Identification and validation of recurrence-associated endothelial cells utilizing a 54-gene panel
As step one in exploring the prognostic worth of TECs in PCa, we estimated the abundance of TECs in main tumors by calculating the expression ranges of three traditional endothelial markers, together with PECAM1, ENG, and VWF, and investigated whether or not the TEC abundance is related to tumor recurrence. Within the TCGA-PRAD cohort, Kaplan–Meier analyzes demonstrated that tumors with increased expression ranges of those markers had considerably increased recurrence charges (all log-rank P < 0.05; Fig. 1a). This commentary is additional supported by Cox regression analyzes, as tumors with a better expression stage for every of the three genes have been at a considerably increased threat of creating recurrence (PECAM1 HR: 1.663, 95% CI: 1.098-2.518; ENG HR: 1.706, 95% CI: 1.126–2.584; VWF HR: 1.555, 95% CI: 1.023–2.363). A major affiliation of upper ENG or VWF stage with PCa recurrence was additionally detected within the DKFZ-PRAD cohort (Fig. S1). These outcomes steered that elevated TECs are related to tumor recurrence in main PCa.
a Greater mRNA ranges of PECAM1, ENG, and VWF are related to worse prognosis in prostate most cancers, as decided by the Kaplan–Meier evaluation within the TCGA-PRAD cohort (high tertile versus the low and median tertiles). P values have been calculated utilizing the log-rank take a look at. b UMAP plot of seven main cell varieties utilizing scRNA-seq information of 12 main prostate most cancers samples from the Chen examine [17]. c Dotplot of marker genes’ expression ranges of the most important cell varieties displayed in panel b. The scale of a dot signifies the share of cells that categorical corresponding genes, whereas the colour of a dot displays a gene’s expression stage. d Scissor algorithm-inferred endothelial cells (ECs) related to recurrence within the TCGA-PRAD cohort. Blue dots characterize recurrence-associated ECs (RAECs), related to a worse prognosis, whereas yellow dots characterize ECs related to a greater prognosis. Grey dots mark ECs that don’t present an affiliation with prognosis. e, f Vocalnol plots of differentially expressed genes between RAECs and non-RAECs (i.e., Scissor-ECs and background ECs) and people between ECs and non-ECs (i.e., different main cell varieties as proven in panel b) f Purple dots point out upregulated genes, whereas blue dots point out downregulated genes. g Venn diagrams of 54 RAEC-related genes, as recognized by their differential expression between RAECs and non-RAECs, increased expression ranges in ECs than non-ECs, and decrease expression ranges in non-ECs than ECs. h The next rating of the RAEC 54-gene panel (high tertile versus the remainder) is correlated with a shorter recurrence time within the TCGA-PRAD cohort as decided by Kaplan–Meier evaluation. The P worth was derived from the Log-rank take a look at.
TECs are extremely heterogeneous, so it’s essential to determine which subpopulation(s) of TECs is accountable for TEC’s affiliation with tumor recurrence in PCa. To deal with this query, we first analyzed the Chen scRNA-seq dataset, which comprises 12 untreated main PCa samples from radical prostatectomy. After high quality management, 29,457 cells have been analyzed, and all main cell varieties have been annotated primarily based on particular markers (Fig. 1b, c). A complete of 2457 ECs have been annotated in accordance with the expression of PECAM1, VWF, and ENG. We then leveraged the Scissor algorithm to combine these ECs’ transcriptomics to the TCGA-PRAD cohort and reliably recognized (P < 0.05) a subset of ECs (n = 347) whose expression profiles are related to PCa recurrence. This subpopulation was hereafter known as recurrence-associated ECs (RAECs) (Fig. 1d).
We then utilized differential gene analyzes to determine the genes whose expression profiles outline RAECs and thus can be utilized to estimate RAEC abundance in bulk sequencing datasets. In complete, 165 genes have been upregulated and 122 downregulated in RAECs in comparison with the remaining ECs (non-RAECs) (Fig. 1e; Desk S5); and 801 genes have been upregulated in ECs in comparison with non-ECs (Fig. 1f; Desk S6). Of these 801 genes, 163 have been upregulated in each RAECs and ECs. To reinforce the specificity of those genes to ECs, we used expression cutoffs to take away 109 genes that have been additionally extremely expressed in non-ECs, which left 54 RAEC-related genes (Fig. 1g).
