Inhibition of DUSP18 enhances the perform of tumor-infiltrating CD8+ T cells
To systematically establish genetic targets in most cancers cells whose loss represses immune evasion, a way to establish these genes was developed (Fig. 1a). MC38 CRC cells had been engineered to precise Cas9 (Supplementary Fig. 1a) and single information RNAs (sgRNAs) from a Mouse CRISPR Deletion Library (termed KPD library) generated in opposition to ~2000 genes encoding drug targets, kinases and phosphatases (Supplementary Information 1)39. After in vitro puromycin choice for 7 days to make sure secure gene modifying, the tumor cells had been subcutaneously transplanted into immunodeficient or immunocompetent mice, with the latter being meant to generate an immune-selective stress on the tumor cells. On the time of harvesting 16 days later, tumors grown in immunodeficient hosts had been considerably bigger (Supplementary Fig. 1b). Upon amplification and sequencing of built-in sgRNA inserts from every group, we recognized a quantity that had been depleted in tumors from the immunocompetent cohort based mostly on the MAGeCK algorithm (Fig. 1b and Supplementary Information 2). As proof that sure gRNAs may very well be predictably chosen in opposition to, these focusing on Ptgs1 (Cox1), which promotes PGE2 manufacturing and reduces CD8+ T cell infiltration, had been depleted in tumors engrafted in immunocompetent mice40. A variety of different strategies together with DrugZ41 and DESeq242 for analyzing CRISPR screens outcomes had been additionally adopted to confirm the accuracy of our evaluation (Supplementary Fig. 1c and Supplementary Information 2). The outcomes confirmed that some genes with the potential to advertise tumor immune evasion, comparable to Ptgs1 and Dusp18, and many others., had been considerably enriched in all three analytical strategies, which excludes the impact of various analytical strategies on the variations in outcomes.
a Workflow of in vivo screens to establish potential targets for immune evasion. b MAGeCK evaluation and RRA rating of prime depleted genes within the screens (n = 2). Ranked dot plots of depleted genes in immunocompetent hosts in contrast with immunodeficient hosts are proven. c Venn diagram exhibiting the overlap of 45 screened genes and 735 CTL associated genes within the TCGA-COAD database. d Differential expression evaluation for DUSP18 in tumors and regular tissues. COAD, colon adenocarcinoma (tumor samples (n = 471), regular samples (n = 41) (left); tumor samples (n = 41), paired regular samples (n = 41) (center)). READ, rectal adenocarcinoma (tumor samples (n = 167), regular samples (n = 10) (proper). e Scatterplot exhibiting the correlation of the DUSP18 expression with that of a number of activated T cell-related genes in TCGA-COAD samples (n = 471 tumor samples). Tumor progress curves (f) or tumor weight (g) for 4–6 weeks nude mice bearing MC38 tumors (n = 4). Tumor progress curves (h), tumor weight (i) or survival curves for 4–6 weeks previous C57BL/6 bearing MC38 tumors (n = 5). Tumor photograph (ok), tumor progress curves (l) and tumor weight (m) for C57BL/6 bearing MC38 tumors (n = 6). Mice had been handled with anti-CD8a antibodies on days −1, 0, 7, and 14, n FACS evaluation of tumor-infiltrating CD4+ and CD8+ T cells in shCtrl or shDusp18 MC38 tumors. (n = 3). o Quantification of IFN-γ and GzmB CD8+ TILs. Tumor-infiltrating T cells had been pre-stimulated with PMA, ionomycin and brefeldin A for 3 h (n = 3). IFN-γ and GzmB-producing cells had been decided by movement cytometry. p Quantification of PD-1, TIM-3 and CTLA-4 CD8+ TILs via movement cytometry (n = 3). Information are offered as imply ± SD (f–p). P values had been calculated by MAGeCK-test module utilizing a modified sturdy rating aggregation (α-RRA) evaluation (b), unpaired two-tailed t-tests (d: left and proper boxplot, l, m), paired two-tailed t-tests (d center boxplot), one-way ANOVA (f, g, h, i, n–p) or log-rank (Mantel-Cox) take a look at (j). P values and R had been calculated by Spearman’s correlation evaluation. Two-sided P worth was given (c, e); ns not important. Supply information are supplied as a Supply Information file.
The Most cancers Genome Atlas (TCGA)-colon adenocarcinoma (COAD) dataset was additionally analyzed to establish cytotoxic T lymphocyte (CTL) scores utilizing a set of 5 beforehand reported genes (CD8A, CD8B, GZMA, GZMB, and PRF1) as a surrogate of tumor infiltrating CD8+ T cells43. This evaluation revealed that sufferers with increased CTL scores skilled higher survival (Supplementary Fig. 1d). Subsequent, the correlation between protein-coding genes and CTL scores was calculated and the outcomes confirmed that 735 genes had been negatively related to CTL scores (Supplementary Information 3). Lastly, the intersection between the CRISPR screens chosen genes and people correlating negatively with CTL scores was decided with Dusp18 being probably the most noteworthy (Fig. 1c). Increased DUSP18 mRNA expression degree was additionally present in CRC samples in contrast with regular colo-rectal tissues (Fig. 1d). DUSP18 expression additionally negatively correlated with 3 of the genes used to calculate the CTL scores, specifically CD8A, PRF1 and GZMA in addition to 3 others (TNFRSF18, GZMH and GNLY) (Fig. 1e), whereas additionally correlating with CD8+ T cell infiltration (Supplementary Fig. 1e). Kaplan–Meier survival curves based mostly on excessive or low DUSP18 mRNA expression had been generated utilizing the Tumor Immune Dysfunction and Exclusion (TIDE) device43 to discover the affiliation between CTL scores and total survival in CRC sufferers. In these CRC sufferers with low DUSP18 mRNA expression, excessive CTL scores had been related to higher survival whereas these with excessive DUSP18 mRNA expression and excessive CTL scores confirmed worse survival. Taken collectively, these findings counsel that prime DUSP18 mRNA expression is related to T cell dysfunction (Supplementary Fig. 1f).
