NOTCH4 is important for the survival of FLT3/ITD cells and its knockdown sensitizes FLT3/ITD AML cells to FLT3 TKI therapy
To seek for potential targets to mix with FLT3 TKI in treating FLT3/ITD AML, we carried out a loss-of-function “Achilles heel” RNAi display screen (Fig. 1A). On this display screen, we used a retroviral shRNA library (>12,500 constructs) which consists of small hairpin RNAs (shRNA) with 10–12 distinctive constructs for every of the >1000 genes related to the regulation of proliferation and survival of most cancers cells. Every particular person shRNA has a novel 60 bp barcode related to it. FLT3/ITD+ Molm14 cells had been pool contaminated with the retroviral RNAi library. Expression of the shRNA library was achieved by elimination of Tet repressor.
A Schematic diagram of RNAi survival and sensitization screening targets for FLT3 mutation remedy. The viral RNAi library contained distinctive shRNA barcodes for transfection of the FLT3/ITD+ Molm14. For RNAi survival screening, genomic DNA was extracted at 0, 5 and 21 days after shRNA transfection. For sensitization screening, FLT3 was inhibited by both knockdown with shRNA or by therapy with FLT3 inhibitors (10 nM lestaurtinib, 10 nM sorafenib or 25 nM midostaurin) adopted by genomic DNA extraction after 3 days. The barcodes had been then amplified by PCR, and the abundance of barcodes was decided by sequencing. Barcodes are largely depleted when (B) they’re poisonous to cells or (C) act synergistically with FLT3 inhibition. D Genes with excessive barcode fold depletion had been recognized as important genes for the survival of Molm14 cells upon FLT3 inhibition. E Western blot of NOTCH 1-4 intracellular area(ICD) in Molm14, MV4;11, THP-1 and HB11;19 cells and (F) quantification of protein expression in every cell line relative to Molm14 (normalized to ACTIN, n = 3 for every cell line). P-values for NOTCH4 expression is proven. G NOTCH4 mRNA expression (log2) in non-ITD (WT) and FLT3/ITD AML samples analyzed utilizing knowledge from dataset GSE9476. H NOTCH4 mRNA expression quantified by qRT-PCR in bone marrow (BM), Lin- and KSL cells from 12-week previous wild-type (WT, n = 3), Flt3/ITD heterozygous mutant (Flt3+/ITD, n = 3) and Flt3/ITD hemizygous mutant (Flt3-/ITD, n = 3) C57BL/6 mice (normalized to ribosomal protein S16 gene). I NOTCH1-4 mRNA expression quantified by qRT-PCR evaluation in MPP cells from BM of 12-week previous wild sort (WT), Flt3/KD and Flt3/ITD mutant C57BL/6 mice (n = 4 for every group). Information are introduced because the imply ± SD. *p < 0.05; ***p < 0.001.
For the survival display screen, barcodes had been depleted when expression of the corresponding shRNA exhibited cytotoxicity to Molm14 cells. The abundance of barcodes for consultant shRNA which demonstrated poisonous results on the cells, together with CYLD, ELK4, IL31RA, LYK5 and NOTCH4, had been all considerably diminished in a time-dependent method when in comparison with these earlier than induction (day 0) (Fig. 1B). For the sensitization display screen, upon doxycycline-induction of the shRNAs, we inhibited FLT3/ITD exercise within the shRNA library-transfected Molm14 cells with FLT3 inhibitors (10 nM lestaurtinib, 10 nM sorafenib or 25 nM midostaurin) or knocked down FLT3 expression by FLT3 shRNA induction through doxycycline addition to the tradition medium. The abundance of barcodes for the aforementioned consultant shRNAs had been enormously diminished as in comparison with the degrees with out FLT3 inhibition (Fig. 1C, D). These outcomes signify potential gene targets (and pathways) important for FLT3/ITD mutant cell survival, whose inhibition is poisonous to those cells and sensitizes them to FLT3 TKI therapy.