Within the TCGA-PRAD cohort, expression ranges of the 54 RAEC-related genes have been summed, and Kaplan–Meier and Cox regression analyzes have been carried out. PCa with increased scores of the 54 genes confirmed a considerably worse disease-free survival (log-rank P = 0.008, HR: 1.741, 95% CI:1.150–2.635; Fig. 1h). These findings verify the robustness of the Scissor choice.
RAECs are attribute of tip ECs and elevated angiogenic actions
To additional characterize RAECs, we carried out sub-clustering and fantastic annotation of ECs (Fig. S2A, b) within the Chen dataset, wherein ECs have been labeled primarily based on purposeful states as an alternative of organic cell varieties [17]. We recognized 7 organic subsets of ECs, together with arterial outlined by FBLN5 and ENPP2; postcapillary vein (PCV) by ACKR1, and SELP; activated PCV by POSTN and CCL14; intermediate; immature by APLNR; and tip by ESM1 and APLN (Fig. S2C). Arteries, PCVs, and capillaries are completely different EC subtypes belonging to conventional vascular beds. Activated PCV is beforehand recognized in lung most cancers and choroid neovascularization, and is taken into account to be the EC subtype from which neovessels originate [19]. Immature cells resemble stalk-like cells, which elongate vessel sprouts whereas Tip cells information and navigate vessel sprouts throughout neovascularization [18, 19]. Intermediate cells are thought of a plastic phenotype probably transitioning from activated PCV to angiogenic cells [18, 19]. RAECs consisted of all annotated EC subtypes (Fig. S2D), tip cells have been most considerable (n = 78), adopted by immature cells (n = 60) and intermediate cells (n = 58). In comparison with non-RAECs, RAECs contained extra tip cells (22.5% vs. 14%), immature cells (17.3% vs. 8.7%), and intermediate cells (16.7% vs. 5.8%, Fig. S2E).
We then evaluated whether or not completely different EC subtypes have distinct gene expression profiles between RAECs and non-RAECs. Among the many 5 subtypes of ECs, tip cells had probably the most DEGs (Fig. S2F). Particularly, tip cells in RAECs expressed increased ranges of gene signatures related to tip cell markers, migration, and extracellular matrix (ECM) modeling than their counterparts in non-RAECs (Fig. S2G). This discovering suggests {that a} extra differentiated state of tip cells performs crucial function in RAEC-associated PCa recurrence. GSEA demonstrated that RAECs expressed increased ranges of tip cell markers, collagen, and VEGFRs (Fig. S2H) and have been enriched in angiogenesis, migratory, and ECM modeling pathways in comparison with non-RAECs (Fig. S2I). For potential regulators of RAECs, the SCENIC evaluation demonstrated that SOX4 and ZEB1 have been probably the most particular regulons of RAECs (Fig. S2J). Persistently, in comparison with different subtypes of RAECs, tip cells had the best mRNA ranges of SOX4 and ZEB1 (Fig. S2F). Taken collectively, RAECs primarily comprise tip cells and immature cells, and such tip cells are extremely differentiated with pronounced angiogenic and ECM modeling actions.
Improvement of a sturdy 18-gene recurrence-associated endothelial cell signature (RACEsig)
Since RAECs have been prognostic for PCa recurrence, we sought to assemble a sturdy gene signature that represents RAECs and might predict tumor recurrence. Univariate Cox regression evaluation demonstrated that 18 of the 54 RAEC-specific genes have been related to disease-free survival (P < 0.1) within the TCGA-PRAD cohort (Fig. 2a and Desk S7). In step with the RAEC traits described above, marker genes of tip cells (ESM1, FSCN1) and immature cells (APLNR) have been among the many 18 genes.
