Our analysis complies with all related moral rules and has been accredited by the committee of ethics of the analysis of the College of Navarra and follows their prescribed moral tips.
Cell tradition
KASUMI-1 (ACC 220, DSMZ), HL-60 (ACC 3, DSMZ), NB-4 (ACC 207, DSMZ), OCI-AML3 (ACC 582, DSMZ), MV4-11 (ACC 102, DSMZ), MOLM-13 (ACC 554, DSMZ), HEL (ACC 11, DSMZ), GF-D8 (ACC 615, DSMZ), TF-1 (CRL-2003, ATCC), THP-1 (TIB-202, ATCC) and M-O7e (ACC 104, DSMZ) cell strains have been cultured in RPMI-1640 medium supplemented with 20% fetal bovine serum (FBS). GM-CSF was added to GF-D8 (50 ng/µL), TF-1 (50 ng/µL) and THP-1 (10 ng/µL) tradition medium. M-O7e cells have been additionally supplemented with 10 ng/µL of IL-3. OCI-AML2 (ACC 99, DSMZ) cells have been cultured with DMEM medium + 20% FBS. The MONO-MAC-6 (ACC 124, DSMZ) cell line was maintained in RPMI-1640 medium supplemented with 20% FBS, 2 mM L-glutamine, non-essential amino acids, 1 mM sodium pyruvate, and 10 µg/ml human insulin. The UT-7 (ACC 137, DSMZ) cell line was cultured with Alpha-MEM medium supplemented with ribo- and deoxyribonucleosides, 20% FBS, and 5 ng/ml GM-CSF. All cell strains have been maintained at 37 °C in a moist environment containing 5% CO2. The cell strains have been obtained from the DSMZ or the American Sort Tradition Assortment (ATCC). All cell strains have been authenticated by performing a brief tandem repeat allele profile and have been examined for mycoplasma (MycoAlert Pattern Package, Cambrex).
Compounds
Panobinostat (HY-10224-CS-0267), Quisinostat (HY-15433) and Molibresib (HY-13032) have been bought from MedChemExpress (Monmouth Junction, NJ, USA). Entinostat (S1053) and JQ1 (S7110) have been obtained from Selleck Chemical substances (Houston, TX, USA). Vorinostat (10009929), Tubastatin (6270) and Retinoic acid (R2625) have been bought from Cayman Chemical (MI, USA), Tocris (Germany) and Sigma-Aldrich (St Louis, Missouri, USA), respectively.
Synthesis of compounds CM-444 and CM-1758
Common chemistry info
Except in any other case famous, all beginning supplies, reagents, and solvents have been bought from industrial suppliers and used with out additional purification. Air-sensitive reactions have been carried out underneath N2. The NMR spectroscopic knowledge have been recorded on a Bruker AV400 or VARIAN 400MR spectrometer with customary pulse sequences. Chemical shifts (δ) are reported in elements per million (ppm). The abbreviations used to elucidate multiplicities are s = singlet, d = doublet, t = triplet, q = quadruplet, m = multiplet and br s = broad singlet. The coupling fixed, J, is reported in Hertz items. Melting factors have been decided on a Mettler FP82 scorching stage managed by a Mettler FP80 central processor.
Protocol for analytical excessive efficiency liquid chromatography (HPLC)
The purity of ultimate compounds was measured by HPLC. HPLC-analysis was carried out utilizing a Shimadzu LC-20AB with a Luna-C18(2), 5 µm, 2.0 × 50 mm column at 40 °C with a diode-array detector. Solvent A: water with 0.037% trifluoroacetic acid; Solvent B: acetonitrile with 0.018% trifluoroacetic acid. Gradient: After 0.01 min on the preliminary situation of 90% A and 10% B, Solvent B was elevated to 80% over 4 min, maintained at 80% for 0.9 min, after which a linear gradient to preliminary circumstances was utilized for 0.02 min and maintained for 0.58 min to re-equilibrate the column, giving a cycle time of 5.50 min. The stream fee was 0.8 mL/min from 0.01 to 4.90 min, elevated to 1.2 mL/min in 0.03 min, and maintained till the tip of the run.
Protocols for preparative HPLC purification strategies
The HPLC measurement was carried out utilizing a SHIMADZU preparative HPLC system, an autosampler, and a UV detector. The fractions have been detected by LC-MS. The MS detector was configured with an electrospray ionization supply. The supply temperature was maintained at 300 °C–350 °C. Reversed-phase HPLC was carried out on a Luna C18 column (250×50 mm; 10 µm). Solvent A: water with 0.1% trifluoroacetic acid (TFA); Solvent B: acetonitrile.
Technique 1: Gradient: At room temperature, 25% to 55% B over 20 min; then 100% B over 15 min. Stream fee: 80 mL/min.
Technique 2: Gradient: At room temperature, 10% to 40% B over 20 min; then 100% B for 15 min. Stream fee: 80 mL/min.
Excessive-resolution mass spectrometry of ultimate compounds
Excessive-resolution mass spectrometry (HRMS) m/z was carried out on an Agilent Applied sciences 1200 liquid chromatographic system geared up with a 6220 Correct-Mass time of flight (TOF) LC/MS, operated in optimistic electrospray ionization mode (ESI+) managed by MassHunter Workstation 06.00 software program (Agilent Applied sciences, Barcelona, Spain). The separation was carried out on a Zorbax SB-C18 (15 cm × 0.46 cm; 5 μm) from Agilent Applied sciences with a SB-C18 precolumn from Teknokroma (Barcelona, Spain). Solvent A: water with 0.1% formic acid; Solvent B: acetonitrile. The gradient elution was 5–100% B from 0–25.0 min; 100% B from 25–26 min; 100–5% B from 26–28 min; and 5% B from 28–30 min. The injection quantity was 10 μL, and the stream fee was 0.5 mL min−1 in the course of the full run. Chromatography was carried out at 40 °C. The ESI circumstances have been as follows: gasoline temperature, 350 °C; drying gasoline, 10 L min−1; nebulizer, 45 psig; capillary voltage, 3500 V; fragmentor, 175 V; and skimmer, 65 V. Acquisition was from 100–1000 m/z at a fee of 1.03 spectra s−1. All solvents used have been liquid chromatography–mass spectrometry (LC-MS)-grade and obtained from Scharlau (Scharlab, Sentmenat, Spain). Water (18.2 MΩ) was obtained from an Ultramatic system from Wasserlab (Barbatáin, Spain).
The artificial means of the small molecules CM-444 and CM-1758 is printed in Supplementary Fig. 10.