Since NOTCH4 has been reported to play roles in hematopoiesis, differentiation and the event of hematolymphoid neoplasms [16,17,18], we determined to deal with exploring NOTCH4 as a possible goal within the mixture examine. First, we investigated NOTCH4 expression in FLT3/ITD cell strains and affected person samples. In comparison with THP-1 (wild-type FLT3) or HB11;19 cells (FLT3 D835H+), expression of NOTCH4 was larger in Molm14 and MV4;11, two human FLT3/ITD AML cell strains (Fig. 1E, F, Supplementary Fig. S1A). Since expression of different members of NOTCH receptors have additionally been noticed in AML [19], we subsequent examined expression of the opposite NOTCH receptors in FLT3/ITD AML cells. In distinction, Molm14 and MV4;11 cells didn’t present considerably larger expression of NOTCH1, 2 or 3 in comparison with THP-1 or HB11;19 cells (Fig. 1E, F). Moreover, evaluation of information accessible within the Gene Expression Omnibus repository (GEO accession: GSE9476, GSE14479) [23, 24] demonstrated that main AML samples even have larger NOTCH4 expression than regular bone marrow (BM) or ALL samples (p < 0.001) (Supplementary Fig. S1B, C). Moreover, NOTCH4 expression was considerably larger in FLT3/ITD AML samples as in comparison with AML samples carrying wild-type (WT) FLT3 (p < 0.05) (Fig. 1G).
Subsequent, to grasp how varied FLT3 mutations impression NOTCH4 expression, we investigated expression of NOTCH4 within the genetically engineered mouse fashions with Flt3/ITD or Flt3/D838Y kinase area (Flt3/KD) mutations [25,26,27]. qRT-PCR evaluation confirmed that NOTCH4 expression was highest in Equipment+/Sca-1+/Lin– (KSL) cells in comparison with lineage unfavourable or complete BM, suggesting an affiliation between NOTCH4 expression and extra primitive hematopoietic cells. Expression of NOTCH4 within the KSL fraction was larger in each Flt3+/ITD (heterozygous) and Flt3-/ITD (hemizygous) mice in comparison with WT management (Fig. 1H). Expression of NOTCH4, however not the opposite NOTCH members of the family (NOTCH1, 2 or 3), in multipotent progenitor (MPP, Lin–Sca-1+CD117+CD34+CD135+) cells was considerably larger in Flt3/ITD mice in comparison with Flt3/KD or WT Flt3 mice (Fig. 1I). Greater expression of NOTCH4 in FLT3/ITD mouse strains is thus per the information seen for human FLT3/ITD AML cells. All these knowledge point out that NOTCH4 is upregulated in FLT3/ITD AML cells.
NOTCH4 knockdown sensitizes FLT3 TKI resistant FLT3/ITD mutant cells to TKI therapy
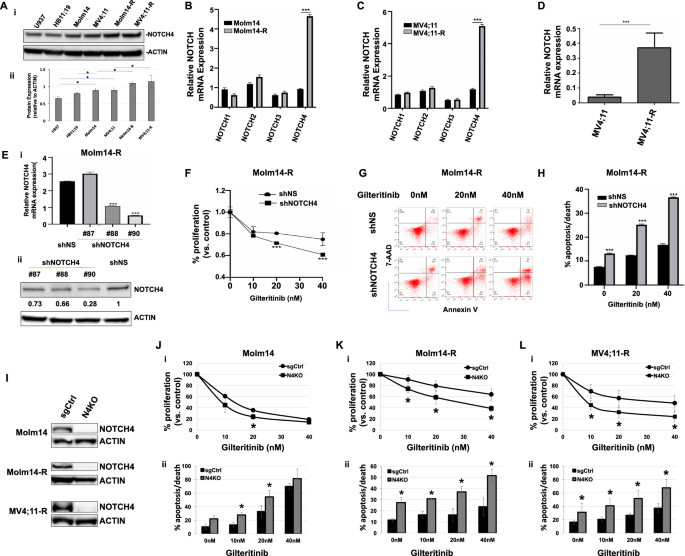
Since NOTCH4 is upregulated in FLT3/ITD AML cells, we subsequent investigated NOTCH4 expression and its impression on the proliferation and survival of TKI-resistant FLT3/ITD cells. Our outcomes confirmed that in comparison with HB11;19 cells or the FLT3-negative U937 cells, expression of NOTCH4 protein was larger in MV4;11 and Molm14 cells. Even larger ranges of NOTCH4 had been detected in TKI-resistant Molm14-R and MV4;11-R cells (Fig. 2A, Supplementary Fig. S1D). Additionally, qRT-PCR confirmed elevated expression of NOTCH4, however not NOTCH1-3 in Molm14-R and MV4;11-R cells, as in comparison with the parental Molm14 or MV4;11 cells, respectively (Fig. 2B, C). Persistently, elevated NOTCH4 expression was additionally noticed in a sorafenib-resistant MV4;11 cell line current in an unrelated GEO repository (GSE74666) (Fig. 2D) [23, 27]. These knowledge recommend that FLT3 TKI-resistant FLT3/ITD AML cells have additional elevated NOTCH4 expression.