a Forest plot of 18 prognostic genes recognized by univariate Cox regression evaluation of 54 RAEC-related genes offered in Fig. 1g. P < 0.1 for all 18 genes. b Warmth map of relative mRNA ranges of the 18 prognostic RAEC-related genes throughout main cell varieties within the Chen dataset. c RAECs had the best rating when the 18 genes’ z-scores have been summed for several types of cells. d Warmth map of the 18 genes’ mRNA expression sample in TCGA-PRAD. e C-index and IBS in outer 10-fold validation utilizing the 11 hyperparameter-tuned fashions. Dots point out every mannequin’s imply of 10 C-indices or built-in Brier scores (IBS). The mannequin with the best common C-index and lowest common IBS was chosen and termed recurrence-associated endothelial cell signature (RAECsig). f Bar plot displaying function significance of the 18 prognostic RAEC-related genes inferred by the eXtreme Gradient Boosting (XGBoost). Higher significance suggests extra contributions to the XGBoost mannequin when predicting PCa development. g RAECsig scores amongst main cell varieties within the Chen dataset. SVM, help vector machine; Enet, elastic community; Lasso, Least Absolute Shrinkage and Choice Operator; plsRcox, partial least squares regression for Cox; StepCox, stepwise Cox regression; GBM, generalized boosted regression modeling; RSF, random survival forest; SuperPC, supervised principal parts.
Every of the 18 RAEC genes was extremely expressed in ECs, with 6 genes downregulated and 12 upregulated in RAECs (Fig. 2b). The z-scores for these 18 genes have been summed throughout different main cell varieties, and the sum scores have been the best within the RAECs (P < 0.001, Fig. 2c), additional indicating the facility of those genes in distinguishing RAECs from different sorts of cells. The expression patterns of the 18 RAEC genes have been confirmed within the Ge dataset (Fig. S3A), wherein virtually all RAEC genes confirmed increased mRNA ranges in ECs than non-ECs, and the 18 genes’ scores have been the best in ECs (Fig. S3B, C).
Importantly, these prognostic genes derived from scRNA-seq information might be utilized within the bulk RNA-seq information. By measuring gene coexpression and hierarchically clustering these genes’ coexpression patterns, we discovered that the mRNA expression ranges of 12 upregulated RAEC genes have been extremely correlated with however separated from these of the 6 downregulated RAEC genes (Fig. 2nd and S4). Subsequently, the scRNA-seq-derived gene markers might be leveraged to deduce RAEC’s abundance in bulk RNA-seq information.
Primarily based on the 18 genes, we benchmarked 11 survival-related machine-learning algorithms by nested CV within the TCGA-PRAD. As proven in Fig. 2e and Desk S8, the XGBoost survival mannequin achieved the perfect performances with the best imply C-index (0.692) and lowest imply IBS (0.151). This mannequin with tuned hyper parameters was then fitted on the complete TCGA-PRAD dataset and termed RAECsig hereafter. The function significance of the 18 genes is proven in Fig. 2f, the place the highest 5 options included FAM107A, FSCN1, TMEM255B, GABRD, and DOCK6. Utilizing the RAECsig, we then calculated threat scores of various cell varieties within the Chen dataset and located that RAECs ranked first amongst all main cell varieties (P < 0.001, Fig. 2g), demonstrating that the RAECsig is indicative of RAECs. Moreover, the RAECsig rating was the best in tip cells of RAECs (Fig. S5A) and will considerably discriminate them from different ECs (AUC = 0.844; Fig. S5B).