Synthesis of two,4-Dichloro-6,7-dimethoxy-quinoline (2)
A combination of commercially accessible 3,4-dimethoxyaniline (1) (20.00 g, 130.57 mmol) and malonic acid (20.38 g, 195.85 mmol, 20.38 mL) in POCl3 (100 mL) was degassed and purged with N2 thrice, after which the combination was stirred at 100 °C for 16 h. The combination was cooled to twenty °C and concentrated in diminished strain at 40 °C. The residue was poured into H2O (300 mL) and extracted with DCM (200 mL × 3). The mixed natural part was washed with brine (300 mL × 2), dried with anhydrous Na2SO4, filtered and concentrated underneath a vacuum. The residue was purified by silica gel column chromatography (petroleum ether/EtOAc = 20:1 to 1:1) to afford pure compound 2 (20.00 g, 59%) as a yellow strong. 1H NMR (CDCl3, 400 MHz): δ 7.38 (s, 1H), 7.37–7.36 (m, 2H), 4.06 (s, 3H), 4.03 (s, 3H). ESI-MS m/z calcd. for C11H9Cl2NO2: 257.0, discovered 258.0 [M + H]+.
Synthesis of 4-Chloro-6,7-dimethoxy-2-(5-methyl-2-furyl)quinoline (3a)
A combination of compound 2 (5 g, 19.37 mmol), commercially accessible 4,4,5,5-tetramethyl-2-(5-methyl-2-furyl)-1,3,2-dioxaborolane (4.03 g, 19.37 mmol), Okay2CO3 (5.35 g, 38.74 mmol), Pd(PPh3)4 (1.12 g, 968.62 μmol) in 1,4-dioxane/H2O (8:1, 90 mL) was degassed and purged with N2 thrice, after which the combination was stirred at 100 °C for 12 h. The combination was filtered, and the filtrate was concentrated underneath vacuum. The residue was purified by flash silica gel chromatography (ISCO®; 80 g SepaFlash® Silica Flash Column, eluent of 0%–40% EtOAc/petroleum ether gradient at 100 mL/min) to afford pure compound 3a (4.20 g, 71%) as a light-yellow strong. 1H NMR (CDCl3, 400 MHz): δ 7.74 (s, 1H), 7.46 (s, 1H), 7.38 (s, 1H), 7.04 (d, J = 3.1 Hz, 1H), 6.18 (d, J = 2.4 Hz, 1H), 4.07 (s, 3H), 4.05 (s, 3H), 2.46 (s, 3H). ESI-MS m/z calcd. for C16H14ClNO3: 303.1 discovered 304.1 [M + H]+.
Synthesis of Ethyl 2-[4-[[[6,7-dimethoxy-2-(5-methyl-2-furyl)-4-quinolyl]amino]methyl]-1-piperidyl]pyrimidine-5-carboxylate (4a)
A combination of compound 3a (1 g, 3.29 mmol), commercially accessible ethyl 2-[4-(aminomethyl)-1-piperidyl]pyrimidine-5-carboxylate (1.13 g, 4.28 mmol), Cs2CO3 (3.22 g, 9.88 mmol), BINAP (615.00 mg, 987.69 μmol) and Pd2(dba)3 (602.96 mg, 658.46 μmol) in 1,4-dioxane (30 mL) was degassed and purged with N2 thrice, after which the combination was stirred at 120 °C for 12 h. The combination was filtered and the filtrate was concentrated underneath vacuum. The residue was purified by flash silica gel chromatography (ISCO®; 40 g SepaFlash® Silica Flash Column, eluent of 0%–100% EtOAc/Petroleum ether gradient at 100 mL/min) to afford pure compound 4a (1.5 g, 86%) as a light-yellow strong. The purity was 97.21% in keeping with the HPLC analytical technique (described above), and the retention time (tR) was 2.948 min. 1H NMR (MeOD, 400 MHz): δ 8.81 (s, 2H), 7.71 (s, 1H), 7.53 (d, J = 3.4 Hz, 1H), 7.46 (s, 1H), 7.01 (s, 1H), 6.44 (d, J = 2.8 Hz, 1H), 4.99 (d, J = 13.4 Hz, 2H), 4.34 (q, J = 7.2 Hz, 2H), 4.05 (d, J = 3.0 Hz, 6H), 3.59 (d, J = 6.9 Hz, 2H), 3.05 (t, J = 12.0 Hz, 2H), 2.53 (s, 3H), 2.26 (br s, 1H), 2.01 (d, J = 11.8 Hz, 2H), 1.40–1.36 (m, 5H). ESI-MS m/z for C29H33N5O5 calcd.: 531.2, discovered: 532.4 [M + H]+.
Synthesis of 2-[4-[[[6,7-dimethoxy-2-(5-methyl -2-furyl)-4-quinolyl]amino]methyl]-1-piperidyl]pyrimidine-5-carboxylic acid (5a)
A combination of compound 4a (1.5 g, 2.82 mmol) and LiOH•H2O (591.98 mg, 14.11 mmol) in THF/H2O (2:1, 30 mL) was stirred at 25 °C for 12 h. Then, the response combination was adjusted to pH 5 with aqueous HCl (2.0 M) at room temperature, and the precipitate was filtered to afford compound 5a (1.2 g, 84%) as a yellow strong. The purity was 97.20% in keeping with the HPLC analytical technique (described above); and the tR was 2.373 min. 1H NMR (MeOD, 400 MHz): δ 8.80 (s, 2H), 7.69 (s, 1H), 7.51 (d, J = 3.4 Hz, 1H), 7.44 (s, 1H), 6.99 (s, 1H), 6.43 (d, J = 3.3 Hz, 1H), 4.97 (d, J = 13.1 Hz, 2H), 4.03 (s, 3H), 4.02 (s, 3H), 3.57 (d, J = 7.2 Hz, 2H), 3.03 (t, J = 11.7 Hz, 2H), 2.51 (s, 3H), 2.24 (br s, 1H), 1.99 (d, J = 12.7 Hz, 2H), 1.41–1.32 (m, 2H). ESI-MS m/z calcd. for C27H29N5O5: 503.2, discovered 504.3 [M + H]+.