To find out whether or not Dusp18 inhibition impacts T cell-mediated antitumor perform, shRNA-mediated inhibition of Dusp18 was carried out in MC38 CRC cells and ovalbumin (OVA)-expressing B16 melanoma (B16-OVA) cells (Supplementary Fig. 1g, h). This didn’t impair tumor cell proliferation in vitro (Supplementary Fig. 1i) or tumor progress in immunodeficient nude mice (Fig. 1f, g and Supplementary Fig. 1j). Comparable findings had been noticed with the human HCT116 CRC cell line (Supplementary Fig. 1k–m). Nonetheless, Dusp18 inhibition did impair tumor progress and extended survival in immunocompetent mice (Fig. 1h-j) though depleting CD8+ T cells fully eradicated this progress drawback (Fig. 1k–m and Supplementary Fig. 1n). Thus, our findings advised that CD8+ T-cell-mediated immunity is concerned in some facet(s) of DUSP18-regulated tumor progress.
The impact of Dusp18 inhibition on the tumor immune panorama was additional explored utilizing movement cytometry. This confirmed that absolutely the numbers and share of innate cells in shCtrl and shDusp18 tumors had been equivalent (Supplementary Fig. 1o, p). Though each CD8+ and CD4+ T cell numbers had been elevated in shDusp18 tumors, the proportion of those immune cells was enhanced solely by CD8+ however not CD4+ T cells (Fig. 1n and Supplementary Fig. 1p). Immuno-histochemical staining additionally confirmed that inhibition of Dusp18 considerably upregulated CD8+ T cell infiltration with little impact on CD4+ T cells and NK cells (Supplementary Fig. 2a). Infiltrating CD8+ T cells from shDusp18 tumors confirmed elevated expression of cytotoxic molecules IFN-γ and granzyme B (Fig. 1o), alongside lowered expression of exhaustion-associated transcripts PD-1, TIM-3, and CTLA-4 (Fig. 1p and Supplementary Fig. 2b). After MC38 cells overexpressing wild-type (WT) or phosphatase useless mutant (Lifeless) Dusp18 had been inoculated into mice, solely the previous promoted tumor progress in immunocompetent mice however not in nude mice (Supplementary Fig. 2c, d). Moreover, solely WT Dusp18 overexpression inhibited CD8+ T cell infiltration (Supplementary Fig. 2e, f) and cytotoxic perform (Supplementary Fig. 2g). The expression of exhaustion-associated molecules in CD8+ T had been additionally increased in WT Dusp18 overexpressing tumors (Supplementary Fig. 2h). Thus, the above outcomes confirmed that DUSP18’s inhibition of CD8+ T cell infiltration and its related cytotoxicity relies on its phosphatase exercise.
DUSP18 inhibition impairs ldl cholesterol biosynthesis
To find out whether or not the attenuation of DUSP18 expression alters intestinal tumorigenesis, we crossed Dusp18flox/flox mice with Villincre mice to generate mice with intestinal epithelium-specific depletion of Dusp18 (termed CKO mice) (Supplementary Fig. 2i). Wild-type (WT) and CKO mice had been then used to generate CRCs utilizing the AOM/DSS protocol (Fig. 2a). Surprisingly, CKO mice developed considerably fewer CRCs (Fig. 2b) and survived longer (Fig. 2c). To reveal that Dusp18 loss was triggering an adaptive immune response, WT and CKO tumors had been examined for proof of infiltration by CD8+ T cells, CD4+ T cells and NK cells. Immunofluorescence research confirmed WT tumors to be immune deserts that had been largely devoid of CD8+ T cells (Fig. second) whereas the isogenic CKO tumors had been extremely infiltrated by CD8+ T cells (Fig. second). In distinction, CD4+ T cell and NK cell infiltration didn’t change considerably (Supplementary Fig. 2j). Expression of the cell proliferation markers Ki-67 additionally didn’t differ within the two tumor tissues (Supplementary Fig. 2j). Tumors from CKO mice had barely increased ranges of JNK phosphorylation (Phospho-JNK (Tyr185)), whereas ERK and p38 remained virtually unchanged amongst a number of MAPKs (Supplementary Fig. 2k). The expression of c-Myc, Cyclin D1, and Cox-2 was not considerably completely different in these two tumor tissues (Supplementary Fig. 2k).
a Scheme for the AOM/DSS-induced colon most cancers mannequin in WT and CKO mice. b Colon tumor numbers in mice from (a), n = 6. c Survival curves, n = 10. d Consultant IF photos and quantification of CD8+ T cells in AOM/DSS-induced colon most cancers shaped by WT and CKO mice. Scale bar, 10 μm. For every mouse, 5 fields from one unbiased tumor of every mouse had been analyzed (n = 6). GO (e) and KEGG (f) analyses present the altered pathways after Dusp18 inhibition. g Heatmap of RNA-seq expression values of ldl cholesterol biosynthesis genes based mostly on shDusp18 and shCtrl MC38 cells (n = 3). h GSEA evaluation for ldl cholesterol homeostasis pathway genes in shCtrl versus shDusp18 cells. i qRT-PCR evaluation of the indicated ldl cholesterol biosynthetic pathway genes in shDusp18 and shCtrl MC38 cells (n = 3). j Western blot evaluation for the indicated proteins in shDusp18 and shCtrl MC38 cells. The IB information are consultant of three unbiased experiments. ok Immunoblot (IB) evaluation of the indicated proteins in CRCs from AOM/DSS-treated mice, n = 3 unbiased experiments. l qRT-PCR evaluation of the indicated ldl cholesterol biosynthetic pathway genes in CRCs from WT and CKO mice, n = 3 unbiased experiments. m Heatmap exhibiting differential expression of genes within the ldl cholesterol metabolism and antigen presentation within the proteomics of shCtrl and shDusp18 MC38 cells (n = 3). n Scatterplot exhibiting the correlation between DUSP18 expression ranges with these of ldl cholesterol biosynthesis-related genes in TCGA-COAD samples (n = 473). Spearman correlation coefficient (r) and p worth are marked. Information are offered as imply ± SD (b, d, i, l). P values had been calculated by modified Fisher’s actual checks (e, f), Kolmogorov–Smirnov checks (h), unpaired two-tailed t checks (b, d, l), one-way ANOVA (i) or log-rank (Mantel-Cox) take a look at (c). P values and R had been calculated by Spearman’s correlation evaluation. Two-sided P worth was given (n); ns not important. Supply information are supplied as a Supply Information file.
