Clonal variety performs a key function in poor therapeutic outcomes in acute myeloid leukemia (AML) and myeloproliferative neoplasm (MPN) sufferers [1, 2]. Subsequently, concentrating on all clones is required to eradicate these ailments. To attain this aim, we built-in genetic clonal panorama of particular person affected person samples with the response to DNA double-strand break restore (DSBR) inhibitors to trace particular person clones’ sensitivity to those medication. We determined to focus on DSBR pathways as a result of AML and MPN accumulate excessive numbers of DSBs, essentially the most deadly of all DNA lesions ensuing from altered metabolism [3, 4]. Thus, survival and proliferation of AML and MPN cells depend upon DSBR mechanisms which signify a promising targetable vulnerability (Supplementary Fig. S1) [5].
We reported that AML/MPN – related somatic mutations, e.g., FLT3(ITD) and JAK2(V617F) accompanied by TET2mut and/or DNMT3Amut may be utilized to trace clonal sensitivity to PARP and Polθ inhibitors [4, 6]. Thus, AML and MPN clones might reply otherwise to DSBR inhibitors relying on their mutational profile, and the remedy ought to be tailor-made to the mutation units carried by malignant clones.
To combine clonal composition with response to DSBR inhibitors we developed a single-cell focused DNA sequencing (sctDNA-seq) myeloid panel which illustrated clone-by-clone response to DSBR inhibitors and unraveled the clonal panorama of AML and MPN at a single-cell decision earlier than and after the remedy (Supplementary Fig. S2). We used AML and MPN somatic mutations as trackable clonal biomarkers to design a patient-tailored therapeutic routine (“clonal assault”) using DSBR inhibitors concurrently concentrating on all malignant clones.
Incubation of Lin-CD34 + cells from AML-MD2 affected person with six DNA harm response (DDR) inhibitors had been total delicate to PARPi, ATMi, ATRi and RAD52i, however immune to Polθi and DNA-PKi (Fig. 1A). sctDNA-seq adopted by phylogenetic tree evaluation revealed branched multi-clonal structure with eight clones carrying particular units of mutations (Fig. 1B-left). Fish plot evaluation demonstrated staggering variations between the sensitivity of varied clones to ATRi and to PARPi, ATMi and RAD52i (Fig. 1B-right, C). For instance, three clones carrying EZH2(V679M), EZH2(V679M) + TET2(L1721W) and EZH2(V679M) + TET2(L1721W) + FLT3(D835Y) had been immune to ATRi ( ~ 8%, 61% and 26% survivors, respectively) however extra delicate to PARPi ( ~ 4%, 33% and 12% survivors, respectively) and ATMi (~3%, 39% and 15% survivors, respectively). Conversely, three different clones carrying EZH2(V679M) + TET2(L1721W) + RUNX1(D160Y), EZH2(V679M) + TET2(L1721W) + RUNX1(D160Y) + EZH2(E54*), and EZH2(V679M) + TET2(L1721W) + RUNX1(D160Y) + EZH2(E54*) + BCOR1(R1334Tfs*32) + NRAS(G13R) had been delicate to ATRi (~1%, 2% and 0.5% survivors, respectively) whereas much less conscious of PARPi (~4%, 44% and 1% survivors, respectively) and ATMi (~2%, 38% and 1% survivors, respectively).