Synthesis of 2-[4-[[[6,7-dimethoxy-2-(5-methyl-2-furyl)-4-quinolyl]amino]methyl]-1-piperidyl]pyrimidine-5-carbohydroxamic acid (6a,
CM-444)
A combination of compound 5a (1.2 g, 2.38 mmol), O-tetrahydropyran-2-ylhydroxylamine (558.34 mg, 4.77 mmol), HOBt (644.01 mg, 4.77 mmol), EDCI (913.68 mg, 4.77 mmol) and DIEA (1.54 g, 11.92 mmol, 2.08 mL) in DMF (30 mL) was degassed and purged with N2 thrice, after which the combination was stirred at 25 °C for 12 h. Subsequent, 0.5 M aqueous HCl (5 mL) was added in a single portion at room temperature. The combination was concentrated underneath vacuum to offer a residue, which was purified by preparative HPLC (Common process described above, technique 1) to afford pure compound 6a (CM-444, 794.3 mg, 62%) as a light-yellow strong, m.p. 136 °C–137 °C. The purity was 96.65% in keeping with the HPLC analytical technique (described above), and the tR was 2.077 min. 1H NMR (MeOD, 400 MHz): δ 8.65 (s, 2H), 7.69 (s, 1H), 7.51 (d, J = 3.5 Hz, 1H), 7.44 (s, 1H), 6.99 (s, 1H), 6.43 (d, J = 3.3 Hz, 1H), 4.94 (br s, 2H), 4.03 (s, 3H), 4.02 (s, 3H), 3.57 (d, J = 7.2 Hz, 2H), 3.01 (t, J = 11.7 Hz, 2H), 2.51 (s, 3H), 2.23 (br s, 1H), 1.98 (d, J = 11.9 Hz, 2H), 1.40–1.35 (m, 2H). 13C NMR (DMSO-d6, 100 MHz): δ 161.2, 157.0 (2 C), 156.7, 154.0, 153.8, 148.6, 144.0, 139.1, 134.3, 116.1, 114.1, 109.9 (2 C), 102.2, 100.3, 91.7, 56.3, 55.9, 47.9, 43.3 (2 C), 35.5, 29.3 (2 C), 13.6. HRMS m/z: [(M + H)]+ calcd. for C27H30N6O5, 519.2350; discovered, 519.2400.
Synthesis of 4-chloro-2-(2,5-dimethyl-3-furyl)-6,7-dimethoxy-quinoline (3b)
A combination of compound 2 (2.00 g, 7.75 mmol), commercially accessible 2-(2,5-dimethyl-3-furyl)-4,4,5,5-tetramethyl-1,3,2-dioxaborolane (1.72 g, 7.75 mmol), Okay2CO3 (2.14 g, 15.50 mmol), Pd(PPh3)4 (895.43 mg, 775.00 μmol) in 1,4-dioxane/H2O (10:1, 55 mL) was degassed and purged with N2 thrice, after which the combination was stirred at 90 °C for 16 h. The response combination was concentrated underneath vacuum to offer a residue. The residue was purified by silica gel column chromatography (petroleum ether/EtOAc = 20:1 to five:1) to afford pure compound 3b (1.9 g, 77%) as a white strong. 1H NMR (CDCl3, 400 MHz): δ 7.46 (s, 1H), 7.39–7.36 (m, 2H), 6.39 (s, 1H), 4.06 (m, 6H), 2.69 (s, 3H), 2.31 (s, 3H). ESI-MS m/z calcd. for C17H16ClNO3: 317.1, discovered 318.1 [M + H]+.
Synthesis of ethyl 2-[4-[[[2-(2,5-dimethyl-3-furyl)-6,7-dimethoxy-4-quinolyl] amino]methyl]-1-piperidyl]pyrimidine-5-carboxylate (4b)
A combination of compound 3b (1 g, 3.15 mmol), ethyl 2-[4-(aminomethyl)-1-piperidyl]pyrimidine-5-carboxylate (1.08 g, 4.09 mmol), Cs2CO3 (3.08 g, 9.44 mmol), BINAP (587.86 mg, 944.09 μmol) and Pd2(dba)3 (576.35 mg, 629.39 μmol) in 1,4-dioxane (30 mL) was degassed and purged with N2 thrice, after which the combination was stirred at 120 °C for 12 h. The combination was filtered and the filtrate was concentrated underneath vacuum. The residue was purified by flash silica gel chromatography (ISCO®; 40 g SepaFlash® Silica Flash Column, eluent of 0%–100% EtOAc/petroleum ether gradient at 100 mL/min) to afford pure compound 4b (1.2 g, 70%) as a light-yellow strong. 1H NMR (CDCl3, 400 MHz): δ 8.84 (s, 2H), 7.37 (s, 1H), 6.87 (br s, 1H), 6.51 (br s, 1H), 6.38 (br s, 1H), 4.99 (d, J = 13.2 Hz, 2H), 4.73 (br s, 1H), 4.35 (q, J = 7.1 Hz, 2H), 4.03 (s, 6H), 3.32 (t, J = 6.2 Hz, 2H), 2.99 (t, J = 11.8 Hz, 2H), 2.65 (s, 3H), 2.32 (s, 3H), 2.14 (br s, 1H), 2.00 (d, J = 11.5 Hz, 2H), 1.41–1.32 (m, 5H). ESI-MS m/z calcd. for C30H35N5O5: 545.3, discovered 546.3 [M + H]+.
Synthesis of 2-[4-[[[2-(2,5-dimethyl-3-furyl)- 6,7-dimethoxy-4-quinolyl]amino]methyl]-1-piperidyl]pyrimidine-5-carboxylic acid (5b)
A combination of compound 4b (1.2 g, 2.20 mmol) and LiOH•H2O (461.41 mg, 11.00 mmol) in THF/H2O (2:1, 30 mL) was stirred at 25 °C for 12 h. Then, the response combination was adjusted to pH 5 with aqueous HCl (2.0 M) at room temperature, and the precipitate was filtered to afford compound 5b (1.1 g, 97%) as a yellow strong. ESI-MS m/z calcd. for C28H31N5O5: 517.2, discovered 518.3 [M + H]+.
Synthesis of 2-[4-[[[2-(2,5-dimethyl-3-furyl)-6,7-dimethoxy-4-quinolyl]amino]methyl] -1-piperidyl]pyrimidine-5-carbohydroxamic acid (6b,
CM-1758)
A combination of compound 5b (1.1 g, 2.13 mmol), O-tetrahydropyran-2-ylhydroxylamine (497.94 mg, 4.25 mmol), HOBt (574.34 mg, 4.25 mmol), EDCI (814.84 mg, 4.25 mmol) and DIEA (1.37 g, 10.63 mmol, 1.85 mL) in DMF (15 mL) was degassed and purged with N2 thrice, after which the combination was stirred at 25 °C for 12 h. The residue was poured into ice water (w/w = 1/1) (30 mL). The aqueous part was extracted with ethyl acetate (20 mL ×3). The mixed natural part was washed with brine (30 mL) and dried with anhydrous Na2SO4. Then, aqueous HCl (2 M, 3 mL) was added in a single portion at room temperature and concentrated underneath vacuum to offer a residue, which was purified by preparative HPLC (Common process described above, technique 2) to afford pure compound 6b (CM-1758, 503.7 mg, 43%) as an off-white strong. m.p. 176 °C–177 °C. The purity was 96.29% in keeping with the HPLC analytical technique (described above), and the tR was 2.073 min. 1H NMR (MeOD, 400 MHz): δ 8.65 (s, 2H), 7.71 (s, 1H), 7.32 (s, 1H), 6.68 (s, 1H), 6.47 (s, 1H), 4.92 (d, J = 13.2 Hz, 2H), 4.03 (s, 6H), 3.53 (d, J = 7.1 Hz, 2H), 2.99 (t, J = 11.9 Hz, 2H), 2.56 (s, 3H), 2.35 (s, 3H), 2.21 (br s, 1H), 1.96 (d, J = 11.0 Hz, 2H), 1.38–1.30 (m, 2H). 13C NMR (DMSO-d6, 100 MHz): δ 161.2, 157.0 (2 C), 154.1, 153.9, 151.4, 150.6, 148.8, 144.3, 134.8, 115.3, 114.2, 109.8, 106.3, 102.1, 100.2, 96.3, 56.3, 56.0, 47.8, 43.3 (2 C), 35.2, 29.3 (2 C), 13.3, 12.9. HRMS m/z: [(M + H)]+ calcd. for C28H32N6O5, 533.2507 discovered, 533.2549.