RAECsig is an impartial threat issue for main PCa recurrence
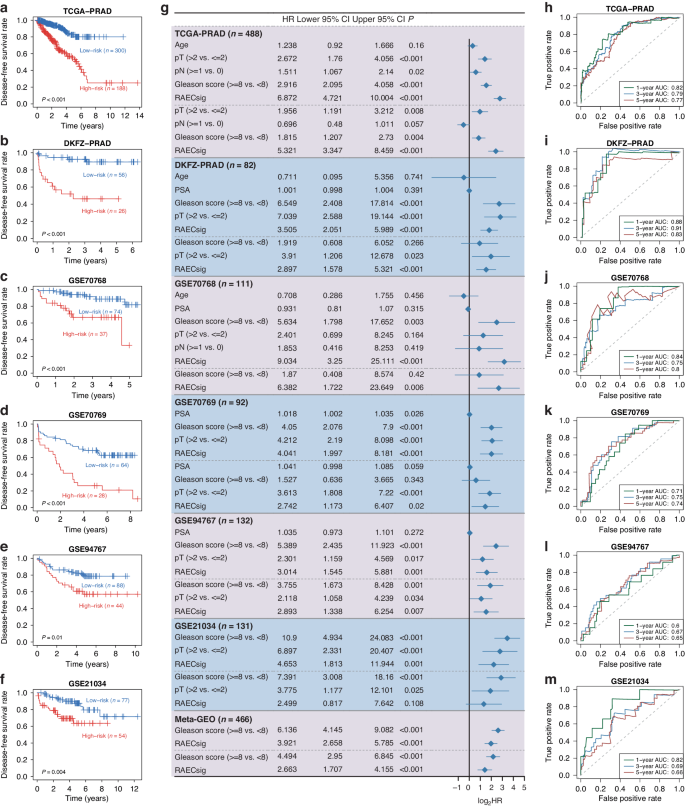
The prognostic worth of the RAECsig was validated utilizing 5 bulk sequencing datasets of main PCa. The RAECsig threat rating was calculated for every case, and the “survminer” package deal was used to outline the optimum threat rating threshold throughout varied datasets. Utilizing the lower off 0.58 derived from the GSE21034 cohort, we assigned sufferers to the high- and low-risk teams in every dataset. The next RAECsig rating was considerably related to shorter disease-free survival in every cohort (log-rank P ≤ 0.01, Fig. 3a–f and S6A). Univariate Cox regression analyzes confirmed that the RAECsig was a threat issue for PCa recurrence in all cohorts (Fig. 3g). After adjusting for age, Gleason rating, serum PSA, and TNM stage, RAECsig remained a statistically vital prognostic issue within the TCGA-PRAD (HR: 5.321, 95% CI: 3.347–8.459, P < 0.001), DKFZ-PRAD (HR: 2.897, 95% CI: 1.578–5.321, P < 0.001), GSE70768 (HR: 6.382, 95% CI: 1.722-23.649, P = 0.006), GSE70769 (HR: 2.742, 95% CI: 1.173–6.407, P = 0.02), GSE94767 (HR: 2.893, 95% CI: 1.338–6.254, P = 0.007), and Meta-Cohort (HR: 2.663, 95% CI: 1.707–4.155, P < 0.001) besides in GSE21034 (HR: 2.499, 95% CI: 0.817–7.642, P = 0.108). Moreover, time-dependent ROC evaluation revealed a sturdy discrimination energy of RAECsig in PCa recurrence. Particularly, the AUC values at 1, 3, and 5 years have been 0.82, 0.79, and 0.77 in TCGA-PRAD; 0.88, 0.91, and 0.83 in DKFZ-PRAD; 0.84, 0.75, and 0.80 in GSE70768; 0.71, 0.75, and 0.74 in GSE70769; 0.60, 0.67, and 0.65 in GSE94767; 0.82, 0.69, and 0.66 in GSE21034; and 0.70, 0.70, and 0.68 in Meta-cohort (Fig. 3h–m and S6B). Taken collectively, we discovered {that a} common RAECsig lower off can classify sufferers into both high- or low-risk teams and that RAECsig might be an impartial threat issue for PCa recurrence.
