CEBPA mutational standing
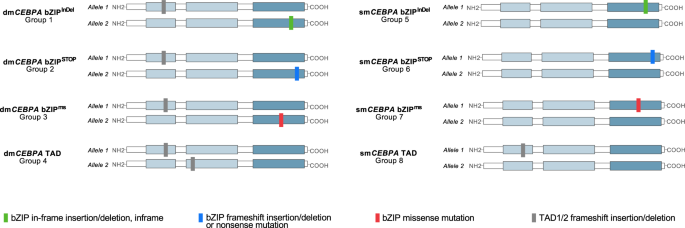
Of the 1010 sufferers, 661 sufferers (65.4%) confirmed mutations affecting the bZIP-domain of CEBPA encompassing amino acids (AA) 272–358 [1]. As illustrated in Fig. 1, mutations within the CEBPA bZIP area have been usually in-frame insertions or deletions (n = 491 pts; 74.3%; known as bZIPInDel), i.e. (multiples of) 3 bp affecting the DNA-binding-, fork- or bZIP-region. Frameshift insertions/deletions or nonsense mutations in bZIP, inflicting untimely termination of transcription (known as bZIPSTOP), in addition to missense mutations (bZIPms), inflicting single AA modifications, have been much less widespread and have been noticed in 81 (12.3%) and 89 (13.5%) of sufferers, respectively. Curiously, the several types of mutations confirmed a non-random distribution, with bZIPms primarily clustering in sure crucial AA positions within the DNA-binding primary area, i.e. AA297 and AA300. In distinction, most bZIPInDel mutations affected the hinge/fork area of the bZIP-domain, i.e., the 14 AAs previous the primary leucine residue of the leucine zipper at place 317 [21], with probably the most frequent InDel mutations at AA312 and AA313 (Fig. 1). In distinction to the bZIP area, the TAD domains virtually uniformly harbored frameshift InDel or nonsense mutations, making a untimely termination codon.
Mutations within the bZIP area (AA272-358) are usually in-frame insertion or deletion mutations (bZIPInDel), whereas frameshift insertions/deletions or nonsense mutations inflicting a untimely translational termination (bZIPSTOP), and single base-pair missense mutations (bZIPms) are much less widespread.
To evaluate the influence of various mutational constellations, 8 subgroups have been generated, making an allowance for each sort and website of mutation (bZIPInDel vs. bZIPms vs. bZIPSTOP vs. TAD) and allelic standing (dmCEBPA vs. smCEBPA). The attribute mutational constellations of the totally different mutational subgroups (dmCEBPA bZIPInDel (Gr1), dmCEBPA bZIPSTOP (Gr2), dmCEBPA bZIPms (Gr3), dmCEBPA TAD (Gr4), smCEBPA bZIPInDel (Gr5), smCEBPA bZIPSTOP (Gr6), smCEBPA bZIPms (Gr7) and smCEBPA TAD (Gr8)) are illustrated in Fig. 2.
On this cohort, sufferers with dmCEBPA predominantly harbored a mixture of a bZIP and a TAD mutation (n = 475, 87%). In uncommon instances (n = 11), a mixture of two bZIP mutations was current, in all of which one of many two mutations was a bZIPInDel mutation. These instances have been assigned to Gr1. 60 dmCEBPA sufferers (11%) confirmed alterations affecting solely the TAD areas, most ceaselessly being a mixture of TAD1 and TAD2 mutations (Gr4). In 10 sufferers with dmCEBPA, greater than two CEBPA mutations have been detected, together with the next mixtures: 2 bZIPInDel+TAD (assigned to Gr1); bZIPInDel+2 TAD (assigned to Gr1, n = 2), bZIPInDel + bZIPms+TAD (assigned to Gr1, n = 2), bZIPSTOP+2 TAD (assigned to Gr2); bZIPms+2 TAD (assigned to Gr3), 2 bZIPms+2 TAD (assigned to Gr3), 3 TAD (assigned to Gr4),
CEBPA mutational subgroups and scientific traits
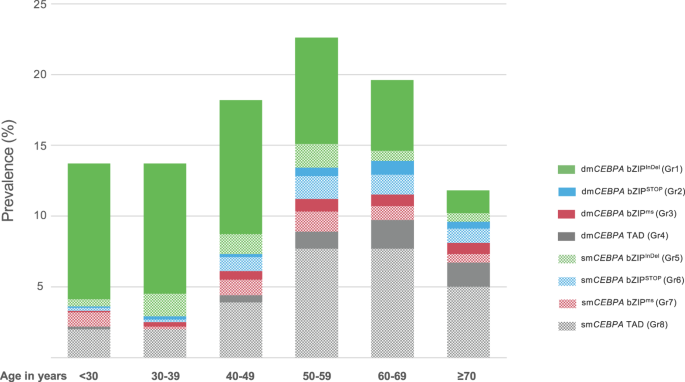
As outlined in Desk 1, the affiliation of a number of scientific parameters differed considerably between the outlined mutational subgroups. Sufferers with bZIPInDel mutations, i.e., Gr1 and Gr5, have been considerably youthful (median age Gr1 42.2 years [IQR 31–54.9]; Gr5 47 years [IQR 39–58]) than these with out bZIPInDel mutations (teams 2–4 and teams 6–8) (median age 52–64 years). Additionally they had a better prevalence of de novo AML (98% and 96%) in comparison with sufferers with out bZIPInDel mutations (teams 2–4 and teams 6–8), with the latter extra ceaselessly evolving as secondary illness after prior MDS or as tAML (charge of de novo AML 81–94%) (Desk 1). Categorizing age in 10-year intervals (Fig. 3, Desk S2), a steady lower within the incidence of bZIPInDel mutations (particularly dmCEBPA bZIPInDel) was seen with growing age, whereas bZIPSTOP and bZIPms mutations and alterations affecting solely the TAD areas have been significantly widespread in older people and fewer prevalent in sufferers as much as the age of 40 years. Different scientific parameters didn’t differ considerably between subgroups.