To ascertain the molecular mechanism(s) by which DUSP18 suppresses antitumor immunity, RNA-seq was carried out utilizing shCtrl and shDusp18 MC38 cells. Gene ontology (GO) and Kyoto Encyclopedia of Genes and Genomes (KEGG) evaluation confirmed that the differentially expressed genes had been concerned in ldl cholesterol biosynthesis and its downstream metabolism and had been expressed at decrease ranges in shDusp18 cells (Fig. 2e–g). Gene set enrichment evaluation (GSEA) in these cells additionally confirmed that genes concerned in ldl cholesterol homeostasis had been positively enriched within the management cells (Fig. 2h). We additional confirmed decreased mRNA expression and protein ranges for chosen genes in shDusp18 MC38 cells and CKO tumor tissues (Fig. 2i-l and Supplementary Fig. 3a). RNA-seq was additionally carried out in shCtrl and shDUSP18 human CRC HCT116 cells to validate these outcomes from mice MC38 cells. Each GO, KEGG and GSEA analyses revealed that inhibition of DUSP18 considerably lowered the expression of genes concerned within the ldl cholesterol biosynthesis pathway (Supplementary Fig. 3b–f). We confirmed decreased mRNA expression and protein ranges for chosen ldl cholesterol biosynthesis genes in shDUSP18 HCT116, SW480 and extra human CRC cells (Supplementary Fig. 3g–j). Proteomic evaluation in MC38 cells additionally validated the above outcomes (Supplementary Information 4). As proven in Supplementary Fig. 4a, inhibition of Dusp18 lowered the degrees of 440 proteins and elevated the degrees of 449 proteins. KEGG signaling pathway enrichment evaluation of those differentially expressed proteins confirmed that these which had been down-regulated in shDusp18 cells had been primarily enriched in metabolic pathways and ldl cholesterol biosynthesis alerts (Supplementary Fig. 4b), whereas upregulated proteins had been primarily enriched in antigen presentation alerts (Supplementary Fig. 4c). These outcomes had been in line with our RNA-Seq outcomes. Of explicit observe, inhibition of Dusp18 decreased the degrees of SREBP2, HMGCR, LSS, and SQLE (Fig. 2m). Collectively above outcomes underscored the significance and conservation of DUSP18 in regulating the ldl cholesterol synthesis pathway.
DUSP18 expression ranges had been positively correlated with ldl cholesterol biosynthesis signaling in TCGA-COAD and GEO datasets (Fig. 2n and Supplementary Fig. 4d). Since DUSP18 appeared to behave as a phosphatase to control the ldl cholesterol synthesis signaling pathway, we hypothesized that DUSP18 regulates a TF that’s concerned in sustaining the pathway. We examined the expression of a number of necessary TFs concerned within the ldl cholesterol synthesis pathway based mostly on the inhibition of Dusp18, together with SREBP2, SREBP1, Ad4BP, Maf, Notch1, XBP1, RORγ, and c-Myc28,36,44,45,46,47,48,49. As proven in Supplementary Fig. 4e, inhibition of Dusp18 solely down-regulated SREBP2 protein ranges and had little impact on the protein ranges of the opposite TFs. Decreased expression of SREBF2 mRNA and SREBP2 protein ranges had been validated in human and murine CRC cells expressing DUSP18 shRNA (Fig. 2i-l and Supplementary Fig. 3a, g–j). Importantly, ranges of each the total (p-SREBP2) and cleaved (n-SREBP2) types of SREBP2 protein28 had been decreased. Moreover, DUSP18 and SREBF2 mRNA ranges had been considerably positively correlated in TCGA-COAD (Supplementary Fig. 4f), and pan-cancer analyses revealed an identical correlation in most tumors, together with CRCs, liver and kidney cancers (Supplementary Fig. 4g). General, these findings help the notion that DUSP18 exerts management over the tumor-associated immune panorama by regulating mobile ldl cholesterol biosynthesis.
USF1 is crucial for DUSP18-mediated regulation of ldl cholesterol biosynthesis
With the intention to elucidate how DUSP18 regulates the expression of SREBF2 in CRC, we requested whether or not DUSP18 interacts with a TF that regulates SREBF2 gene expression. GSEA of our RNA-seq information revealed USF1 transcripts to be most importantly enriched, thus main us to hypothesize that USF1 could be the TF that not directly regulates ldl cholesterol metabolism maybe by way of an affiliation with SREBP2 (Fig. 3a). Utilizing the Human TFDB and JASPAR web sites to foretell the putative binding websites for USF1 within the SREBF2 promoter (Supplementary Fig. 5a), and analyzing of ChIP-seq information from HCT116 cells confirmed a USF1-binding peak within the SREBF2 promoter (Supplementary Fig. 5b). The practical penalties of this had been established by demonstrating that inhibition of USF1 considerably down-regulated each SREBF2 mRNA and SREBP2 protein ranges (Fig. 3b and Supplementary Fig. 5c). ChIP assays additionally revealed that USF1 occupied the SREBF2 gene promoter in HCT116 cells (Fig. 3c). To verify this, luciferase reporters containing SREBF2 promoter components had been rendered USF1-responsive, whereas inhibition of USF1 dramatically decreased luciferase exercise pushed by a WT SREBF2 promoter, however not by a promoter that was unable to bind USF1 (Fig. 3d). These information indicated that USF1 can bind to the SREBF2 promoter and regulate its transcriptional output. In GSEA evaluation, inhibition of DUSP18 led to a notable down-regulation of the USF signature (Fig. 3e).