A (i) NOTCH 1-4 expression in U937, HB11;19, Molm14, MV4;11, Molm14-R and MV4;11-R cells by Western blot and (ii) quantification of protein expression in every cell line (normalized to ACTIN, n = 4). NOTCH1–4 mRNA expression by qRT-PCR in Molm14/ Molm14-R (B) and MV4;11/ MV4;11-R cells (C). D NOTCH4 mRNA expression in MV4;11 and MV4;11-R (MV4;11 cells proof against 500 nM sorafenib) cells analyzed from the GEO repository GSE74666. E Expression of NOTCH4 in Molm14-R cells transduced with shNOTCH4 #87, #88 and #90 decided by quantitative RT-PCR (normalized to GAPDH) (i) and Western blot (ii). Numbers under NOTCH4 bands in (ii) are the values of every particular person band relative to regulate (shNS) (normalized to ACTIN). F MTT assay and (G, H) apoptosis/dying assays of Molm14-R cells transduced with management shNS or shNOTCH4 (#90) +/− gilteritinib. I Western blot displaying affirmation of knockout of NOTCH4 in Molm14, Molm14-R and MV4;11-R cells. J–L MTT assay (i) and annexin V – primarily based apoptosis/dying assays (ii) of Molm14, Molm14-R and MV4;11-R cells with knockout of NOTCH4 (N4KO) or single information RNA controls (sgCtrl). Outcomes from a consultant knockout clone in every cell line are proven. Information are introduced because the imply ± SD. *p < 0.05 relative to regulate; *p < 0.05; ***p < 0.001 relative to regulate.
Subsequent, to judge how elevated NOTCH4 expression contributes to the survival and proliferation of FLT3 TKI-resistant cells, we inhibited NOTCH4 expression in Molm14-R cells utilizing NOTCH4 shRNA (shNOTCH4) knockdown (Fig. 2E). In comparison with the shRNA (shNS)-transduced management cells, shNOTCH4 (sh90)-transduced Molm14-R cells confirmed considerably diminished proliferation when handled with gilteritinib, an FDA accredited FLT3 TKI (Fig. 2F). Apoptosis and cell dying was additionally considerably elevated in shNOTCH4 (#90)-transduced cells each with and with out gilteritinib therapy (Fig. 2G, H).
To additional discover the impression of NOTCH4 expression on FLT3/ITD AML cells’ sensitivity to TKI, we knocked out NOTCH4 in Molm14 and TKI-resistant cells utilizing the CRISPR-Cas9 system (Fig. 2I, Supplementary Fig. S2). NOTCH4-knockout cell strains (Molm14-N4KO, Molm14-R-N4KO and MV4;11-R-N4KO) had been generated utilizing the lentiviral CRISPR-Cas9 system with plasmids acquired by way of Addgene (Addgene #108100 and #108098, each had been kindly offered by Dr. Christopher Vakoc) [28]. In comparison with single-guide management (Molm14-sgCtrl) cells, knockout of NOTCH4 in Molm14 (Molm14-N4KO) cells decreased proliferation and elevated apoptosis when handled with gilteritinib (Fig. 2J). NOTCH4-knockout Molm14-R (Molm14-R-N4KO) cells additionally displayed diminished proliferation and elevated apoptosis in response to gilteritinib as in comparison with management cells (Fig. 2K). Equally, knockout of NOTCH4 in MV4;11-R (MV4;11-R-N4KO) cells additionally results in diminished proliferation and elevated apoptosis upon gilteritinib therapy in comparison with management cells (Fig. 2L). These outcomes, along with the information obtained from the NOTCH4 shRNA knockdown cells, additional validates the discovering that diminished expression of NOTCH4 sensitizes TKI-resistant FLT3/ITD+ cells to FLT3 TKI therapy.