1H-NMR of CM-444
HPLC hint of CM-444
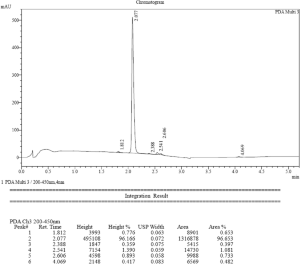
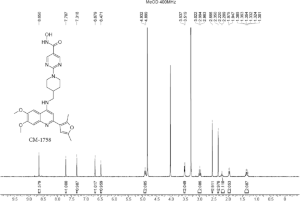
1H-NMR of CM-1758
HPLC hint of CM-1758
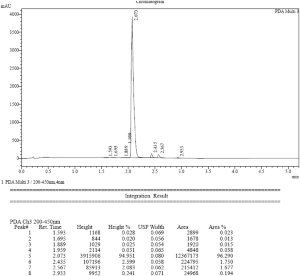
Cell proliferation assay: GI50 calculation
Cell proliferation was analyzed utilizing the CellTiter 96 Aqueous One Resolution Cell Proliferation Assay (Promega, Madison, WI). This can be a colorimetric technique for figuring out the variety of viable cells in proliferation. For the assay, suspension cells have been cultured in triplicate at a density of 1 × 106 cells/mL in 96-well plates (100.000 cells/properly, 100 µL/properly), utilizing solely the 60 interior wells to keep away from any border results. After 48 h of therapy with completely different doses of every compound (0, 0.001, 0.01, 0.1, 0.2, 0.4, 0.6, 0.8, 1, 2, 3, 4, 5 and 10 µM), the plates have been centrifuged at 800 g for 10 min adopted by elimination of the medium. Then, the cells have been incubated with 100 μL/properly of medium and 20 μL/properly of CellTiter 96 Aqueous One Resolution reagent for 1–3 h. The absorbance was recorded at 490 and 650 nm as reference wavelengths utilizing 96-well plate readers till the absorbance of the management cells with out therapy was roughly 0.8 AU. The background absorbance was measured in wells with solely cell line medium and answer reagent. First, the typical of the absorbance from the management wells was subtracted from all different absorbance values. Information have been calculated as the share of whole absorbance of handled cells/absorbance of non-treated cells. The GI50 values of the completely different compounds have been decided utilizing non-linear regression plots made with GraphPad Prism v5 software program.
Stream cytometry
For CD11b / annexin-V detection, 100,000 cells of ML-2, KASUMI-1, HL-60, NB-4, OCI-AML3, MV4-11, MOLM-13, HEL, GF-D8, TF-1, THP-1, M-O7e, OCI-AML2, MONO-MAC-6 and UT-7 cell strains have been cultured at a density of 1 × 106 cells/mL and handled each day for 48 h with CM-444, CM-1758, or ATRA (see Supplementary Desk 2 for GI50 knowledge). The cell strains HL-60, ML-2, MV4-11 and MOLM-13 have been additionally handled each day as much as 8 days with CM-444 and CM-1758 at 25% GI50. HL-60, ML-2, MOLM-13 and MV4-11 cell strains have been additionally handled for 48 h with 25% GI50 of Panobinostat, Quisinostat, Entinostat, Vorinostat, or Tubastatin (Supplementary Desk 9) or with the BRD inhibitors JQ1, Molibresib, and the mixture of CM-444 or CM-1758 with JQ1 and Molibresib (Supplementary Desk 9). Subsequent, the cells have been washed twice with phosphate-buffered saline (PBS) and resuspended in 1X Binding Buffer at a focus of 1 × 106 cells/mL. An 8-µL aliquot of APC-CD11b (BD Pharmingen, Franklin Lakes, NJ, USA) and 1 µL of FITC annexin-V (AV) (BD Pharmingen, Franklin Lakes, NJ, USA) antibodies have been added and incubated for 15 min at room temperature at the hours of darkness. Lastly, after the addition of 400 µL of 1X Binding Buffer to every tube, samples have been collected on a BD FACSCanto stream cytometer (Becton Dickinson, San Jose, CA, USA) and analyzed utilizing FlowJo software program.
For the remainder of experiments, the combo of antibodies (2 µL of BV786-CD13, Cat No, 744748 BD; 1 µL of diluted 1:10 antibody of PB-HLADR, Cat No 307633, Biolegend; 5 µL of APCH7-CD14, Cat No 641394, BD) was added to 10 µL of Good Stain Buffer Plus (Becton Dickinson, San Jose, CA, USA). Then, 1 million cells (quantity of 250 µL) have been added and incubated for 30 min at room temperature in darkness. After 10 min incubation at room temperature with 2 mL of FACSLysing 1X, cells have been centrifuged at 540 g for five min and washed twice with FACSBuffer. Lastly, after centrifugation at 540 g for five min cells have been resuspended in FACSBuffer.
The detailed gating technique for stream cytometry is proven in Supplementary Fig. 11.
Enzymatic assays
HDAC1, HDAC2, HDAC3 and HDAC6 IC50 values of compounds CM-444 and CM-1758 have been decided utilizing a particular fluorescence-labeled substrate (BPS Biosciences, Cat # 50037) after its deacetylation by HDAC, as described beforehand28. Outcomes are the typical of two organic replicates.
The HDAC4, HDAC5, HDAC7, HDAC8, HDAC9, HDAC10 and HDAC11 % inhibition at 10 μM and IC50 values have been decided by Eurofins (https://www.eurofins.com/), in duplicate.
HDAC11 IC50 values of CM-444 and CM-1758 have been decided by a steady and direct exercise assay for HDAC11 based mostly on inner fluorescence quenching48. Outcomes are the typical of two organic replicates.
DNMT1 and G9a IC50 values have been measured utilizing time-resolved fluorescence power switch assays, as described beforehand18,25,26. The outcomes are the typical of two organic replicates.