a–f Recurrence-free survival of high- and low-risk teams stratified by a universe RAECsig worth (0.58), together with TCGA-PRAD (a), DKFZ-PRAD (b), GSE70768 (c), GSE70769 (d), GSE94767 (e), and GSE21034 (f). P values have been derived from log-rank checks and have been < 0.05. g Forest plot displaying hazard ratio (HR) at 95% confidence interval (CI) and the corresponding P values of RAECsig and medical and pathological traits utilizing each the univariate (above the dashed strains) and the multivariate Cox regression analyzes (under the dashed strains) in 6 PCa cohorts. Solely variables with a P worth < 0.05 in univariate analyzes have been included in multivariate analyzes. The Meta-Cohort consists of 4 PCa cohorts (panels c–f) wherein gene expression was detected utilizing the microarray platform. h–m AUROC curve evaluation of RAECsig for predicting recurrence at 1, 3, and 5 years within the cohorts of TCGA-PRAD (h), DKFZ-PRAD (i), GSE70768(J), GSE70769(Okay), GSE94767(l), and GSE21034 (m). pT, pathological tumor stage; PSA, prostate-specific antigen; pN, pathological lymph node stage; AUROC, space below the receiver working attribute curve.
Comparability of RAECsig with clinicopathological options and industrial genetic assays
Clinicopathological variables comparable to age, Gleason rating, and TNM stage are generally used to evaluate PCa prognosis in medical apply. We, subsequently, used the C-index to check the predictive energy of RAECsig to these of frequent clinicopathological options throughout datasets (Desk S9). The C-index [95% CI] of RAECsig was calculated with 0.767 [0.714–0.819] in TCGA-PRAD, 0.836 [0.765–0.907] in DKFZ-PRAD, 0.769 [0.671–0.868] in GSE70768, 0.679 [0.608-0.750] in GSE70769, 0.641 [0.546–0.736] in GSE94767, and 0.669 [0.564–0.774] in GSE21034 (Fig. 4a–f). Total, RAECsig carried out higher than age, Gleason rating, and TNM stage in TCGA-PRAD and GSE70768 however was statistically non-inferior in different datasets.
a–f C-indices of the RAECsig, age, pathological tumor stage (pT), pathological lymph node stage (pN), prostate-specific antigen (PSA), and Gleason scores in numerous PCa cohorts, together with TCGA-PRAD (a), DKFZ-PRAD (b), GSE70768 (c), GSE70769 (d), GSE94767 (e), and GSE21034 (f). g Univariate Cox regression evaluation of RAECsig and printed fashions throughout PCa cohorts. h C-indices of the RAECsig and printed fashions in varied PCa cohorts. Information in (a–h) are offered as imply ± 95% confidence interval. *P < 0.05; **P < 0.01; ***P < 0.001; ****P < 0.0001.
We additionally in contrast RAECsig with OncotypeDX, Prolaris, and Decipher. There have been no overlapping genes between RAECsig and every of those 3 assays. We calculated OncotypeDX-like, Prolaris-like, and Decipher-like scores to samples in all datasets. Just like the RAECsig scores, the OncotypexDX-like, Prolaris-like, and Decipher-like scores confirmed constant constructive associations with recurrence in all datasets (Fig. 4g). Notably, the univariate HRs of RAECsig have been better than these from the three assays in all datasets. Moreover, we calculated C-indices for these signatures and located that RAECsig considerably outperformed the opposite three in TCGA-PRAD (OncotypeDX-like: 0.661 [0.592–0.729]; Prolaris-like: 0.654 [0.586–0.721]; Decipher-like: 0.588 [0.518–0.659]). In different datasets, RAECsig was akin to the opposite three assays (Fig. 4h; Desk S10). Moreover, we confirmed that RAECsig carried out higher than the endothelial signature in predicting PCa recurrence (Fig. S7). Altogether, RAECsig displayed strong predictive powers in comparison with medical options and gene panels in numerous cohorts and, subsequently, had higher extrapolation potential.