Lin-CD34+ AML affected person cells had been handled with PARPi olaparib, RAD52i 6-hydroxy-DL-dopa, ATMi KU-60019, ATRi VE-821 and DNA-PKi NU7026 for six days following cell survival evaluation and sctDNA-seq. A Sensitivity to the indicated concentrations of the inhibitors. Outcomes signify imply % ± SD of dwelling cells in comparison with vehicle-treated management. B Left – The phylogenetic tree visualizes the expected clonal construction based mostly on sctDNA-seq information. Proper – The proportion of clones with a special mixture of mutations after the remedy with the red-marked concentrations of the inhibitors indicated in panel A. C The fish plots mirror variety of cells earlier than (0 days) and 6 days after the remedy and the inferred clonal evolution sample based mostly on sctDNA-seq information. Chi-Sq. goodness of match p-value = 0.01578 exhibiting treatment-induced clonal variety. D Sensitivity of Lin-CD34+ AML-MD2 cells and regular counterparts to 2.5 μM olaparib, 30 μM 6-hydroxy-DL-dopa, 0.2 μM VE-821, 25 μM ART558, 2.5 μM KU-60019, and the indicated mixtures. Outcomes signify imply variety of colonies ± SD; ***p < 0.001 and **p < 0.01 in comparison with different teams utilizing one-way Anova. E Imply % of γH2AX-positive cells in Ki67+ cells ± SD; ***p < 0.001 and *p < 0.05 in comparison with corresponding particular person inhibitors teams utilizing one-way Anova and ###p < 0.001, ##p < 0.01 and #p < 0.05 in comparison with car utilizing one-way Anova.
Primarily based on this commentary, we hypothesized that simultaneous remedy with ATRi + PARPi or ATRi + ATMi ought to end in elimination of quite a few, if not all AML-MD2 clones. In settlement with this speculation, these drug mixtures had been 20–30× simpler in eliminating clonogenic progress of AML-MD2 cells when in comparison with particular person inhibitors (Fig. 1D). Remarkably, mixtures of those inhibitors (ATRi + PARPi and ATRi + ATMi) had been solely modestly poisonous to regular hematopoietic cells (Fig. 1D) bolstering their therapeutic potential. Furthermore, mixture of the inhibitors displaying comparable clonal concentrating on specificity reminiscent of PARPi + RAD52i was solely 2× higher than particular person inhibitors in opposition to clonogenic exercise of AML-MD2 cells. The impact would possibly end result from the induction of intracellular twin artificial lethality [7].
As anticipated, AML cells collected roughly 3× extra spontaneous DSBs detected by γH2AX immunofluorescence when in comparison with regular counterparts (Fig. 1E, Automobile). Solely the DSBR inhibitors which lowered survival of AML-MD2 cells (PARPi, ATRi, ATMi, RAD52i) elevated the proportion of γH2AX-positive leukemia cells. Furthermore, the magnitude of sensitivity of leukemia cells to the mixtures of PARPi + ATRi and PARPi + RAD52i was proportional to detection of cells with DSBs (γH2AX+). A low variety of γH2AX + cells was detected in regular hematopoietic cell populations handled with these DSBR inhibitors.
PARPi and ATRi – mediated accumulation of γH2AX+ AML-MD2 cells and discount of cell survival was accompanied by induction of differentiation (elevated of CD14+/CD11b+, CD11c+, and HLA-DR+ dwelling cells) (Supplementary Fig. S3) which could contribute to sturdy anti-leukemia exercise of the mix [8].
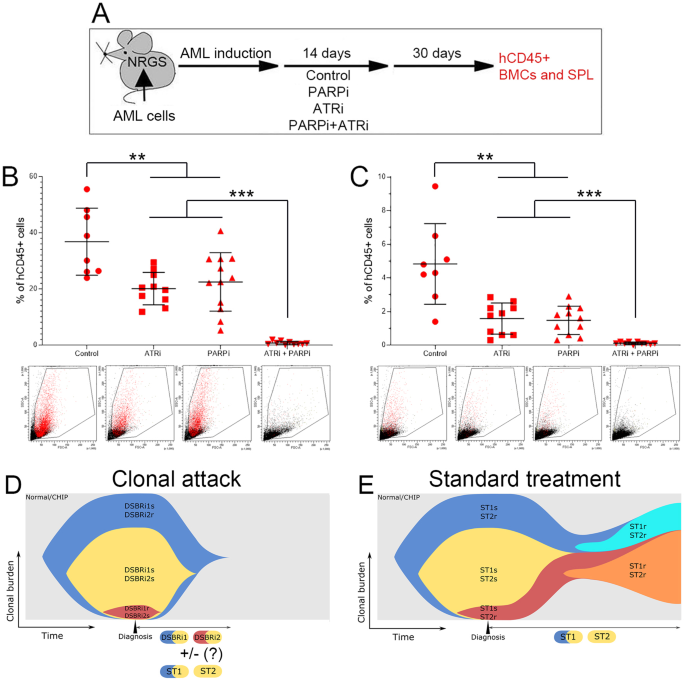
To find out if the “clonal assault” with DSBR inhibitors eradicates AML in vivo, we handled mice bearing AML-MD2 main leukemia xenografts with PARPi and/or ATRi (Fig. 2A). Particular person medication lowered the variety of hCD45+ AML-MD2 cells in bone marrow and spleen by roughly 2× and three×, respectively (Fig. 2B, C). Remarkably, the mix of PARPi +ATRi eradicated > 99% of leukemia cells in bone marrow and in spleen of seven/10 and 10/10 mice, respectively. No apparent toxicity in mice was attributed to remedy with PARPi + ATRi in concordance with different research exhibiting excessive efficacy and tolerable toxicity of low doses of ATRi + PARP1i [9].