The selectivity of CM-444 and CM-1758 towards different epigenetic targets, together with histone demethylase (JMJD2A, JMJD2B, JMJD2C, JMJD2D, JMJD2E, JMJD3, JMJD1A, LSD1, Jarid1A, Jarid1B, Jarid1C, FBXL10, FBXL11, JHDM1D, JMJD1B, PHF8, LSD2 and UTX); histone acetyltransferases (GCN5, P300); histone methyltransferases (G9a, GLP, MLL1, SET7/9, SUV39H1, SUV39H2, PRMT1, PRMT3, PRMT4, PRMT5, PRMT6, PRMT8, PRMT9, NSD1, NSD2, EZH1, EZH2, SETDB1, SETD2, SET8, SUV4-20H1, SMYD2, SMYD3, DOT1L, EZH2 (A677G), EZH2 (A677G), EZH2 (A738T), EZH2 (Y641S), EZH2 (P132S), EZH2 (Y641C), EZH2 (Y641F), EZH2 (Y641H), EZH2 (Y641N)); bromodomains (ATAD2B, ATAD2A, BRD1, BRD9, BPTF/FALZ, CeCR2, BRPF3, BRD3 (BD1 + BD2), BAZ2B, WDR9, TAF1 (BD2), TAF1L (BD2), TAF1L (BD1 + BD2), BRD2 (BD1), BRD2 (BD2), BRD2 (BD1 + BD2), BRD3 (BD1), BRD3 (BD2), BRD4 (BD1), BRD4 (BD2), BRD4 (BD1 + BD2), BRDT (BD1), BRDT (BD1 + BD2), CREBBP, SMARCA2A, SMARCA4/BRG1); DNA methyltranferases (DNMT1, DNMT3A/3L and DNMT3B/3L), and sirtuin histone deacetylases (SIRT1, SIRT2, SIRT3, SIRT5 and SIRT6), have been decided by BPS Bioscience (http://www.bpsbioscience.com/index.ph). Binding experiments have been carried out in duplicate at a focus of 10 μM. If the share of inhibition was >50%, dose-response IC50 values have been decided (in duplicate).
Docking of CM-444 and CM-1758 towards HDAC isoforms
The GoldSuite 5.3.6 program (Cambridge Crystallographic Information Centre, https://www.ccdc.cam.ac.uk/pages/Residence.aspx) was used to carry out docking of CM-444 and CM-1758 to completely different HDAC isoforms with accessible crystallographic constructions: HDAC1 (PDB entry 4BKX49), CD2 area of HDAC6 complexed with trichostatin A (PDB entry 5EDU50), and HDAC7 complexed with trichostatin A (PDB entry 3C1051). The binding web site was outlined as a 20 Å sphere across the Zn atom of the catalytic cavity of HDACs and the remaining configuration parameters as described beforehand52.
Western blot
After treating HL-60, ML-2, MV4-11 and MOLM-13 cell strains each day for 48 h with 25% GI50 of CM-444 and CM-1758.48 h, the cells have been washed twice with PBS, the final centrifugation of 1520 g for 10 min at 4 °C. Histone extraction was carried out with a Histone extraction Package (Abcam, Cambridge, UK) following the producer’s directions. The histone focus within the extract was measured utilizing the dye-binding assay of Bradford. A ten µg quantity of histone was separated on 15% sodium dodecyl sulfate-polyacrylamide gel electrophoresis and transferred to a nitrocellulose membrane. The membrane, after being blocked with Tropix I-block blocking reagent (Cat No AI300, Tropix) in PBS with 0.1% Tween 20 and 0.02 NaN3 was incubated with the first antibody towards Acetyl H3 (rabbit polyclonal antibody, Cat No 06-599, Millipore) diluted 1:50,000 o/n at 4 °C or towards H3K27 me3 (mouse monoclonal antibody to histone H3 trimethyl K27, Cat No ab6002, Abcam) diluted 1:2000 in bovine serum albumin o/n at 4 °C after which with alkaline phosphatase-conjugated secondary antibodies. Sure antibodies have been revealed by a chemiluminiscent reagent (Tropix) and visualized utilizing the Chemidoc Imaging Techniques (Bio-Rad Laboratories). Whole H3 was used as a loading management (diluted 1:50,000 o/n at 4 °C or for 1 h at room temperature) (Anti-Histone H3, CT, pan, rabbit polyclonal, Cat No 07-690, Millipore). Pictures have been cropped for presentation.
Dot blot
After treating HL-60, ML-2, MV4-11 and MOLM-13 cell strains each day for 48 h with 25% GI50 of CM-444 and CM-1758.48 h, the cells have been washed twice with PBS, and genomic DNA was extracted utilizing a DNA Package (Nucleo Spin Tissue, Cat No 74095250, Macherey-Nagel) following the producer’s directions. DNA purity and focus have been measured utilizing a NanoDrop spectrophotometer (Thermo Scientific). A 500 ng quantity of genomic DNA was loaded onto a nitrocellulose membrane (Amersham Hybond_N+, RPN203B, GE Healthcare), pre-wetted in 6X SSC for 10 min, utilizing the Bio-Dot microfiltration equipment (Cat No 170-6545, Bio-Rad) following the producer’s directions. Subsequent, the membrane was incubated with 2X SSC for five min and cross-linked for two h at 80 °C. The membrane, after being blocked with Tropix I-block blocking reagent (Cat No AI300, Tropix) in PBS with 0.1% Tween 20 and 0.02 NaN3, was incubated with the first antibody towards 5-methylcytosine (monoclonal antibody 5-methylcytidine, Cat No BI-MECY-1000, Eurogentec) diluted 1:4000 o/n at 4 °C after which with alkaline phosphatase-conjugated secondary antibody. Sure antibodies have been revealed by a chemiluminiscent reagent (Tropix) and visualized utilizing the Chemidoc Imaging Techniques (Bio-Rad Laboratories). To make sure equal loading of whole DNA on the membrane, the identical blot was stained with 0.02% methylene blue in 0.3 M sodium acetate (pH 5.2).