Gilteritinib and γ-secretase inhibitors have a synergistic impact on FLT3/ITD AML cells
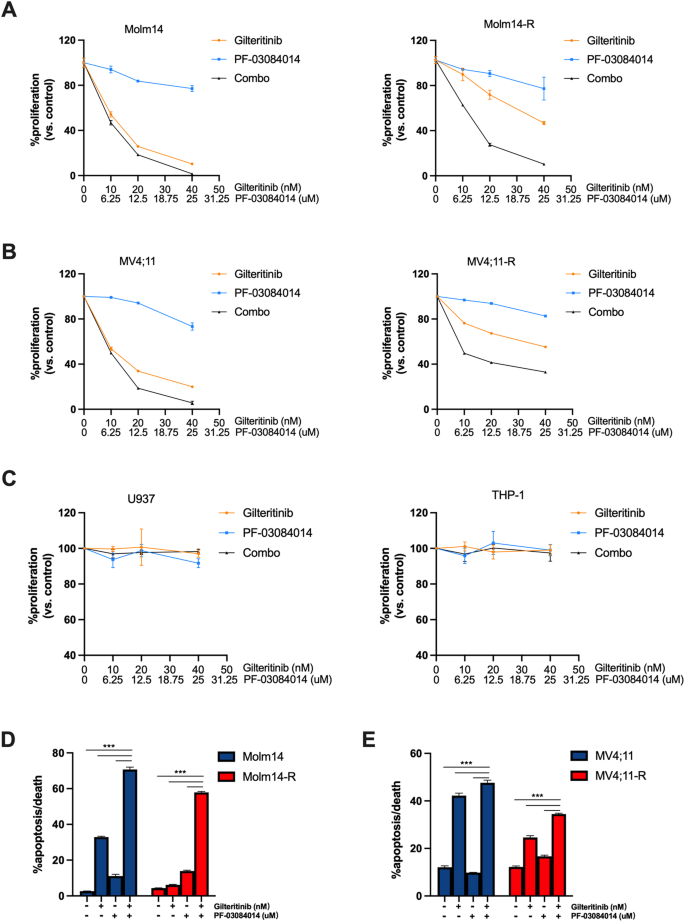
To additional discover the combinatorial results of NOTCH4 inhibition with FLT3 TKIs, we used PF-03084014 or DAPT, each γ-secretase inhibitors, to pharmacologically inhibit NOTCH4 exercise in FLT3/ITD cells. MTT assay confirmed enormously diminished proliferation of Molm14 or MV4;11 cells when handled with the mixture of gilteritinib and PF-03084014 (Fig. 3A, B, left panels; Supplementary Desk S5), with each CI values lower than 1.0, demonstrating synergy for the mixture. Moreover, mixture therapy resulted in considerably diminished proliferation of Molm14-R or MV4;11-R cells, with even decrease CI values than their parental cells, once more demonstrating synergy between the 2 medication (Fig. 3A, B, proper panels; Supplementary Desk S5). Equally, the mixture of gilteritinib with DAPT was extremely synergistic in lowering the proliferation of Molm14-R and MV4;11-R cells (CI values of 0.364 and 0.391, respectively) (Supplementary Fig. S3A, B). In the meantime, no response was noticed in U937 or THP-1 cells when handled with gilteritinib and/or γ-secretase inhibitors (Fig. 3C, Supplementary Fig. S3C).