a Candidate transcription issue prediction was carried out by GSEA of regulatory goal gene units. The highest 8 TFs ranked by GeneRation scores and p values are proven. b Inhibition of USF1 suppresses the indicated protein ranges. c qPCR ChIP evaluation of USF1 binding to SREBF2 promoter areas in HCT116 cells (n = 3). d A luciferase vector pushed by a WT SREBF2 promoter, however not by a non-USF1-binding mutant promoter, is aware of inhibition of USF1 (n = 3). e GSEA evaluation for USF signature in shCtrl versus shDUSP18 cells. f DUSP18 inhibition suppresses USF1 protein ranges. g USF1 threonine dephosphorylation mediated by DUSP18. HA-USF1 was co-transfected with Flag-DUSP18 (WT or phosphorylase Lifeless) into HEK293T cells, and the cell lysates had been subjected to immunoprecipitation. h Dephosphorylation of USF1 T100 by DUSP18. Flag-USF1s (WT, T10A, T100A, T131A, T153A and T194/195A) had been co-transfected with HA-DUSP18WT or DUSP18Lifeless into HEK293T cells, and the cell lysates had been subjected to immunoprecipitation. i Stability of Flag-USF1 (WT, T100A and T100D) proteins in HEK293T cells (left) within the presence of CHX block. Flag-USF1 protein quantified by densitometry, with β-actin as a loading management (proper) (n = 3). j Decreased Flag-USF1 (T100A) polyubiquitination. Myc-Ub was co-transfected with Flag-USF1 (WT, T100A and T100D) into HEK293T cells, and the cell lysates had been subjected to immunoprecipitation. ok Overexpression of Flag-DUSP18 has no impact on the SREBP2 protein degree mediated by USF1 inhibition. l Subcutaneous xenograft experiments in C57BL/6 had been carried out within the indicated MC38 cells group. Tumor progress curves and weights are proven (n = 4). The share of infiltrating CD8+ T cells (n = 3) (m), effector molecules (n = 3) (n), and exhausted molecules (n = 3) (o) within the indicated group. Information are offered as imply ± SD (c, d, i, l–o). P values had been calculated by modified Fisher’s actual checks (a), Kolmogorov–Smirnov checks (e), unpaired two-tailed t-tests (c, i), one-way ANOVA (d, l–o); ns, not important. All IB information are consultant of three unbiased experiments. Supply information are supplied as a Supply Information file.
Primarily based on the above conjecture, we subsequent requested whether or not DUSP18 interacts with USF1 in order to have an effect on the expression of SREBF2. Coimmunoprecipitation assays demonstrated interactions between each endogenous and exogenous DUSP18 and USF1 (Supplementary Fig. 5d, e). Deletion analyses confirmed that the N terminal area of USF1 and the DSPc area of DUSP18 had been essential for this interplay (Supplementary Fig. 5f, g). Nonetheless, inhibition of DUSP18 decreased USF1 protein with out affecting its mRNA degree (Fig. 3f and Supplementary Fig. 5h). In keeping with this, inhibition of DUSP18 considerably lowered the half-life of endogenous USF1 (Supplementary Fig. 5i).
For the reason that earlier outcomes indicated that DUSP18’s skill to dam T cell activation requires its phosphatase exercise (Supplementary Fig. 2c–h), its skill to dephosphorylate and activate USF1 was investigated. To this finish, Flag-tagged WT and enzymatically useless DUSP18 and HA-tagged USF1 had been co-expressed in HEK293T cells adopted by anti-HA antibody and immunoblotting with anti-pan phospho-Tyr, Ser and Thr antibodies. This confirmed that DUSP18 selectively dephosphorylates USF1 at a number of Thr residues and advised that that is liable for stabilizing USF1 and permitting it to transcriptionally activate SREBF2 (Fig. 3g). After serially mutating particular person Thr residues in USF1 and co-expressing these with HA-DUSP18 in HEK293T cells, we discovered that the USF1 mutant T100A not served as a dephosphorylation substrate for HA-DUSP18 (Fig. 3h). In keeping with the above speculation, cycloheximide chase experiments indicated that USF1T100A had an prolonged half-life relative to that of both WT USF1 or the phospho-mimetic USF1T100D mutant (Fig. 3i). USF1T100A was additionally much less topic to ubiquitination (Fig. 3j), in addition to having extra binding with USF2 (Supplementary Fig. 5j), whose binding is necessary for USF1 to carry out its transcriptional perform as beforehand famous50. In distinction to this, binding to the transcriptional repressor Cha was not considerably altered (Supplementary Fig. 5k).
Casein kinase 2 (CK2) is a identified kinase for USF1 T10050. To look at the cross-talk between DUSP18 and CK2 on USF1 threonine phosphorylation, DUSP18 and CK2β had been co-expressed or co-depleted in CRC cells. The research confirmed that DUSP18 decreased CK2β-mediated phosphorylation of USF1 (Supplementary Fig. 5l, m). Subsequently, DUSP18 and CK2β seem to perform antagonistically to control USF1 stability.
Lastly, we discovered that, whereas DUSP18 overexpression didn’t upregulates the SREBP2 in CRC cells with inhibition of USF1 (Fig. 3k), USF1 overexpression may largely rescue the SREBP2 protein degree decline mediated by DUSP18 inhibition (Supplementary Fig. 5n). These outcomes counsel that USF1 performs an indispensable function within the regulation of SREBP2 by DUSP18. As well as, overexpression of USF1T100A however not USF1T100D rescued the inhibition of CRC cell progress mediated by Dusp18 inhibition (Fig. 3l). USF1T100A overexpressing tumors exhibited fewer CD8+ T cells infiltration (Fig. 3m) and decrease expression ranges of the cytotoxic molecules IFN-γ and granzyme B (Fig. 3n). In distinction, they expressed increased ranges of T cell exhaustion molecules comparable to PD-1, TIM-3 and CTLA-4 (Fig. 3o). Collectively, these findings level to the existence of a transcriptional cascade during which DUSP18 dephosphorylates and stabilizes USF1, which in flip upregulates SREBP2 and ldl cholesterol biosynthetic signaling.
DUSP18 ablation induces T cell activation in vitro
As a result of CD8+ T cells should acknowledge MHC-I molecules on the floor of tumor cells previous to initiating tumor cell killing (MHC restriction), the impact of Dusp18 inhibition on tumor cell antigen presentation was examined. Two well-established mouse synergic tumor fashions, MC38-OVA and B16-OVA, had been chosen. The next degree of MHC-I (H2-Kb) was detected in MC38-OVA and B16-OVA cells with Dusp18 inhibition (Fig. 4a, c), in addition to elevated MHC-I-bound SIINFEKL (OVA epitope peptide) complicated expression (Fig. 4b, d). Since Dusp18 mediates tumor immune evasion in a CD8+ T cell-dependent method, the cytotoxic killing of B16-OVA-Luc and MC38-OVA-Luc cells (expressing Ovalbumin and Luciferase) by OT-I T cells was measured in an in vitro killing system (Fig. 4e). This demonstrated that apoptosis and cell loss of life had been extra pronounced in shDusp18 cells (Fig. 4f, g). We subsequent co-cultured tumor cells and CD8+ T cells to discover the impact of Dusp18 inhibition on CD8+ T cell activation, effector perform, and expression of exhausted-related molecules (Fig. 4h). Inhibition of Dusp18 considerably enhanced expression of activation molecules CD69 and CD25, IFN-γ and GzmB manufacturing of co-cultured CD8+ T cells (Fig. 4i, j and Supplementary Fig. 6a). As well as, the proportion of PD-1+, TIM-3+, CTLA-4+ and PD-1+TIM-3+ CD8+ T cells decreased within the shDusp18 cohort (Fig. 4k, l and Supplementary Fig. 6b, c). Collectively, these findings demonstrated that DUSP18 in tumor cells suppresses CD8+ T cell activation and cytotoxicity and promotes CD8+ T cell exhaustion.