LINE-1 pyrosequencing
DNA methylation of the repetitive aspect LINE-1 was analyzed utilizing a pyrosequencing approach. First, HL-60, ML-2, MV4-11 and MOLM-13 cell strains have been each day handled for 48 h with 25% GI50 of CM-444 and CM-1758. Cells have been washed twice with PBS, after which genomic DNA was extracted utilizing a DNA Package (Nucleo Spin Tissue, Cat No 74095250, Macherey-Nagel) following the producer’s directions. DNA purity and focus have been measured utilizing a NanoDrop spectrophotometer (Thermo Scientific). A 1 µg quantity of genomic DNA was handled and modified utilizing a CpGenome DNA modification Package (Cat No S7820, Chemicon Worldwide) following the producer’s directions. After bisulfite modification, a “scorching begin” polymerase chain response (PCR) (PyroMark PCR Package, Cat No 978703, Qiagen) was carried out with denaturalization at 95 °C for 15 min, adopted by 45 cycles consisting of denaturation at 94 °C for 1 min, annealing at 55 °C for 1 min, and extension at 72 °C for 1 min adopted by a remaining 10-min extension. This PCR was carried out utilizing 2 µL of modified DNA, 12.5 µL of 2X Buffer, 1 µL of 10 µM of every particular primer (remaining focus of 0.4 µM) (LINE-1-F: 5′- TTTTGAGTTAGGTGTGGGATATA -3′ and LINE-1-R: 5′-Biotin- AAAATCAAAAAATT CCCTTTC -3′) in a remaining quantity of 25 µL. The ensuing biotinylated PCR merchandise have been immobilized to Streptavidin Sepharose ® Excessive-Efficiency beads (GE Healthcare) and processed to yield high-quality ssDNA utilizing the PyroMark Vacuum Prep Workstation (Biotage) in keeping with the producer’s directions. The pyrosequencing reactions have been carried out utilizing the PyromarkTM ID (Biotage), and sequence evaluation was carried out utilizing the PyroQ-CpG evaluation software program (Biotage).
Major samples
Peripheral blood (PB) specimens have been obtained from preliminarily identified AML sufferers who had not acquired any therapy earlier than specimens have been collected. All sufferers gave knowledgeable consent for his or her participation on this research, which was accredited by the Scientific Analysis Ethics Committee of Clínica Universidad de Navarra. Written knowledgeable consent was obtained from every participant. PB was combined with an ammonium chloride-potassium (ACK) buffer lysis (0.15 mol/L NH4Cl, 10 mmol/L KHCO3, 0.1 mmol/L ethylenediaminetetraacetic acid, pH 7.2) at a 1:2 quantity ratio by light inversion, incubated for 10 min and centrifuged at 200 g for 10 min. The cell pellet was washed twice with PBS and at last resuspended in EGM2 medium (Cambrex Bioscience, Walkersville, MD). EGM2 accommodates 2% fetal calf serum, hydrocortisone, heparin, antibiotics, epidermal progress issue, human fibroblast progress issue, insulin-like progress issue, vascular endothelial progress issue, and ascorbic acid. The cells have been plated at 1 × 106 cells/mL and handled each day with 200 and 500 nM of CM-444 and CM-1758 for 48 h. Subsequent, the differentiation assay was carried out as described above.
Quantitative polymerase chain response (q-RT-PCR)
The expression of MYC, CDKN2A, CDKN1A, GATA2, TAL1, CEBPA and SPI1 have been analyzed by q-RT-PCR in HL-60, ML-2, MOLM-13 and MV4-11 cell strains after 48 h of CM-444, CM-1758, JQ1, Molibresib and the mixtures of CM-444 or CM-1758 with JQ1 and Molibresib remedies. RNA was extracted with TRIzol Reagent (Invitrogen) in keeping with the producer’s directions. First, cDNA was synthesized from 1 µg of whole RNA utilizing the PrimeScript RT reagent Package (Good Actual Time) (Cat No RR037A, TaKaRa) following the producer’s directions. The standard of cDNA was checked by a multiplex PCR that amplifies PBGD, ABL, BCR and β2-MG genes. A QuantStudio 5 Actual-Time PCR System (Utilized Biosystems) utilizing 20 ng of cDNA in 2 µL, 1 µL of every particular primer at 10 µM and 5 µL of SYBR Inexperienced PCR Grasp Combine 2X (Cat No 4334973, Utilized Biosystems) in a 10-µL response quantity was used to carry out q-RT-PCR. The next program circumstances have been used for the q-PCR: 50 °C for two min, 95 °C for 60 s following by 45 cycles at 95 °C for 15 s and 60 °C for 60 s; melting program, one cycle at 95 °C for 15 s, 40 °C for 60 s, and 95 °C for 15 s. The relative expression of every gene was quantified by the log 2(−ΔΔCt) technique utilizing the gene GUS as an endogenous management. The sequence of primers used will be present in Supplementary Desk 10.
Cell-cycle evaluation
For cell-cycle evaluation, 250,000 cells of HL-60, ML-2, MV4-11 and MOLM-13 cell strains have been cultured at a density of 1 × 106 cells/mL and handled for twenty-four h with CM-444 and CM-1758. Subsequent, the cells have been washed twice with PBS and resuspended in 0.2% Tween 20 in PBS and 0.5 mg/mL Rnase A (Ribonuclease A Sort III-A from bovine pancreas, Cat No. R5125, Sigma), after which incubated for 30 min at 37 °C. Subsequently, the cells have been stained with 25 μg/mL of propidium iodide (Cat No P4170, Sigma) and analyzed utilizing a BD FACSCanto stream cytometer (Becton Dickinson, San Jose, CA, USA).
Could–Grünwald Giemsa staining
After therapy of the HL-60, ML-2, MV4-11 and MOLM-13 cell strains each day with CM-444 and CM-1758 at 25% GI50 focus for 96 h, the cells have been washed twice with PBS and resuspended at 20,000 cells in 100 µL. Cytospins have been ready utilizing a cytospin centrifuge and we stained slides with Could–Grünwald Giemsa (Sigma, St. Louis, MO, USA). We examined mobile morphology utilizing a Nikon Eclipse 90i microscope (Nikon, Melville, NY, USA).
RNA-seq
Transcriptomic analyses have been carried out following low-input 3′ finish RNA sequencing approximation for characterization of transcriptional modifications in ML-2 and HL-60 samples handled with CM-444 and CM-1758.
Low-input 3′ finish RNA-seq was carried out as beforehand described53,54 with minor modifications. Briefly, RNA from 0.1 ×106 cells from every experimental situation was purified utilizing a Dynabeads mRNA DIRECT Purification Package (Thermo Fisher Scientific). For barcoded cDNA era, polyadenylated RNA was retrotranscribed utilizing an Affinity Script cDNA Synthesis Package (Agilent Applied sciences), and barcoded primers additionally harboring T7 promoter and a partial Read2 sequence for illumine sequencing (CGATTGAGGCCGGTAATACGACTCACTATAGGGGGCGACGTGTGCTCTTCCGATCTXXXXXXNNNNTTTTTTTTTTTTTTTTTTTT, the place XXXXXX is the cell barcode and NNN is a singular molecular identifier or UMI, which represents the identical preliminary RNA molecule for future quantification). At this level, the samples have been pooled (≤6 samples per pool) at equimolar ratios assessed by q-PCR.