MTT assays of (A) Molm14/Molm14-R, (B) MV4;11/MV4;11-R, (C) U937 and THP1 cells handled with gilteritinib and/or PF-03084014. Apoptosis assay of (D) Molm14/Molm14-R and (E) MV4;11/MV4;11-R cells handled with gilteritinib (20 nM) and/or PF-03084014 (12.5 μM). n = 3 for every cell line / therapy group. Information are introduced because the imply ± SD. ***p < 0.001.
Movement cytometry evaluation confirmed that the mixture of gilteritinib and PF-03084014 considerably elevated the share of Molm14 and MV4;11, in addition to Molm14-R and MV4;11-R cells present process apoptosis, when in comparison with both drug alone (Fig. 3D, E). Related outcomes had been seen for the mixture gilteritinib and DAPT (Supplementary Fig. S3D, E).
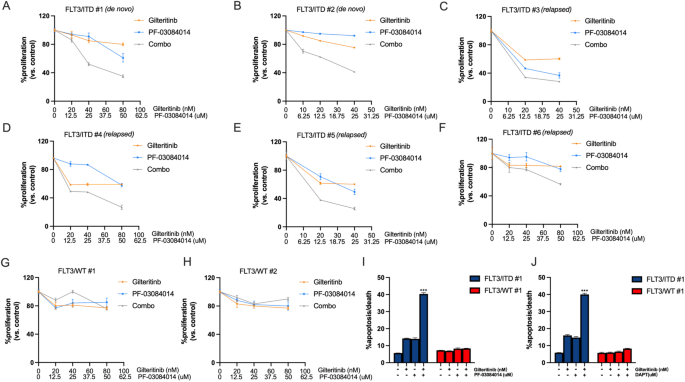
We then examined whether or not the synergistic results additionally current in main FLT3/ITD AML blast cells. When FLT3/ITD AML samples had been handled with the mixture of gilteritinib and PF-03084014, synergistic anti-proliferative results had been noticed in all 6 samples (2 de novo and 4 relapse) at efficient dose 50 (ED50) (Fig. 4A–F, Desk 1). WT FLT3 leukemic samples confirmed variable sensitivity to PF-03084014, however no additive or synergistic impact upon mixture therapy (Fig. 4G, H, Desk 1). The mix of gilteritinib and DAPT additionally resulted in the same impact in a main FLT3/ITD AML pattern (Supplementary Fig. S4). As well as, FLT3/ITD AML cells displayed considerably larger apoptosis when handled with the mixtures in comparison with both drug alone (Fig. 4I, J). In distinction, no important enhance in apoptosis was noticed in WT FLT3 leukemic samples handled with both single-drug or the mixture. Collectively, these outcomes recommend that FLT3/ITD AML cells, together with TKI-resistant FLT3/ITD cells, are very delicate to the mixed therapy of gilteritinib with γ-secretase inhibitors.
Major FLT3/ITD AML blast cells had been handled with gilteritinib and/or PF-03084014 for 72 h. A–H MTT assay of main FLT3/ITD (A–F) and FLT3/WT (G, H) AML affected person samples handled with gilteritinib and/or PF-03084014. Apoptosis assay of main FLT3/ITD and FLT3/WT AML affected person samples handled with gilteritinib/ PF-03084014 (I) or gilteritinib/ DAPT (J). n = 3 for every pattern/therapy group. Information are introduced because the imply ± SD. ***p < 0.001.