Expression ranges of H2-Kb and H2-Kb-SIINFEKL on shDusp18 and shCtrl MC38-OVA (n = 3) (a, b) and B16-OVA (n = 3) (c, d) cells had been decided by FACS. (MFI, imply fluorescence depth). e Schematic illustration of the in vitro T cell killing assay. MC38-OVA-Luc or B16-OVA-Luc shCtrl and shDusp18 cells had been co-cultured with splenic CD8+ T cells from OVA-specific T cell receptor transgenic (OT-I) mice. Cytotoxic results of OT-I had been measured by Annexin V/propidium iodide staining (n = 3) and bioluminescence signaling (n = 3) from MC38-OVA (f) and B16-OVA (g) cells after being co-cultured with OT-I for 48 h. h Schematic illustration of the ex vivo T cell co-culture assay. Splenic CD8+ T cells had been activated with the CD3 and CD28 antibodies, and co-cultured with shDusp18 or shCtrl tumor cells. After 24 h of coculture, cells had been harvested and processed for movement cytometry evaluation. i Quantification of CD69 (left) or CD25 (proper) expression in CD8+ T cells after co-culture with MC38 cells. (n = 3). j Quantification of IFN-γ (left) or GzmB (proper) manufacturing in CD8+ T cells after co-culture with MC38 cells (n = 3). ok, l Quantification of PD-1+, TIM-3+, CTLA-4+ and PD-1+ TIM-3+ share in CD8+ T cells after co-culture with MC38 cells (n = 3). Information are offered as imply ± SD (a–d, f, g, i–l). P values had been calculated by one-way ANOVA; ns, not important. Supply information are supplied as a Supply Information file.
Tumor-cell-derived lanosterol within the TME diminishes CD8+ T cell activation by inhibiting KRAS-ERK signaling
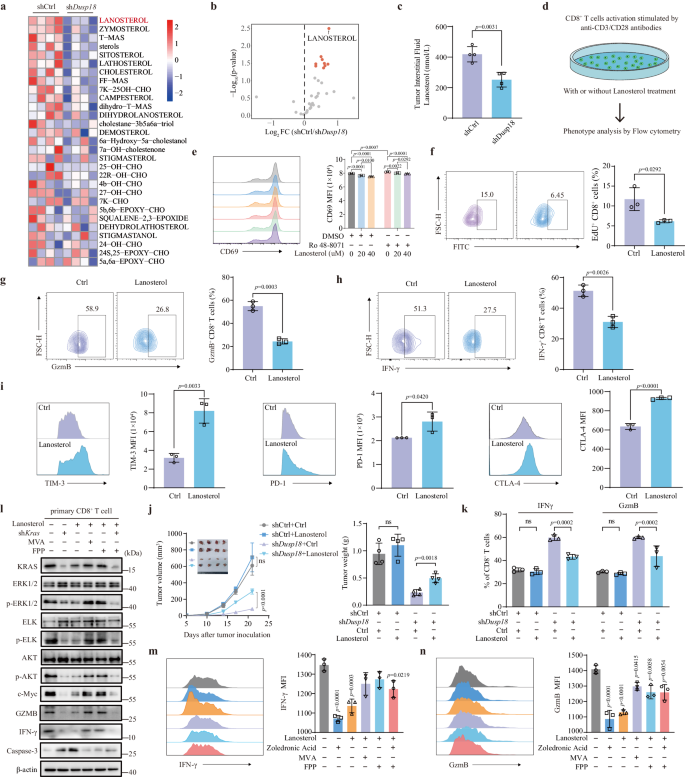
As a result of the previous work confirmed that DUSP18 and USF1 collaborate to upregulate ldl cholesterol biosynthesis, we explored the likelihood that a number of intermediates within the ldl cholesterol biosynthesis pathway may be liable for suppressing CD8+ T cell perform. We due to this fact decided the degrees of those intermediates in tumor interstitial fluid of shDusp18 and shCtrl MC38 tumors utilizing ldl cholesterol metabolomics. The degrees of ldl cholesterol itself, quite a few ldl cholesterol synthesis intermediates, oxysterols and different derivatives had been considerably decrease within the fluid obtained from shDusp18 tumors (Fig. 5a and Supplementary Information 5). Probably the most down-regulated of those (40%) was lanosterol, a little-studied ldl cholesterol synthesis intermediate (Fig. 5b, c). To find out the relevance of this straight, main CD8+ T cells had been handled with completely different concentrations of lanosterol and the consequence confirmed that lanosterol lowered expression of the CD8+ T cell activation markers CD69 in a dose dependent method (Fig. 5e). In one other set of experiments, we added Ro 48-8071, an inhibitor of lanosterol synthase (LSS) to dam endogenous lanosterol manufacturing. Ro 48-8071 remedy enhanced CD69 expression in CD8+ T cells, and a gradient of lanosterol was capable of lower CD69 expression when the CD8+ T cells with or with out the addition of Ro 48-8071 (Fig. 5e). Lanosterol remedy not solely led to a lower in CD25 expression (Supplementary Fig. 7a) but additionally successfully inhibited CD8+ T cell proliferation (Fig. 5f and Supplementary Fig. 7b). Moreover, it notably suppressed cytokine manufacturing, together with IFN-γ and GzmB (Fig. 5g, h), whereas concurrently selling the expression of immune checkpoint molecules comparable to PD-1, TIM-3, and CTLA-4 (Fig. 5i and Supplementary Fig. 7c), in addition to inducing mobile apoptosis (Supplementary Fig. 7d).