Affiliation of CEBPA mutations with different molecular and cytogenetic abnormalities
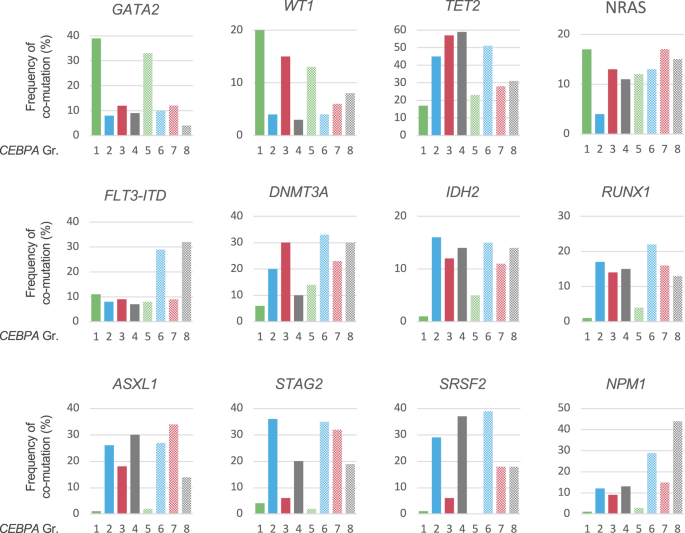
Further mutations have been recognized in 861/1010 CEBPA-mutant sufferers (85.2%). Determine 4 illustrates the distribution of co-mutations within the totally different CEBPA mutational subgroups. Important variations have been noticed for a number of genes, probably the most placing being GATA2, which was discovered mutated in 39 and 33% of sufferers in Gr1 and Gr5, however solely 4–12% in sufferers carrying non-bZIPInDel mutations (teams 2–4 and 6–8), and NPM1, with only one and three% of sufferers affected in Gr1 and Gr5, however 10–44% in teams 2–4 and 6–8 (p < 0.001). On the whole, the spectrum of co-mutations of sufferers with bZIPInDel mutations differed markedly from the opposite CEBPA subgroups, the latter extra ceaselessly carrying mutations in genes related to AML after prior MDS, comparable to spliceosome mutations (i.e. SRSF2, SF3B1, U2AF1 or ZRSR2) in addition to alterations related to DNA-methylation (i.e. DNMT3A, TET2, IDH1 and IDH2) (Fig. 4, Desk S3). Moreover GATA2, sufferers with bZIPInDel mutations have been extra more likely to harbor mutations in WT1 (Gr1 bZIPInDel 20%, Gr5 13%, all different teams 3–15%).
Most CEBPA-mutant sufferers confirmed a traditional karyotype (Desk 1). In sufferers with bZIPInDel mutations, the predominant cytogenetic aberrations have been del 9q (Gr1 n = 24/92; Gr5 n = 5/20) and +21 (Gr1 n = 19/92; Gr5 n = 4/20). Among the many different CEBPA mutational subgroups, chromosomal abnormalities have been extra various and included −7, −5, +8 and several other others, thereby reflecting the spectrum of modifications seen in AML normally.