Key RAEC genes improve angiogenic exercise and PCa cell proliferation
Of the 12 upregulated RAEC-associated genes, FSCN1 was ranked the highest one by XGBoost (Fig. 2f), adopted by TMEM255B and GABRD. Moreover, these 3 genes’ expression ranges have been considerably related to PCa recurrence and better Gleason scores (Fig. S6). We thus selected FSCN1, TMEM255B, and GABRD to judge the RAECsig in PCa development. FSCN1 is a marker of tip cells and was considerably upregulated in ECs from samples with increased Gleason scores (Fig. 5a). The expressions of TMEM255B and GABRD in ECs have been additionally confirmed on the protein stage (Fig. S7).
a Consultant immunostaining micrographs of FSCN1 and CD31 in human PCa samples with increased and decrease Gleason scores. Nuclei have been stained with DAPI. Photos to the fitting are magnified boxed areas. Scale bars, 20 μm. Information are offered as means± s.e (n = 28, proven in the fitting panel). P worth was calculated by the Wilcoxon rank sum take a look at. b Gene set enrichment evaluation revealed high dysregulated HALLMARK pathways in PCa sufferers with increased RAECsig scores. c Validation of RNAi-mediated knockdown of FSCN1, TMEM255B, and GABRD in HUVECs, as measured by real-time qPCR. Information are offered as means ± s.d (n = 3). Statistical significance was decided utilizing two-tailed Scholar’s t-tests. d Tube formation of HUVECs was affected by silencing GABRD and TMEM255B. Information are offered as means ± s.d (n = 3, proven in proper panels). e Knockdown of GABRD by 2 siRNAs in HUVEC, as measured by real-time qPCR. Information are offered as means ± s.d (n = 3). f GABRD knockdown inhibited HUVEC migration and invasion, as decided by the transwell assay. Information are denoted as means ± s.d (n = 3). g GABRD and TMEM255B knockdown induced HUVEC apoptosis. Information are denoted as means ± s.d (n = 3). h The variety of PCa cells was decreased by conditioned media from HUVECs with the knockdown of FSCN1, TMEM255B, and GABRD, as decided by the CCK-8 assay. Information are proven as means ± s.d (n = 3). Statistical significance was decided by one-way ANOVA with Dunnett’s take a look at for (d–h). *P < 0.05; **P < 0.01; ***P < 0.001.
As a better RAECsig rating steered extra considerable RAECs, notably extra tip cells, we reasoned that PCa with increased RAECsig scores may manifest elevated angiogenic actions to boost tumor progress by supplying extra oxygen and vitamins. In step with this concept, PCa with increased RAECsig scores have been enriched in pathways of angiogenesis and cell cycle development primarily based on GSEA of consensus DEGs from the majority RNA-seq datasets (Fig. 5b and S8; Desk S11). Subsequently, we assessed whether or not FSCN1, TMEM255B, and GABRD functionally improve angiogenic actions. In HUVECs, silencing GABRD or TMEM255B considerably inhibited their tube formation, an indicator of angiogenesis (Fig. 5c, d). GABRD silencing was stronger than different genes within the tube formation assay. Persistently, GABRD silencing considerably diminished the migration and invasion of HUVECs (Fig. 5e, f). Additionally, we discovered that silencing GABRD or TMEM255B considerably induced apoptosis in HUVEC cells, which can additionally clarify the dramatic discount of their tube formation (Fig. 5g).
Accumulating proof signifies that ECs may regulate PCa development by way of interacting with tumor cells [44,45,46]. We thus used collected media from HUVECs with the silencing of FSCN1, TMEM255B, and GABRD to deal with human PCa cells and carried out cell proliferation evaluation. We discovered that silencing FSCN1, TMEM255B, or GABRD in HUVECs considerably inhibited the proliferation of PC-3 and 22Rv1 cells (Fig. 5h). These findings recommend that RAECs may additionally promote PCa development by instantly modulating tumor cell proliferation.
The RAECsig additionally predicts castration resistance
Cell cycle development (CCP) is inversely correlated with AR actions and might predict main abiraterone resistance in metastatic PCa [47]. We seen that the downregulation of the androgen response pathway was accompanied by the enrichment of cell cycle-related pathways in RAECsig-high tumors (Fig. 5b). We thus investigated whether or not RAECs additionally play a task in castration resistance in PCa. We calculated the CCP scores, AR actions, and CRPCsig51 scores for tumor samples within the TCGA-PRAD, DKFZ-PRAD, and Meta-Cohort and located that increased RAECsig scores have been considerably correlated with each CCP and CRPCsig51 scores whereas weakly correlated with unfavourable androgen actions (Fig. 6a–c and Fig. S9A, B). To additional take a look at whether or not a rise in RAECs is related to the event of castration resistance in PCa, we curated scRNA-seq information of ECs from each main PCa and mCRPC primarily based on the expression of PECAM1 and PROX1 (Fig. S10A–M). After eradicating samples with lower than 50 ECs (Fig. S10N) and correcting batch results (Fig. S11A–D), a complete of 16645 ECs from 28 samples have been used to generate a transcriptional atlas of ECs throughout PCa phases (Fig. 6d). All EC subtypes described above have been recognized primarily based on canonical markers (Fig. 6e and S10F–11E).