a Ldl cholesterol metabolism-related metabolites and absolute lanosterol concentrations in interstitial fluids from subcutaneous shDusp18 MC38 tumors in 4–6 weeks previous C57BL/6J mice (n = 4). b Volcano plot exhibiting variations in lanosterol and different metabolites in shCtrl and shDusp18 tumors. c Boxplot of absolute lanosterol concentrations in tumor interstitial fluid (n = 4). d Schematic illustration of CD8+ T cells phenotypes handled by lanosterol. e FACS evaluation of CD8+ T cell activation marker CD69 with indicated remedy (n = 3). f FACS evaluation of CD8+ T cell proliferation with lanosterol remedy. Quantification of IFN-γ (n = 3) (g) and GzmB (n = 3) (h) manufacturing in CD8+ T cells after lanosterol remedy. i Quantification of PD-1+ (n = 3), TIM-3+ (n = 3) and CTLA-4+ (n = 3) MFI in CD8+ T cells after lanosterol remedy. 1 × 106 shCtrl or shDusp18 MC38 cells had been subcutaneously injected into C57BL/6J mice (n = 4). Mice had been injected with 30 mg/kg of lanosterol each 2 days from day 0 till the time of sacrificed. Tumor progress curves and photographs of tumors are proven in (j); Move cytometric quantification of IFN-γ (left) and GzmB (proper) CD8+ T cells from completely different teams is proven in (ok) (n = 3). l The indicated protein ranges had been confirmed by Western blotting following the indicated remedies. The IB information are consultant of three unbiased experiments. Quantification of IFN-γ (m) and GzmB (n) MFI in CD8+ T cells following the indicated remedy (n = 3). Information are offered as imply ± SD (c, e–i, j, ok, m, n). P values had been calculated by unpaired two-tailed t checks (b, c, f–i, j, ok), one-way ANOVA (e, m, n); ns not important. Supply information are supplied as a Supply Information file.
Relative to manage CD8+ T cells, these handled with lanosterol confirmed impaired skill of their killing of MC38-OVA and B16-OVA tumor cells. As well as, the proportion of apoptotic tumor cells was considerably lowered within the lanosterol-treated group, as was LDH launch and the variety of viable tumor cells was considerably elevated (Supplementary Fig. 7e). To evaluate the impression of lanosterol on in vivo tumor progress, we subcutaneously inoculated equal numbers of shCtrl and shDusp18 MC38 cells. Notably, silencing Dusp18 resulted in a deceleration of xenograft tumor progress (Fig. 5j), accompanied by elevated ranges of IFN-γ and GzmB expression (Fig. 5k), and a lower within the share of PD-1+, TIM-3+, and CTLA-4+ CD8+ T cells (Supplementary Fig. 7f). Intriguingly, this impact was partially counteracted by intraperitoneal (i.p.) injection of lanosterol (30 mg/kg mice) (Fig. 5j, ok and Supplementary Fig. 7f).
Lanosterol has been reported to advertise ubiquitination and proteasomal degradation of HMG-CoA reductase (HMGCR), the rate-limiting enzyme within the mevalonate pathway51. In keeping with this, lanosterol lowered HMGCR protein ranges in a dose-dependent method each in mouse main CD8+ T cells and in human Jurkat T lymphoblastic leukemia cells (Supplementary Fig. 7g).
The mevalonate pathway is crucial for the synthesis of quite a lot of isoprenoids derived from acetyl-CoA, together with prenyl teams, that are important for the in vivo organic exercise of RAS proteins30. Lanosterol remedy considerably inhibited KRAS protein abundance in addition to ERK, and AKT signaling in mouse main CD8+ T cells, whereas activating apoptosis-associated caspase-3 cleavage as beforehand reported (Fig. 5l)52. Zoledronic Acid, an inhibitor of farnesyl pyrophosphate synthase, blocked the synthesis of farnesyl pyrophosphate (FPP) and geranylgeranyl pyrophosphate whereas lowering KRAS prenylation and attenuating downstream signaling. Like lanosterol, zoledronic acid additionally inhibited IFN-γ and GzmB manufacturing by CD8+ T cells. Supplementation with both mevalonic acid or FPP largely reversed the signaling inhibition mediated by each (Fig. 5m, n).
KRAS signaling-mediated T cell receptor (TCR) signaling activation and cell proliferation are stipulations for CD8+ T perform, which can clarify why lanosterol inhibits intratumoral CD8+ T cell perform. Collectively, all of the above outcomes counsel that tumor-derived lanosterol serves as an immune suppressive metabolite that limits the cytotoxic perform of CD8+ T cells by lowering HMGCR protein degree and thereby impairing KRAS-ERK signaling.
DUSP18 is overexpressed in CRC sufferers and predicts immune deserts
To additional examine the scientific relevance of the above-described DUSP18-USF1-SREBP2 axis, 20 pairs of human CRC samples (T) and adjoining regular colon tissues (N) had been collected and examined for expression of the above proteins and a number of other others. Usually, the degrees of DUSP18, USF1, SREBP2, HMGCR and LSS had been considerably upregulated in CRC samples (Fig. 6a and Supplementary Fig. 8a). All of those besides USF1 additionally positively correlated with DUSP18. Whereas USF1 phosphorylation negatively correlated with DUSP18 and was considerably decreased in CRC samples (Fig. 6b). DUSP18 mRNA expression was additionally upregulated in a number of colorectal most cancers GEO datasets and another most cancers sorts (Fig. 6c). Extra research revealed that CRCs contained excessive ranges of DUSP18 protein (Supplementary Fig. 8b). Receiver Working Attribute Curve evaluation indicated that elevated DUSP18 expression was an efficient estimate customary of CRC affected person’s survival (Fig. 6d). The DUSP18 gene additionally tended to be hypomethylated in CRC samples, thus explaining the premise for its overexpression (Supplementary Fig. 8c, d). Lastly, excessive DUSP18 expression additionally positively correlated with scientific and TMN stage (Supplementary Fig. 8e).
