. Cells had been harvested on the indicated time factors post-infection and whole-cell extracts ready underneath denaturing lysis circumstances. D Cells from C and two further organic replicates had been counted utilizing the trypan blue exclusion technique on an automated cell counter (Vi-Cell Blu, Beckman Coulter) on the indicated time factors. E Cells from C and two further organic replicates had been subjected to MTS viability assays utilizing the CellTiter 96® Aqueous One Resolution on the indicated time factors. Leads to D and E are normalized and introduced in relation to sgCtrl. ***P < 0.001; **P < 0.01 ; *P < 0.05, by One pattern t-test. F Differential gene expression evaluation of 1041 Ubiquitin (Ub)-related genes within the MLL AML affected person cohort (n = 762) in comparison with wholesome bone marrow (BM) controls (n = 64). Log2FC (fold change) and the related p-value are proven. G Particular person values from F for FBXL6 mRNA expression in AML sufferers and wholesome BM. CPM counts per million reads mapped. ****P < 0.0001, by Pupil’s t check. H FBXL6 mRNA expression throughout AML and different most cancers cell strains derived from the DepMap public 23Q4 dataset. TPM transcripts per million reads mapped. ***P < 0.001 by Pupil’s t check.
To cross validate FBXL6 as a possible dependency in AML, we analyzed RNAseq knowledge of a big AML affected person cohort encompassing greater than 700 instances [11]. Throughout the over 1,000 ubiquitin-related genes investigated, FBXL6 was considered one of most extremely overexpressed genes (Fig. 1F). Importantly, throughout all AML subtypes in accordance with the WHO classification, over 90% of AML instances displayed greater FBXL6 mRNA ranges than wholesome management bone marrow (Fig. 1G, Supplementary Fig. 1B). Likewise, evaluation of the RNAseq Expression Public 23Q4 dataset from DepMap, protecting roughly 1400 totally different most cancers cell strains, revealed notably excessive FBXL6 mRNA expression ranges in AML cell strains as in comparison with different most cancers cell strains (Fig. 1H). Furthermore, evaluation of the BEAT AML 2.0 cohort [12] revealed excessive FBXL6 mRNA expression ranges to correlate with antagonistic prognosis in accordance with the ELN pointers [2] in additional help of an oncogenic position (Supplementary Fig. 1C).
When continuing with the purposeful characterization of FBXL6, immunoblot analyses revealed a second type of FBXL6 working at a decrease molecular weight (MW) of roughly 54 kDa (FBXL6-low_MW) subsequent to the wild sort (WT) kind on the predicted molecular weight of 59 kDa (FBXL6-WT) (Fig. 2A, B). Importantly, FBXL6-low_MW was recognized solely in AML cell strains, notably of the M4 and M5 subtypes (in accordance with the FAB classification) and never some other of the investigated hematological or strong most cancers entities (Fig. 2A, B). FBXL6-low_MW was additionally present in AML patient-derived xenograft (PDX) cells (Fig. 2C). Knockdown experiments confirmed the specificity of this extra band (Supplementary Fig. 2A). Strikingly, FBXL6-low_MW was depleted upon TPA-induced myeloid differentiation whereas FBXL6-WT was enriched (Fig. 2D, Supplementary Fig. 2B–D). This shift to FBXL6-WT was additionally noticed upon ATRA-induced differentiation (Supplementary Fig. 2E), suggesting a possible position of FBXL6-low_MW in sustaining the undifferentiated state in AML blasts. As no described isoforms of FBXL6 match the molecular weight of FBXL6-low_MW, we speculated that the decrease kind may end result from proteolytic cleavage. To