CEBPA mutational subgroups and response to remedy
Therapy response knowledge have been accessible for 992 sufferers (98.2%). Final result evaluation confirmed comparability of CR charges in addition to RFS and OS between research teams (Desk S1). Concerning preliminary response to remedy inside the eight totally different subgroups, sufferers with bZIPInDel mutations achieved the very best charges of CR1, with 94.3% in Gr1 (OR 6.38 [3.83–10.63], p < 0.001) and 92.1% in Gr5 (OR 4.51 [1.71–11.86], p = 0.002) in comparison with CR-rates of 73.1–79.6% noticed within the different subgroups (Desk 1, Desk S4).
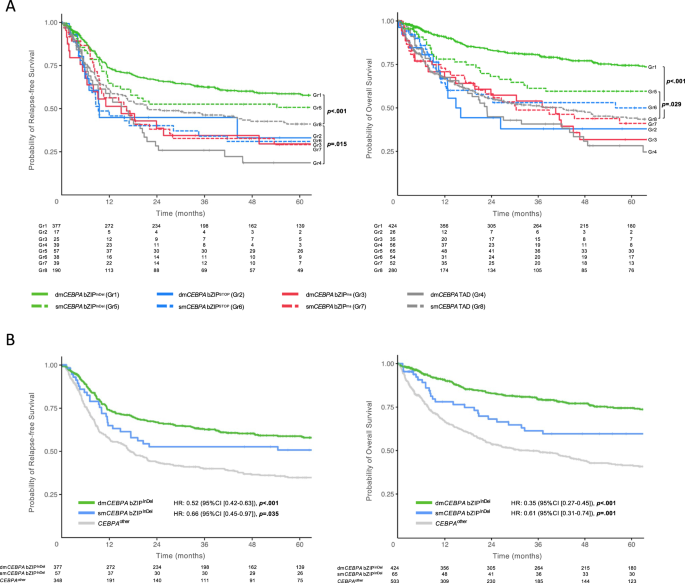
When analyzed in accordance with the eight totally different mutational subgroups, sufferers carrying bZIPInDel mutations had a extra favorable final result (RFS and OS) than sufferers with out these mutations (Fig. 5A). Intimately, sufferers in Gr1 confirmed the longest RFS (median 152 months (HR 0.60 [0.47–0.77]); p < 0.001) and OS (median 215 months, HR 0.37 [0.29–0.46], p < 0.001). Final result of Gr5 sufferers was much less favorable than Gr1 (median RFS 64 months (HR 0.77 [0.51–1.15]), p = 0.21; median OS 126 months HR = 0.65 [0.44 to 0.96], p = 0.029), however nonetheless higher than for the opposite subgroups (median RFS ranged between 9.4 and 21.8 months, median OS 15.7 and 70.9 months) (Tables S5 and S6).
In a subsequent evaluation combining teams 2–4 and 6–8 as CEBPAdifferent (reference group), sufferers with dmCEBPA bZIPInDel in addition to smCEBPA bZIPInDel each demonstrated considerably higher CR1, RFS and OS in contrast with CEBPAdifferent sufferers, which was confirmed in multivariate evaluation making an allowance for the person research teams and the affected person age (CR1: dmCEBPA bZIPInDel OR 5.82 [3.62–9.36], p < 0.001), smCEBPA bZIPInDel OR 4.13 [1.59–10.68], p = 0.003; RFS: dmCEBPA bZIPInDel HR 0.52 [0.42–0.63], p < 0.001, smCEBPA bZIPInDel HR 0.66 [0.45–0.97], p = 0.035; OS: dmCEBPA bZIPInDel HR 0.35 [0.28–0.43], p < 0.001, smCEBPA bZIPInDel HR 0.61 [0.42–0.89], p = 0.011 (Fig. 5B, Tables S7–S9).
AlloHCT carried out in CR1 confirmed no profit in bZIPInDel sufferers (HR 1.19 [0.81 to 1.75], p = 0.178) (Fig. S1, Desk S10).
Affect of co-mutations on final result in CEBPA bZIPInDel mutant sufferers
A number of research have reported on the impact of co-mutations [22,23,24,25,26,27,28] in CEBPA-mutant AML. In sufferers with dmCEBPA bZIPInDel, a possible concurrent impact on final result was assessed for the commonest alterations beforehand related to final result, i.e. GATA2, TET2, WT1, CSF3R and FLT3-ITD. The RFS within the few CSF3R-mutant sufferers (n = 15) was considerably lowered in univariate evaluation (dmCEBPA bZIPInDel /CSF3Rwt: RFS HR 0.24 [0.14–0.44], p < 0.001) (Fig. S2), which remained vital in multivariate evaluation (Desk S11), nonetheless, this impact didn’t translate right into a distinction in total survival (OS HR 1.42 [0.52–3.89]; p = 0.491). The presence of a co-mutation in TET2 was additionally related to worse RFS (dmCEBPA bZIPInDel /TET2wt: RFS HR 0.61 [0.4–0.95], p = 0.028) and OS, (dmCEBPA bZIPInDel /TET2wt: OS HR 0.56 [0.34–0.93], p = 0.025) (Fig. S3), although this impact misplaced its significance in multivariate evaluation (Desk S12). FLT3-ITD-positive sufferers had a shorter RFS in univariate (dmCEBPA bZIPInDel with out FLT3-ITD: RFS HR 0.60 [0.38–0.96], p = 0.031) (Fig. S4), however not in multivariate evaluation (Desk S13). In distinction, the presence of GATA2 and WT1 co-mutations didn’t considerably have an effect on RFS or OS on this group (Figs. S5, S6).