γ-secretase inhibitors repress the sustained activation of MAPK/ERK noticed in resistant FLT3/ITD AML cells regardless of FLT3 TKI inhibition and impacts expression of genes related to cell biking, differentiation and DNA restore
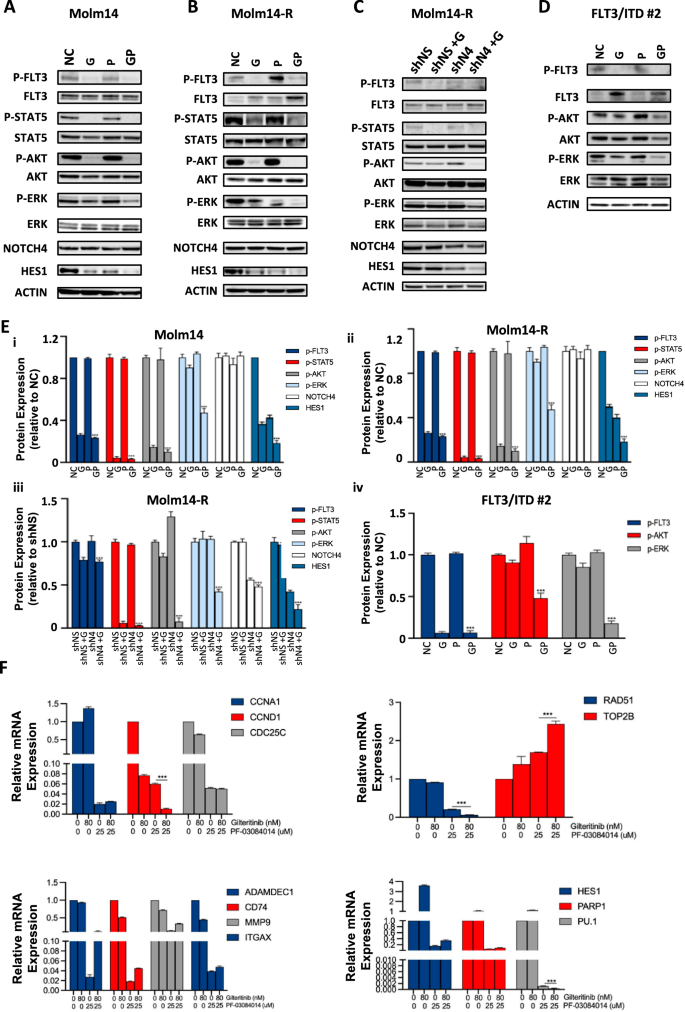
To grasp the mechanism(s) underlying the noticed mixture drug synergy, we regarded for any potential modifications in signaling pathways downstream of FLT3 in FLT3/ITD AML cells upon mixture therapy. 24 h after the mixture therapy of gilteritinib and PF-03084014, p-AKT and p-ERK ranges in Molm14 and Molm14-R cells had been considerably decreased (Fig. 5A, B, E). Mixture of gilteritinib and DAPT repressed p-ERK alerts within the Molm14-R however not parental Molm14 cells (Supplementary Fig. S5). Since γ-secretase inhibitors have an affect on all members of the NOTCH household in addition to probably having off-target results, we investigated the signaling modifications upon FLT3 TKI therapy in shNOTCH4 knockdown Molm14-R cells. As proven in Fig. 5C, E, each p-AKT and p-ERK had been considerably decreased upon gilteritinib therapy in shNOTCH4-knockdown Molm14-R cells in comparison with management. In line with what we noticed in Molm14 cells, in main FLT3/ITD AML cells p-AKT and p-ERK ranges had been additionally decreased with mixture therapy (Fig. 5D, E).
Western blots of (A) Molm14 and (B) Molm14-R cells handled with gilteritinib (G, 40 nM) and/or PF-03084014 (P, 12.5 μM) for twenty-four h. C Western blots of management shRNA (shNS) or NOTCH4 shRNA (shN4, shNOTCH4 #90)-transduced Molm14-R cells handled with gilteritinib (G, 80 nM). D Western blots of a FLT3/ITD main AML pattern handled with gilteritinib (G, 40 nM) and/or PF-03084014 (P, 12.5μM) for twenty-four h. NC: Unfavourable management. A consultant of three experiments is proven. E Graphs displaying the quantification of the phosphorylated protein bands proven in A–D (relative to the corresponding protein expression and normalized to ACTIN management) or protein expression (for NOTCH4 and HES1, normalized to ACTIN management) (Picture lab V5.2.1); ***p < 0.001 in comparison with the remainder of the samples in the identical group. F Expression of genes in main FLT3/ITD AML cells from a affected person handled with gilteritinib (80 nM) and/or PF-03084014 (25 μM) for 48 h.