check this speculation, we cloned N- or C-terminally FLAG-tagged FBXL6 constructs and expressed them in AML cells. Apparently, FBXL6-low_MW was solely detectable by anti-FLAG antibodies upon C-terminal and never N-terminal tagging (Fig. 2E). This discovering was strongly suggestive of proteolytic cleavage within the N-terminal area of FBXL6, leading to a small N-terminal fragment not detectable in SDS-PAGE (Fig. 2E). Subsequent, we carried out in-vitro-cleavage assays based mostly on extended incubation of lysates in presence of normal protease inhibitors or an expanded inhibitor cocktail moreover containing the protease inhibitors AEBSF, bestatin, E-64 and pepstatin. Diminished cleavage upon therapy with the expanded inhibitor cocktail additional confirmed proteolytic processing of FBXL6 because the supply for FBXL6-low_MW (Fig. 2F). Separate testing of the inhibitors recognized AEBSF, an irreversible serine protease inhibitor, as the one inhibitor abrogating FBXL6-processing in vitro (Supplementary Fig. 3A, B), pointing towards cleavage by a protease of the serine-type. To determine the precise cleavage web site, a twin proteomic method combining top-down and bottom-up proteomics was utilized (Supplementary Fig. 4A). The underside-up method, which measures the molecular weight of peptides ensuing from trypsin digest, revealed the lack of peptides amongst amino acid (aa) 21 and aa 58 within the samples akin to FBXL6-low_MW (Supplementary Fig. 4B, C). High-down proteomics, which determines the mass of the intact protein, resulted within the detection of three totally different fragments corresponding nearly completely to the calculated molecular weight of cleavage merchandise beginning at Val48, Leu49 and Ser50 of FBXL6, thus mapping the cleavage web site to aa 47-50 (Fig. 2G, Supplementary Fig. 4D, E). With a purpose to determine the accountable protease, mass spectrometry-based interactome screening for FBXL6 interplay companions was carried out subsequent (Fig. 2H). Cross-validation of the FBXL6 interactome with an inventory of serine-proteases with an analogous cleavage motif extracted from the MEROPS database [13] revealed cathepsin G (CatG) as essentially the most promising candidate (Supplementary Fig. 5A). Certainly, CatG was discovered to work together with FBXL6-WT however not with FBXL6∆N, a mutant of FBXL6 missing the primary 47 aa to characterize FBXL6-low_MW (Fig. 2I). Notably, a CatG-specific inhibitor effectively abrogated in vitro FBXL6-processing, whereas an inhibitor for neutrophil elastase (NE), one other myeloid serine-protease, had no impact (Fig. 2J). Furthermore, we discovered that FBXL6 cleavage correlated with CatG expression throughout the cell line and PDX panels described in Fig. 2A–C (Supplementary Fig. 5B–D). CatG ranges additionally correlated with adjustments in FBXL6 cleavage upon TPA or ATRA-induced differentiation (Supplementary Fig. 5E, F), confirming CatG because the accountable protease. Literature interrogation on CatG nevertheless made us conscious of chosen observations, during which aberrant proteolytic exercise of CatG in direction of particular substrates was described as an artifact in commonplace lysis buffers with out having a correlate in dwelling cells [14, 15]. We subsequently examined FBXL6 cleavage underneath denaturing circumstances or in commonplace lysis buffer supplemented with extreme quantities of CatG inhibitor. Unexpectedly, cleaved FBXL6 was utterly absent underneath these circumstances (Fig. 2K), arguing towards a physiological existence of cleaved FBXL6.