A Experimental protocol. Humanized NRGS immunodeficient mice bearing main AML-MD2 xenograft had been handled for 14 consecutive days with car, PARPi olaparib (30 mg/kg i.p.), ATRi VE-822 (40 mg/kg oral gavage) and the mix of those medication. The effectivity of those medication was evaluated by detecting hCD45+ AML-MD2 cells in bone marrow and spleen one month after the top of the remedy to permit regrowth of cells that survived remedy. hCD45+ AML-MD2 affected person cells had been detected by immunofluorescence in bone marrow (B) and spleen (C). Outcomes signify imply variety of colonies ± SD; ***p < 0.001 and **p < 0.01 utilizing one-way Anova. D “Clonal assault” by DDR1 inhibitor + DDR2 inhibitor (e.g., PARPi + ATRi) finally mixed with normal remedy (ST) medication (ST1, ST2). E Normal remedy with ST1 drug + ST2 drug (e.g., quizartinib + azacytidine). Regular/CHIP regular hematopoiesis/clonal hematopoiesis of indeterminate potential.
Altogether, these outcomes spotlight the exceptional effectivity of a “clonal assault” with the DSBR inhibitors tailor-made to assault all AML clones in a affected person thus lowering the chance of improvement of time-dependent acquired resistance (Fig. 2D). As compared, normal remedy (Fig. 2E) typically employs chemotherapeutics with or with out tyrosine kinase inhibitor (TKi). Chemotherapy dosages are restricted by uncomfortable side effects and TKi requires extended remedy to successfully eradicate malignant clones. Thus, clones which can be inherently immune to the remedy, not detected by subsequent era sequencing, and/or these rising in the course of the extended remedy trigger the illness relapse. The potential combinatorial results of DSBR inhibitors and FDA accredited medication (e.g., tyrosine kinase inhibitors, and hypomethylating, genotoxic and pro-apoptotic brokers) have to be examined.
To check if clonal concentrating on by DSBR inhibitors might have broader applicability, extra affected person samples had been examined. Lin-CD34+ cells from MPN P349 affected person had been delicate to PARPi, ATMi, ATRi, RAD52i and DNA-PKi (Supplementary Fig. S4A). sctDNA-seq adopted by fish plot evaluation revealed similarities and variations between the sensitivity of the clones to those inhibitors (Supplementary Fig. S4B, C). For instance, the clone carrying KMT2A(L2373H) + SETBP1(H1100R) was extra delicate to ATRi (24% survivors) than RAD52i (59% survivors), conversely clones with KMT2A(L2373H) and KMT2A(L2373H) + SETBP1(H1100R) + FLT3(R834L) had been extra delicate to RAD52i (~40% and 1% survivors, respectively) than ATRi (72% and 4% survivors, respectively). Remarkably, the mix of RAD52i + ATRi was >100× simpler than particular person inhibitors in opposition to clonogenic progress of P349 cells (Supplementary Fig. S4D). Alternatively, mixture of RAD52i + ATMi, the 2 DSBR inhibitors displaying comparable sample of clonal concentrating on was solely 2x higher than particular person inhibitors.