a Relative protein degree of DUSP18, USF1, pSREBP2, nSREBP2, HMGCR and LSS (tumor samples dimension: 20, paired regular samples dimension: 20). Sign depth of those proteins was quantified by Picture J, after which normalized to β-actin band depth. b Correlation between the expression ranges of DUSP18 and USF1, pSREBP2, nSREBP2, HMGCR and LSS, as decided by Pearson’s r evaluation. c Differential expression evaluation for DUSP18 in tumor and regular tissues. LIHC, liver hepatocellular carcinoma (tumor samples (n = 374), regular samples (n = 50)); KIRP, kidney renal papillary cell carcinoma (tumor samples (n = 289), regular samples (n = 32)); KIRC kidney renal clear cell carcinoma (tumor samples (n = 535), regular samples (n = 72)); CHOL, cholangiocarcinoma (tumor samples (n = 36), regular samples (n = 9)); STAD, abdomen adenocarcinoma (tumor samples (n = 375), regular samples (n = 32)). For GSE23878 (tumor samples (n = 35), regular samples (n = 24)); GSE35279 (tumor samples (n = 74), regular samples (n = 5)); GSE32323 (tumor samples (n = 17), regular samples (n = 17)); GSE25070 (tumor samples (n = 26), regular samples (n = 26)); GSE146587 (tumor samples (n = 29), regular samples (n = 30)). d ROC plot exhibiting the AUC of the DUSP18 expression from TCGA-COAD (n = 473) and GSE23878 (n = 35) databases. e UMAP plot from excessive (left, n = 15 samples) or low (proper, n = 15 samples) DUSP18 expression sufferers. f Bar plot of proportional variations in immune cells between the DUSP18excessive (n = 15 samples) and DUSP18low (n = 15 samples) teams. g Re-clustering of CD8+ T lymphocytes, UMAP visualization and marker-based annotation of CD8+ T lymphocyte subtypes, coloured by cluster identification (n = 12528 cells). h Bar plot of proportional variations in CD8+ T lymphocytes between the DUSP18excessive (n = 2264 cells) and DUSP18low (n = 2804 cells) teams. CD8_T_ reminiscence, reminiscence CD8+ T cells; CD8_T_ naive, naive CD8+ T cells; CD8_T_Tex, exhausted CD8+ T cells; CD8_T_Teff, effector CD8+ T cells; CD8_T_Trm, tissue-resident reminiscence CD8+ T cells. i Enrichment of various gene signature scores between the DUSP18excessive (n = 2264 cells) and DUSP18low (n = 2804 cells) teams in single-cell transcriptomes from re-clustered CD8+ T cells. Information are offered as imply ± SD (a). P values had been calculated by unpaired two-tailed t-tests (a, c), modified Fisher’s actual checks (i). P values and R had been calculated by Pearson’s correlation evaluation. Two-sided P worth was given (b); ns, not important. Supply information are supplied as a Supply Information file.
To additional discover the connection between DUSP18 and the TME of CRC sufferers, single-cell RNA sequencing information had been analyzed from 62 CRC sufferers (Supplementary Fig. 6f). CD8+ T cell signatures had been considerably elevated in tumors with low DUSP18 expression (Fig. 6e, f and Supplementary Fig. 8g) and DUSP18 expression negatively correlated with the variety of CD8+ T cells (Supplementary Fig. 8h, i).
Primarily based on classical markers, CD8+ T cells had been reclassified into 5 subpopulations: exhausted T cells (Tex), effector T cells (Teff), tissue-resident reminiscence T cells, naive T cells, and reminiscence T cells (Fig. 6g). The proportion of Teffs was dramatically upregulated in CRCs with low DUSP18 expression, whereas the relative ratio of Texs was decrease (Fig. 6h). Gene signatures of CD8+ T cells additionally revealed that the IFN-γ and TCR signaling pathways had been all enriched in low DUSP18 expression tumors (Fig. 6i).
Beforehand reported immune-related gene signatures calculated by the single-sample gene-set enrichment evaluation (ssGSEA) algorithm53, had been used to characterize immune-related indicators based mostly on the DUSP18 expression ranges within the TCGA-COAD dataset. Tumor samples with excessive DUSP18 expression had decrease immune-related indicators scores (Supplementary Fig. 9a–d), decrease tumor mutation burden, microsatellite instability, tumor neoantigens and mutant-allele tumor heterogeneity (Supplementary Fig. 9e). In distinction tumor samples with low DUSP18 expression confirmed increased enrichment scores of T cell inflammatory gene expression profile, innate anti-PD-1 resistance, immuno-predictive rating signatures and PD-1 response signatures (Supplementary Fig. 9f). As well as, DUSP18 was extremely related to a lot of the antitumor immunity course of (Supplementary Fig. 9g).
To additional validate DUSP18’s function in regulating tumor immunity in CRC on the transcriptional degree, GSEA, GO, and KEGG analyses had been utilized to TCGA-COAD information. The outcomes confirmed that low DUSP18-expressing tumor samples had been considerably enriched for immune-related features pertaining to antigen presentation, chemokine signaling, T-cell receptor signaling (Supplementary Fig. 10a-c), whereas high-expressing samples had been enriched for ldl cholesterol biosynthetic pathways (Supplementary Fig. 10d). The usage of a number of algorithms confirmed that DUSP18 expression negatively correlated with infiltrations by CD8+ T cells and quite a lot of different immune cells (Supplementary Figs. 10e, f and 11). These findings present that DUSP18 was considerably upregulated in human CRC samples and was predictive of much less immune cell infiltration and the non-inflammatory TMEs.
Lumacaftor is a potent DUSP18 inhibitor
On condition that DUSP18 performs an necessary function in tumor immune evasion, we tried to establish DUSP18 inhibitors to make use of as potential CRC therapies. Using a digital display screen based mostly on the AutoDock4 algorithm, the putative DUSP18 inhibitor Lumacaftor was recognized as having the best binding vitality to DUSP18 from a library of about two thousand FDA-approved medicine within the ZINC15 compound library (Fig. 7a, b). Lumacaftor has been beforehand used within the remedy of cystic fibrosis the place it normalizes the trafficking of some mutant CFTR proteins to the outer membrane54. Primarily based on the crystal construction DUSP18, nonetheless, our molecular docking examine confirmed that Lumacaftor interacts with DUSP18 residues Ala105, Ala106, Ser109, Arg110 and Arg142 (Supplementary Fig. 12a). In help of this mannequin, the outcomes of Microscale Thermophoresis (MST) confirmed Lumacaftor to have a stronger affinity to WT DUSP18 than to DUSP18 containing mutations in any of the above residues (Fig. 7c). In mobile thermal shift assays (CETSAs), Lumacaftor stabilized the thermal denaturation of the DUSP18 protein in each HCT116 and SW480 cells, thereby indicating that DUSP18 is a direct goal of Lumacaftor (Fig. 7d). Lumacaftor additionally binds to DUSP18 residues which might be important for its phosphatase exercise. Unsurprisingly, Lumacaftor remedy lowered the protein and mRNA ranges of genes within the ldl cholesterol biosynthesis, whereas having virtually no impact on DUSP18 protein (Fig. 7e, f and Supplementary Fig. 12b, c). In vitro dephosphorylation experiments confirmed that Lumacaftor inhibited the power of purified DUSP18 to dephosphorylate WT USF1 however had no impact on USF1T100A (Fig. 7g). As well as, Lumacaftor remedy in shDUSP18 CRC cell strains didn’t additional down-regulate the protein ranges of USF1 and SREBP2, indicating that the impact of Lumacaftor is DUSP18-dependent (Fig. 7h).