a–c Associations of upper RAECsig values with increased cell cycle development (CCP) scores (a), increased values of a CRPC signature (CRPCsig51, b), and decrease AR actions (c). The correlation coefficient R and corresponding P values have been decided utilizing Spearman’s rank correlation evaluation. d UMAP plot of ECs collected from 4 PCa scRNA-seq datasets of main PCa (blue) and metastatic CRPC (mCRPC, purple). e UMAP plot of EC subtypes. EC (LS) represents cells with decrease sequencing depth. f Featureplot displaying RAECsig scores of ECs. RAECsig scores have been calculated utilizing scaled information as enter for the RAECsig mannequin. g UMAP plot of ECs stratified by RAECsig scores. The lower off RAECsig worth for discretion was decided utilizing the Youden index that predicts tip cells from RAECs with maximized sensitivity and specificity. h, i Boxplot displaying proportions of RAECsig-high and RAECsig-low cells (h) and EC subtypes (i) in main PCa and mCRPC. Dots characterize the cell proportion from a pattern. Statistical significance was decided utilizing the Wilcoxon rank sum take a look at. ns, not vital; *, P < 0.05; **, P < 0.01; ***, P < 0.001; ****, P < 0.0001. (j) Kaplan–Meier evaluation displaying general survival with a first-line ARSI between RAECsig-high (purple) and RAECsig-low (blue) mCRPC. ok–n Boxplots displaying baseline RAECsig (ok), AR exercise (l), CCP rating (m), and CRPCsig51 rating (n) between enzalutamide responders and non-responders in main PCa. Statistical significance was decided utilizing the Wilcoxon rank sum take a look at. (o) ROC curves of RAECsig, AR exercise, CCP rating, and CRPCsig51 rating in predicting the likelihood of main resistance to enzalutamide within the GSE197780 dataset. CCP, cell cycle development; AR, androgen receptor; PCV, postcapillary vein; LS, decrease sequencing depth; mCRPC, metastatic castration-resistant prostate most cancers; ARSI, AR-signaling inhibitors; ROC, receiver working attribute.
We then calculated the RAECsig rating for every subtype of ECs. Once more, ECs with increased RAECsig scores have been principally tip cells (Fig. 6f). RAECsig-high ECs have been considerably enriched in mCRPC (Fig. 6g) whereas RAECsig-low ECs have been extra enriched in main tumors (Fig. 6h). In step with the enrichment of tip and immature cells in RAECs, each tip and immature cells have been additionally enriched in mCRPC (Fig. 6i). As well as, mCRPCs with increased RAECsig scores had considerably worse prognoses after remedies with first-line AR-signaling inhibitors (ARSI) (Fig. 6j).
We additionally examined whether or not increased RAECsig scores can predict main resistance to ARSI in main PCa utilizing the GSE197780 dataset, wherein sufferers underwent bulk RNA sequencing earlier than 3 months of neoadjuvant enzalutamide therapy [35]. Non-responders had considerably increased RAECsig scores than responders in these main tumors (Fig. 6k). Nonetheless, we didn’t detect vital variations within the main tumors’ AR actions, CCP scores, and CRPCsig51 scores (Fig. 6l–n). ROC evaluation confirmed that RAECsig may markedly discriminate non-responders from responders of enzalutamide (AUC = 0.704; Fig. 6o). These findings recommend that RAECsig can determine sufferers who could not reply to ADT.