Lin-CD34+ cells from MPN P350 affected person had been delicate to ATMi, ATRi, and RAD52i, and modestly delicate to PARPi (Supplementary Fig. S4E). Once more, sctDNA-seq adopted by fish plot evaluation revealed clonal similarities and variations in response to DSBR inhibitors (Supplementary Fig. S4F, G). Clone carrying TET2(P363L) + NRAS(G12D) was extra delicate to ATRi (~51% survivors) than PARPi (~ 79% survivors), whereas clones with TET2(P363L) + NRAS(G12D) + DNMT3A(W330C) and TET2(P363L) + NRAS(G12D) + DNMT3A(W330C) + IDH1(R132C) responded higher to PARPi (~0.1% and 0% survivors, respectively) than ATRi (~22% and three% survivors, respectively). The mixture of PARPi + ATRi was >9x simpler in inhibiting clonogenic progress of P349 cells, whereas RAD52i + ATMi displaying comparable sample of clonal concentrating on had been solely 2× higher than particular person inhibitors (Supplementary Fig. S4H).
Clonal concentrating on by DSBR inhibitors will not be relevant to all AML and MPN samples. For instance, Lin-CD34+ cells from AML V18 affected person had been delicate to ATMi, ATRi and DNA-PKi (Supplementary Fig. S5A) however no vital variations in main clones’ response had been detected (Supplementary Fig. S5B, C). As anticipated on this context, mixtures of ATMi + ATRi and ATMi + DNA-PKi didn’t exert stronger anti-leukemia impact than particular person inhibitors (Supplementary Fig. S5D).
Furthermore, Lin-CD34+ cells from AML V20 affected person pattern had been total immune to all examined DSBR inhibitors (Supplementary Fig. S5E) and mixtures of the inhibitors didn’t exert vital anti-leukemia impact when in comparison with particular person inhibitors (Supplementary Fig. S5F–H). AML V20 cells collected spontaneous DSBs (Supplementary Fig. S5I) due to this fact resistance to DSBR inhibitors was possible related to a DSB restore mechanism(s) not focused by these inhibitors.
The mechanistic features of clonal response to DSBR inhibitors are scarcely out there. For instance, the presence of DNMT3A(R882S) in AML V20 was related to resistance to DSBR inhibitors of just about all clones (Supplementary Fig. S5E–H). DNMT3A mutations (e.g., on the R882) are continuously detected in AML and had been related to resistance to anthracyclines [10]. Remarkably, acquisition of TET2(P562Qfs*6) reversed the resistant phenotype and rendered the clone to be delicate to a number of DSBR inhibitors. This commentary is supported by our report that whereas DNMT3Amut promoted resistance to PARPi in FLT3(ITD)-positive cells, DNMT3Amut;TET2mut;FLT3(ITD)-positive counterparts had been HR-deficient and extremely delicate to PARPi [6]. In conclusion, we postulate that acquisition of a mutation identified to modulate DSBR could also be a biomarker of the clonal response to DSBR inhibitor(s).
Altogether, “clonal assault” by the mixtures of DSBR inhibitors revealed exceptional effectivity in simultaneous eradication of malignant clones from a cohort of AML and MPN sufferers. This novel method which, as an alternative of utilizing genotoxic brokers to induce DNA harm, takes benefit of metabolic/replication stress-induced DSBs and targets clone-specific vulnerabilities in DSBR pathways. Whereas PARP inhibitors are broadly used to deal with sufferers with homologous recombination-deficient tumors together with AML [11], ATR, Polθ, ATM, DNA-PKcs kinase inhibitors have been evaluated in most cancers medical trials (NCT04991480) [12,13,14] and RAD52 inhibitor nonetheless awaits medical improvement. Furthermore, newly developed inhibitors ought to broaden the spectrum of AML and MPN clones delicate to DSBR inhibitors [15].