The samples have been then handled with Exonuclease I (Thermo Fisher Scientific following a 1.2x optimistic SPRI-selection. Second-strand cDNA synthesis was then carried out utilizing a NEBNext Extremely II Directional RNA Second-Strand Synthesis Module (New England Biolabs) for two h at 16 °C, adopted by a 1.4x optimistic solid-phase reverse immobilization (SPRI) cleanup. The samples have been linearly amplified by T7 in vitro transcription utilizing a T7 RNA Polymerase (New England Biolabs) for 16 h at 37 °C. After DNAse therapy (TURBO DNAse I, Thermo Fisher Scientific) for 15 min at 37 °C to take away dsDNA, samples have been cleaned with a optimistic 1.2x SPRI-selection. RNA measurement, high quality, and focus have been decided by evaluation with a Qubit Fluorometer (Termo Fisher Scientific) and TapeStation (Agilent 4200 TapeSation System, Agilent Applied sciences). If essential, RNA was fragmented in 200- to 300-bp fragments utilizing Zn2+divalent cations adopted by a 2x optimistic SPRI cleanup. Fragmented RNA was dephosphorylated utilizing FastAP Thermosensitive Alkaline Phosphatase (Thermo Fisher Scientific) for 10 min at 37 °C to boost RNA ligation yield. On this step, a partial Read1 (AGATCGGAAGAGCGTCGTGTAG) sequence for illumine sequencing was ligated to the three′ finish of the fragmented RNA utilizing T4 RNA Ligase I (New England Biolabs). After a 1.5x optimistic SPRI cleanup, the RNA product was retrotranscribed utilizing an Affinity Script cDNA Synthesis Package (Agilent Applied sciences) and a particular retrotranscription primer (TCTAGCCTTCTCGCAGCACATC). The library was then purified by 1.5x optimistic SPRI cleanup, and the yield of the earlier steps was decided by q-PCR. Lastly, PCR utilizing KAPA HiFi HotStar Prepared Combine (Kapa Biosystems) was carried out for second-strand cDNA synthesis, illumine primers addition (Read1-P5: AATGATACGGCGACCACCGAGATCTACACTCTTTCCCTACACGACGCT CTTCCGATCT and Read2-P7: CAAGCAGAAGACGGCATACGAGATGTGACTGGAGTTCAGACGTGTGCTCTTCCGATCT) and library amplification utilizing the really helpful cycle numbers to reduce over-amplification. The product was purified by a 0.7x optimistic SPRI cleanup. The standard of the ultimate library was decided by q-PCR (based mostly on the edge cycle, or Ct, obtained for GAPDH (Fw: CCAGCAAGAGCACAAGAGGAA, Rv: GATTCAGTGTGGTGGGGG) or GUSB (Fw: CAAGTGCCTCCTGGACTGTT, Rv: TCCACCTTTAGTGTTCCCTGC) housekeeping genes calibrated per particular person set of samples, Qubit Fluorometer (Thermo Fisher Scientific) and TapeStation (Agilent 4200 TapeStation System, Agilent Applied sciences). All libraries have been pooled at equimolar ratios and sequenced on a NextSeq500 (Illumina) utilizing 60 bp single-end reads at a minimal sequence depth of 10 million reads per pattern.
Uncooked reads have been demultiplexed utilizing bcl2fastq2 Conversion Software program v2.19 (Illumina) to transform base name (BCL) information into FastQ information. The standard management of FastQ information was carried out utilizing FastQC (Bioinformatics Babraham Institute). Sequencing reads have been aligned to hg19 human reference utilizing Bowtie 2 (Johns Hopkins College) and quantified utilizing quant3p script (github.com/ctlab/quant3p). The DEseq2 bundle (R) was used for filtering, normalization, and evaluation of differential gene expression.
Practical evaluation
The Database for Annotation, Visualization, and Built-in Discovery database (david.ncifcrf.gov/residence.jsp) was used to carry out gene ontology (GO). A false discovery fee (FDR) < 0.05 was thought of to be vital.
The fold change ranked checklist of genes was used as enter to the non-parametric Kolmogorov–Smirnoff rank check as applied within the GSEA software program.
CYP inhibition
This research was performed at Wuxi (http://www.wuxi.com/). The inhibitory results of CM-444 and CM-1758 on 5 human cytochrome P450s (1A2, 2C9, 2C19, 2D6 and 3A4) have been evaluated in human liver microsomes. Check compounds (10 µM remaining focus) or management compounds (α-naphthoflavone, sulfaphenazole, (+)-N-3-benzylnirvanol, quinidine, ketoconazole; 3-µM remaining focus), and the corresponding substrates for every P450 isoform (1A2: phenacetin, 10-µM remaining focus; 2C9: diclofenac, 5-µM remaining focus; 2C19: S-mephenytoin, 30-µM remaining focus, 2D6: dextromethorphan, 5-µM remaining focus; 3A4: midazolam, 2-µM remaining focus) have been incubated with human liver microsomes (0.2-mg/mL remaining focus; Corning Cat # 452117) and NADPH cofactor (1-mM remaining focus, Chem-impex Cat # 00616) for 10 min at 37 °C. The response was terminated by including 400 µL of chilly cease answer (200 ng/mL tolbutamide and labetalol in acetonitrile), and the samples have been centrifuged at 1520 g for 20 min. Subsequent, 200-μL aliquots of supernatants have been diluted with 100 μL of ultra-pure water and shaken for 10 min. The samples have been analyzed by LC-MS/MS utilizing an analyte/inner customary peak space ratio. Check compounds and optimistic controls have been examined in duplicate. The share of inhibition was calculated because the ratio of substrate metabolite detected in handled and non-treated wells.
Plasma protein binding
This research was performed by Equilibrium Dialysis at Wuxi (http://www.wuxi.com/). HT Dialysis plate (Mannequin HTD 96 b, Cat# 1006) and the dialysis membrane (molecular weight cutoff of 12–14 kDa, Cat# 1101) have been bought from HT Dialysis LLC (Gales Ferry, CT). Human plasma (BioIVT, # HUMANPLK2P2N) and CD-1 mouse plasma (Beijing Important River Laboratory Animal Know-how Co., Ltd.) have been thawed previous to experiments underneath working chilly faucet water and centrifuged at 3320 × g for five min to take away any clots, and the pH was checked. Solely plasma from pH 7.0 to eight.0 was used. The dialysis membrane was pretreated in keeping with the producer’s directions. Check compounds and warfarin (management) have been assayed at a remaining focus of two μM. Aliquots of fifty μL loading plasma (matrix) containing check or management compound in triplicate have been transferred to Pattern Assortment Plates, respectively. The samples have been matched with reverse clean buffer to acquire a remaining quantity of 100 μL with a quantity ratio of plasma (matrix) to dialysis buffer (1:1, v-v) in every properly instantly. The cease answer (methanol–acetonitrile (50:50, v-v) containing tolbutamide at 200 ng/mL, labetalol at 200 ng/mL, and metformin at 50 ng/mL) have been added to those T0 samples of check compound and management compound. The plate was sealed and shaken at 800 rpm for 10 min. Then, these T0 samples have been saved at 2 °C–8 °C pending additional processing together with different post-dialysis samples. Aliquots of 150 μL of the loading plasma (matrix) containing check compound or management compound have been transferred to the donor facet of every dialysis properly in triplicate, and 150 μL of the dialysis buffer (100 mM sodium phosphate and 150 mM NaCl, pH 7.4 ± 0.1) was loaded to the receiver facet of the properly. Then, the plate was rotated at roughly 100 rpm in a humidified incubator with 5% CO2 at 37 °C ± 1 °C for 4 h. On the finish of the dialysis, aliquots of fifty μL samples from the buffer facet and plasma (matrix) facet of the dialysis system have been taken into new 96-well plates (Pattern Assortment Plates). An equal quantity of the other clean matrix (buffer or plasma) in every pattern was added to succeed in a remaining quantity of 100 μL with a quantity ratio of plasma (matrix) to dialysis buffer at 1:1 (v:v) in every properly. Samples have been additional processed by protein precipitation. Concentrations of check compounds within the beginning answer (earlier than dialysis), the plasma facet of the membrane, and the buffer facet of the membrane have been quantified by LC-MS/MS utilizing a peak space ratio of analyte/inner customary. The fraction of unbound, sure, and restoration have been calculated. Outcomes are the typical of three organic replicates.