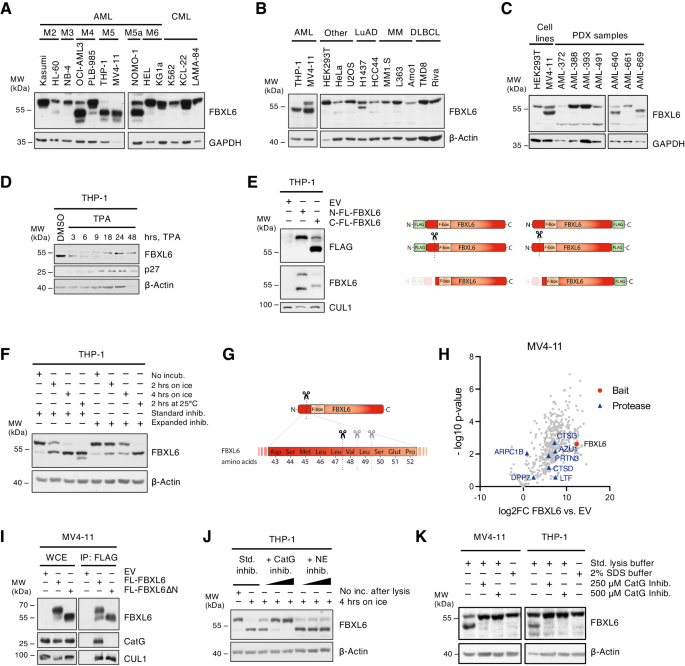
Immunoblot analyses of whole-cell extracts (WCE) ready underneath commonplace lysis circumstances from cell strains of (A) totally different AML subtypes and chosen CML (continual myeloid leukemia) cell strains, and (B) varied different most cancers entities together with lung adenocarcinoma (LuAD), a number of myeloma (MM) and diffuse massive B-cell lymphoma (DLBCL). C Immunoblot analyses of whole-cell extracts (WCE) ready underneath commonplace lysis circumstances from cell strains and totally different AML patient-derived xenograft strains. D Immunoblot evaluation of THP-1 cells handled with 25 ng/ml TPA or DMSO management for the indicated timepoints. E Immunoblot evaluation of THP-1 cells lentivirally transduced to ectopically specific N- or C-terminally FLAG-tagged FBXL6 (N-FL-FBXL6, C-FL-FBXL6) or empty vector (EV) management. WCE of contaminated cells had been ready underneath commonplace lysis circumstances. Proper facet: scheme to visualise the tagged kinds and ensuing cleavage fragments of FBXL6. F In-vitro-cleavage assay utilizing THP-1 lysates which had been incubated on ice or at 25 °C for the indicated durations of time or denatured straight by addition of Laemmli buffer. Normal inhibitor cocktail incorporates aprotinin, leupeptin, soybean trypsin inhibitor, PMSF, TPCK, TLCK; expanded inhibitor cocktail moreover contains AEBSF, bestatin, E-64, and pepstatin. G Scheme of recognized cleavage websites within the N-terminal area of FBXL6 (see additionally Supplementary Fig. 4) H Outcomes from mass spectrometry-based screening for interplay companions of FLAG-purified FBXL6. Log2FC values of co‐immunoprecipitated proteins in FBXL6 versus management samples are plotted towards the p-value. FBXL6 (bait) is depicted in pink and recognized proteases within the interactome are marked with blue triangles. I FLAG-IP of FLAG-tagged full-length FBXL6 and FBXL6∆N, a fraction beginning at Val48 to characterize the cleaved type of FBXL6, with subsequent immunoblot evaluation. J In-vitro-cleavage assay of THP-1 WCE incubated with particular inhibitors for Cathepsin G (CatG) and neutrophil elastase (NE) at rising concentrations (2.5 µM, 25 µM or 250 µM) along with the usual inhibitor cocktail (Std. inhib.). Inc, incubation. Okay Immunoblot evaluation of WCE ready underneath commonplace (Std. lysis buffer) or denaturing circumstances (SDS buffer) from MV4-11 or THP-1 pellets of equal dimension. The place indicated, extreme quantities of CatG-specific inhibitor had been added to the usual lysis buffer.
Taken collectively, we right here nominate the ubiquitin ligase FBXL6 as a possible novel vulnerability in AML, thus offering additional proof for the vital position of PTMs and particularly the ubiquitin system as a possible therapeutic goal in AML. Furthermore, we create consciousness in direction of particular cleavage phenomena simulating organic relevance however finally representing artifacts attributable to excessive protease exercise which isn’t sufficiently coated by commonplace protease inhibitor cocktails. This seems to be an experimental pitfall particularly in lysates of myeloid cells. Data of those phenomena helps to plan in accordance experiments underneath particular precautions, together with applicable inhibitors and controls to stop such artifacts.