In an effort to additional examine the mobile outcomes brought on by the synergistic mixture remedy, we examined the impression of PF-03084014 and gilteritinib on cell biking (CCNA1, CCND1, CDC25C), DNA restore (RAD51, TOP2B), differentiation (ADAMDEC1, CD74, MMP9, ITGAX) and NOTCH signaling pathway (HES1, PARP1 and PU.1) gene expression in main FLT3/ITD AML affected person samples. Notably, expression of CCND1, RAD51 and PU.1 had been considerably downregulated whereas TOP2B was considerably upregulated in cells handled with gilteritinib and PF-03084014 in comparison with management cells or cells handled with both drug alone, indicating these genes could also be concerned within the motion of the mixture therapy (Fig. 5F). In a complementary genetic NOTCH4 knockout experiment, related reductions of CCND1 expression had been noticed in NOTCH4-knockout Molm14 (Molm14-N4KO) in addition to MV4;11-R (MV4;11-R-N4KO) cells when handled with gilteritinib (Supplementary Fig. S6A).
Since CCND1 is called a key cell cycle regulator modulating the G1 to S section transition and performs an essential position within the pathogenesis of most cancers [29], we additional investigated the impression of the drug mixture on cell cycle development. BrdU-incorporation cell cycle assays exhibit that in comparison with management, knockout of NOTCH4 in Molm14 cells blocked the cells from coming into S section. Remedy with gilteritinib leads to additional G1 arrest in a dose-dependent method (Supplementary Fig. S6B). The identical outcomes had been discovered within the MV4;11-R cells (Supplementary Fig. S6C). The block of G1 to S section transition seen in gilteritinib handled NOTCH4-knockout Molm14 and MV4;11-R cells is per the position CCND1 performs within the G1 to S transition.
Altogether, the outcomes recommend that NOTCH4 inhibition might block the adaptive suggestions mechanism that leads to reactivation of ERK and/or AKT signaling that happens in response to FLT3 TKI inhibition in FLT3/ITD AML cells. Overcoming ERK reactivation sensitizes FLT3/ITD cells, particularly these with FLT3 TKI resistance, to FLT3 inhibition. It additionally impacts expression of genes concerned in cell cycle, differentiation and DNA restore and ultimately results in cell dying.
Mixture therapy with FLT3 TKI and NOTCH4 inhibition reduces illness development of FLT3/ITD AML cell strains and a patient-derived FLT3/ITD AML xenograft (PDX) in vivo
To additional consider the results of NOTCH4 inhibition on FLT3/ITD cells in vivo, we transplanted shNOTCH4 and shNS Molm14 cells into NSG mice adopted by therapy with the FLT3 TKI sorafenib (Fig. 6A). The outcomes exhibit a major discount in BM involvement of leukemic cells and spleen weight within the sorafenib-treated shNOTCH4-Molm14 transplanted mice in comparison with vehicle-treated shNOTCH4, sorafenib-treated shNS and vehicle-treated shNS Molm14 recipients, respectively (Fig. 6B, C). As well as, shNOTCH4 Molm14-transplanted mice had a considerably longer median survival (52 days) when handled with sorafenib in comparison with these with out sorafenib therapy or shNS Molm14 transplanted mice, no matter sorafenib therapy (38, 38 and 29 days, respectively) (Fig. 6D). These knowledge recommend that the mixture of FLT3 TKI with NOTCH4 inhibition is efficient in eliminating FLT3/ITD Molm14 cells in vivo.