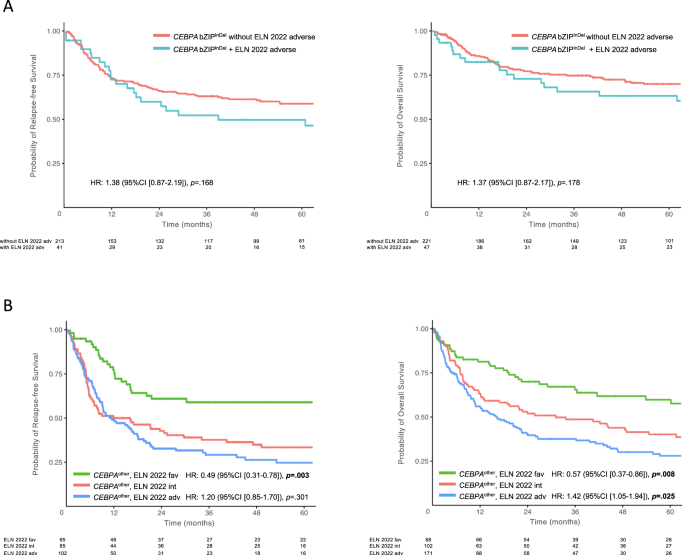
Affect of ELN2022 antagonistic molecular and cytogenetic alterations on final result in CEBPA bZIPInDel mutant AML
To analyze whether or not cytogenetic and molecular threat components in accordance with the novel ELN2022 suggestions had an impact on final result in bZIPInDel sufferers, additional final result evaluation was carried out inside this subgroup. In complete, 48 sufferers harbored a mixture of CEBPA bZIPInDel mutations and antagonistic cytogenetic or molecular abnormalities, together with advanced karyotypes (n = 9), monosomy 5 or 7 (n = 6) or mutations in ASXL1 (n = 5), EZH2 (n = 8), RUNX1 (n = 6), SF3B1 (n = 1), SRSF2 (n = 1), STAG2 (n = 8), TP53 (n = 2) and U2AF1 (n = 2). As depicted in Fig. 6A, the presence of ELN2022 antagonistic genetic components didn’t considerably have an effect on the favorable final result in CEBPA bZIPInDel-mutant sufferers (RFS: HR 1.37 [0.87–2.17], p = 0.178 and OS: HR 1.46 [0.9–2.37], p = 0.129) (Tables S14, S15).
Affect of ELN2022 mutational subgroups in CEBPA mutant sufferers with out bZIPInDel (subgroups 2–4 and 6–8)
Current evaluation steered that sure co-mutations, specifically mutant NPM1, would possibly have an impact in sufferers with smCEBPA [29]. To achieve additional insights on the influence of co-mutations in sufferers with out bZIPInDel mutations, i.e. teams 2–4 and 6–8 (CEBPAdifferent), a mixed evaluation based mostly on the ELN2022 pointers was carried out for these sufferers. A complete of 345 sufferers (63.8%) had ample cytogenetic and molecular knowledge to permit reclassification in accordance with the ELN2022 threat teams. Most of those sufferers (n = 183/345) (53%) have been assigned to the antagonistic threat group, predominantly attributable to ASXL1 (n = 85), RUNX1 (n = 63), SRSF2 (n = 56) and STAG2 (n = 55) mutations, whereas solely 27 sufferers (14.8%) had poor threat cytogenetics.
Within the sufferers re-assigned to the favorable threat group (n = 88), all however one affected person (displaying a t(8;21)) had NPM1-mutations. As depicted in Fig. 6B, the result of those teams confirmed statistically vital variations, with the median RFS not reached and a median OS of 154 months within the CEBPAdifferent/ELN2022 favorable threat group in comparison with 16 months and 31 months within the CEBPAdifferent/ELN2022 intermediate threat group and 12 months and 16 months within the CEBPAdifferent/ELN2022 antagonistic threat group (p < 0.001; for multivariate evaluation see Tables S16 and S17).