a Affinity rank of small molecules with DUSP18 in response to digital screening. b Chemical construction system of Lumacaftor. c The equilibrium dissociation fixed (Okayd) worth was decided because the binding of Lumacaftor to purified human DUSP18 or DUSP18 mutant (A105/106D, S109D, R110D and R142D) proteins utilizing MST (n = 3). d Impact of Lumacaftor on the thermal denaturation of mobile DUSP18 protein. CETSA was carried out on cell lysates from HCT116 and SW480 cell strains. e Western blot for the indicated proteins in MC38 and B16-OVA cell strains following publicity to 40 μM Lumacaftor for twenty-four h. f mRNA ranges of indicated genes from MC38 and B16-OVA cells handled by management or 40 μM Lumacaftor for twenty-four h had been analyzed utilizing RT-qPCR (n = 3). g Recombinant GST-DUSP18-His and Flag-USF1 (WT or T100A) had been used for in vitro dephosphorylation assay, with or with out the addition of 40 μM Lumacaftor. The degrees of USF1 phosphorylation had been measured by immuno-blotting evaluation. h Lumacaftor was added to shCtrl or shDUSP18 HCT116 and SW480 cells, and the associated protein ranges had been detected by immunoblot. i Quantification of apoptosis in MC38-OVA and B16-OVA cells that had been pretreated with 40 μM Lumacaftor for twenty-four h and co-cultured with CD8+ T cells for twenty-four h (n = 3). j Imply Fluorescence Depth (MFI) quantification of PD-1, TIM-3, CTLA-4 in CD8+ T cells co-cultured with MC38 and B16-OVA cells that had been pretreated with 40 μM Lumacaftor for twenty-four h (n = 3). ok Quantification of cytokine manufacturing by CD8+ T cells co-cultured with MC38 and B16-OVA cells pretreated with 40 μM Lumacaftor for twenty-four h (n = 3). Information are offered as imply ± SD (f, i–ok). P values had been calculated by unpaired two-tailed t-tests (f, i–ok); ns not important. All IB information are consultant of three unbiased experiments. Supply information are supplied as a Supply Information file.
Much like the consequences of Dusp18 inhibition, Lumacaftor exerted minimal toxicity on cells (Supplementary Fig. 12d–f). Enhanced launch of cytokines, elevated cytotoxic capability and decreased inhibitory molecules had been present in CD8+ T cells co-cultured with the tumor cells with Lumacaftor remedy (Fig. 7i–ok). General, Lumacaftor was recognized as a potent inhibitor of DUSP18 phosphatase exercise, of the DUSP18-USF1-SREBP2 TF cascade and of ldl cholesterol biosynthesis in CRC cells, with little proof of toxicity.
Inhibition of DUSP18 with Lumacaftor sensitizes most cancers cells to ICB remedy
On condition that Lumacaftor can inhibit DUSP18 exercise and improve the antitumor perform of CD8+ T cells in vitro, the antitumor impact of Lumacaftor had been subsequently assessed on tumor fashions in vivo. The syngeneic mouse MC38 CRC mannequin was utilized to look at how Lumacaftor, with or with out the addition of anti-PD-1 antibody, affected tumor progress and survival. Strikingly, mixture remedy considerably suppressed tumor progress and extended survival of MC38 tumor-bearing immunocompetent C57BL/6 J mice in comparison with both single-agent or control-treated group (Fig. 8a–c). Evaluation of tumor-associated immune cells demonstrated that the mix of Lumacaftor and anti-PD-1 remedy considerably elevated the proportion of CD8+ T cells however had no important impact on CD4+ T cells (Fig. 8d, e). Mixed remedy additionally considerably elevated the expression of GzmB and IFN-γ by these CD8+ T cells (Fig. 8f, g). Comparable outcomes had been additionally noticed in B16-OVA tumor-bearing immunocompetent C57BL/6J mice with the above-combined remedy (Fig. 8h–j). Move cytometric evaluation confirmed that mixture remedy considerably enhanced CD8+ T cell infiltration in tumor tissue and the expression of GzmB and IFN-γ (Fig. 8k–m). These information had been concordant with the outcomes obtained with MC38 tumor research and strengthened the findings that DUSP18 inhibition with Lumacaftor represses tumor immune evasion and enhances responses to immunotherapy.
Therapeutic impact of Lumacaftor on tumorigenesis of MC38 cells or B16-OVA cells in C57BL/6J mice. 4–6 weeks C57BL/6J mice had been injected i.p. with 30 mg/kg Lumacaftor on days 7, 10, and 13, and 100 μg anti-PD-1 on days 7 and 10 following subcutaneous injection of tumor cells. a (n = 6), h (n = 7), Tumor progress curves. b (n = 6), i (n = 7), Tumor weights. c (n = 6), j (n = 7), Survival curves. d (n = 3), ok (n = 3), Proportion of tumor-infiltrating CD8+ T cells decided by movement cytometry. e Proportion of tumor-infiltrating CD4+ T cells (n = 3). f (n = 3), l (n = 3), Proportion of IFN-γ+ CD8+ T cells within the TME. g (n = 3), m (n = 3), Proportion of GzmB+ CD8+ T cells within the TME. n Schematic mannequin of the regulatory pathway and mechanism of DUSP18 in tumor immune evasion. Information are offered as imply ± SD (a–m). P values had been calculated by unpaired two-tailed t checks (a, b, d, f, g, h, i, ok–m), one-way ANOVA (e) or log-rank (Mantel-Cox) take a look at (c, j). ns not important. Supply information are supplied as a Supply Information file.