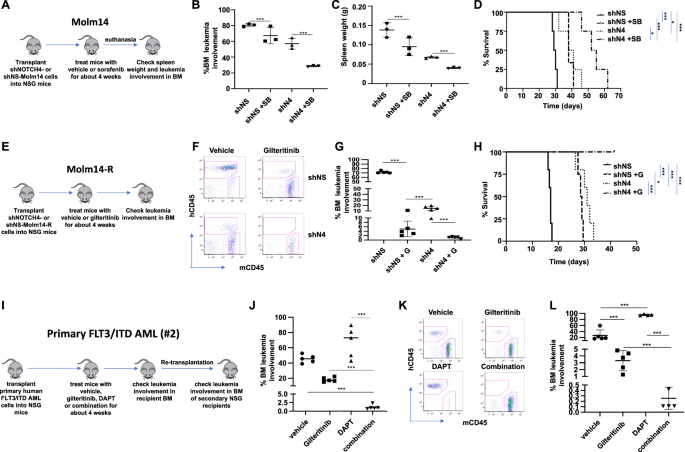
A Mouse xenograft schematic diagram of NOCTH4 knockdown Molm14 mouse mannequin. B Share of BM leukemia involvement in recipient BM, C spleen weight and D general survival in shNOTCH4 (shN4) or management shNS Molm14 teams handled with/with out sorafenib (8 mg/kg/day, oral gavage) for 4 weeks. E Mouse xenograft schematic diagram for NOCTH4 knockdown Molm14-R mouse mannequin. F, G Ranges of BM leukemia involvement and H general survival in shNOTCH4 (shN4) or shNS Molm14-R mice handled with/with out gilteritinib (30 mg/kg/day, oral gavage) for 4 weeks or till the mice succumbed to dying. I Mouse xenograft schematic diagram for PDX mannequin (main FLT3/ITD AML #2) (J) ranges of BM leukemia involvement in PDX mice handled with car, gilteritinib (G, 30 mg/kg/day, oral gavage) and/or DAPT (D, 10 mg/kg/day, oral gavage) for 4 weeks. Ok, L Leve ls of BM leukemia involvement in re-transplanted secondary recipients. BM had been collected and subjected to stream cytometry evaluation for BM involvement evaluation 4 weeks after re-transplantation. *p < 0.05, ***p < 0.001.
We additional investigated the results of NOTCH4 inhibition on sensitizing FLT3 TKI-resistant AML cells to TKI therapy in vivo (Fig. 6E). We discovered that shNOTCH4 Molm14-R cells demonstrated decrease BM leukemia involvement in comparison with management shNS Molm14-R cells. Whereas therapy with gilteritinib diminished BM involvement in shNS Molm14-R recipients, an extra important discount was noticed for shNOTCH4 Molm14-R recipients (Fig. 6F, G). As well as, the gilteritinib-treated shNOTCH4 Molm14-R mice had been all nonetheless alive on the time the experiment was terminated (day 80) in comparison with median survival of 62, 56 and 34 days, respectively for the shNOTCH4 Molm14-R mice with out gilteritinib therapy, or the shNS Molm14-R mice with or with out gilteritinib therapy (Fig. 6H).
To evaluate the results of FLT3 and NOTCH mixture inhibition on main FLT3/ITD AML cells in vivo, we generated a PDX mouse mannequin by transplanting BM cells collected from a de novo FLT3/ITD AML affected person (FLT3/ITD #2) into NSG mice (Fig. 6I). In comparison with vehicle-treated (45.5% ± 6.0%), gilteritinib-treated (18.3% ± 2.9%) or DAPT-treated (67.3 ± 20.0%) recipients, the extent of BM leukemia involvement was remarkably diminished in mice handled with the mixture of the two medication (1.2% ± 0.7%) (Fig. 6J). These outcomes once more help that mixed inhibition of FLT3 and NOTCH reduces the flexibility of FLT3/ITD AML cells to progress in vivo.
Given these outcomes, we needed to additional study the impact of mixed inhibition on leukemic initiating cells (LICs). As human AML LICs are characterised by the flexibility of the cells to propagate AML in immunodeficient mice, we investigated the flexibility of the medication to scale back the LICs in vivo. Transplant of equal numbers of hCD45+ cells harvested from the first handled recipients from every group had been re-transplanted into secondary recipients and allowed to develop for 120 days. Cells from the mixture handled mice resulted in enormously diminished ranges of BM involvement (0.3% ± 0.2%) in secondary recipients in comparison with management (27.1% ± 18.4%), gilteritinib (3.3% ± 1.5%) or DAPT (95.0% ± 1.9%) handled mice (Fig. 6K, L). Taken collectively, these knowledge recommend that NOTCH4 inhibition combines with FLT3 TKI to considerably scale back enlargement and LICs of FLT3/ITD AML cells in vivo.