Kinetic solubility
This research was performed at Wuxi (http://www.wuxi.com/). Inventory options of check compounds (10 mM; 100% DMSO) and 490 μL of PBS buffer at pH = 7.4 have been added within the decrease chamber of Whatman’s Mini-Uniprep vials. After vortex mixing for ≥2 min, the vials have been shaken at 800 rpm for twenty-four h and at last centrifuged at 1520 g for 20 min. The focus of the filtrate was quantified by HPLC (calibration curve from 1.5624 μM to 200 μM). Amiodarone hydrochloride, carbamazepine and chloramphenicol have been used as controls.
CACO-2 permeability
This research was performed at Wuxi (http://www.wuxi.com/). Caco-2 cells (ATCC) have been seeded onto polyethylene membranes in 96-well BD insert plates at 1 × 105 cells/cm2, and the medium was modified each 4 days till the 21st to twenty-eightth day for confluent cell monolayer formation. Check compounds have been diluted with transport buffer (HBSS with 10 mM HEPES at pH 7.4 ± 0.05) to a focus of two µM (remaining DMSO focus adjusted to <1%) and examined bi-directionally to analyze P-glycoprotein-mediated efflux in duplicate. Controls: digoxin (10 μM, bi-directionally), nanodolol and metoprolol (2 μM, apical, A, to basolateral, B). The permeation was assessed over a 120-minute incubation at 37 °C ± 1 °C and 5% CO2 at saturated humidity with out shaking. Samples have been combined with acetonitrile containing inner customary and centrifuged at 1520 g for 10 min. Then, 100-μL aliquots of supernatant options have been diluted with 100 μL of pure water. Concentrations of check and management compounds in beginning answer, donor answer, and receiver answer have been quantified by LC-MS/MS utilizing a peak space ratio of analyte to inner customary. Permeation of lucifer yellow by the monolayer was measured to judge the mobile integrity. The outcomes are the typical of two organic replicates.
Human ether-a-go-go associated gene (hERG) blockade assay
This research was performed at Wuxi (http://www.wuxi.com/), utilizing CHO cells with secure expression of hERG potassium channels (Sophion Biosciences, Ballerup, Denmark). Cells have been cultured (F12 medium, supplemented with 10% FBS, 1% Geneticin® selective antibiotic, 89 μg/mL hygromycin B) in a humidified and air-controlled (5% CO2) incubator at 37 °C ± 2 °C. Check compounds have been diluted at completely different concentrations (0.3 μM–30μM), with a remaining DMSO focus in an extracellular answer of <0.3%. The exterior answer consisted of two mM CaCl2, 1 mM MgCl2, 4 mM KCl, 145 mM NaCl, 10 mM glucose and 10 mM HEPES at pH 7.3–7.4. The inner answer consisted of 5.37 mM CaCl2, 1.75 mM MgCl2, 120 mM KCl, 10 mM HEPES, 5 mM EGTA and 4 mM Na2ATP at pH 7.2–7.3. The assay was carried out at room temperature utilizing the whole-cell patch clamp strategies managed with Patchmaster Professional software program. For high quality management, minimal seal membrane resistance was set at 500 MOhms and the minimal/most particular hERG present (pre-compound) was 400 pA/3000 pA. A 100-nM quantity of cisapride was used because the optimistic management to judge the reliability of the check system. Outcomes are the typical of two organic replicates.
PAMPA permeability
This assay was carried out in triplicate as described beforehand25,26.
Cytotoxicity in THLE-2 cells and PBMCs cells
The cytotoxicity of compounds CM-444 and CM-1758 towards the immortalized human liver THLE-2 cell line (ATCC CRL-2706) following 24-h and 72-h publicity and PBMCs following a 72-h publicity was carried out in duplicate (pLC50 distinction <0.30 and 0.50 log items, for THLE-2 and PBMCs, respectively), following the protocol described beforehand18,25,26.
Human and mouse liver microsomal stability
This research was performed at Wuxi (http://www.wuxi.com/). The information collected have been analyzed to calculate a half-life (t1/2, min) for check compounds at a remaining focus of 1 μM. A 5-μL aliquot of inventory answer of check compound (10 mM; 100% DMSO) was diluted in 495 μL of 1:1 methanol/water (remaining focus of 100 μM, 99% MeOH). Then, 50 μL of this intermediate answer was diluted in 450 μL of 100 mM potassium phosphate buffer to a focus of 10 μM (working answer, 9.9% MeOH). The β-nicotinamide adenine dinucleotide phosphate diminished kind (NADPH) regenerating system additionally contained tetrasodium salt, NADPH•4Na (Sigma, Cat.# No.00616). Human liver microsomes have been obtained from Corning (Cat.#452117), and C57 mouse liver microsomes from Xenotech (Cat.#M5000), at a remaining focus of 0.5 mg protein/mL. A quantity of 10 μL of working answer of check and management compounds (testosterone, diclofenac and propafenone) and 80 μL/properly microsome answer have been added to a 96-well plate and incubated for 10 min at 37 °C. The response was began by the addition of 10 μL of NADPH regenerating system and stopped by the addition of 300 μL of cease answer (acetonitrile at 4 °C, together with 100 ng/mL tolbutamide and 100 ng/mL of labetalol as inner customary) at completely different incubation instances (0, 5, 10, 20, 30, and 60 min). Then, the samples have been shaken for 10 min and centrifuged at 1699 g for 20 min at 4 °C. An aliquot of supernatant (100 μL) was transferred from every properly and combined with 300 μL of ultra-pure water earlier than submitting to LC-MS/MS evaluation. Concentrations of check and management compounds have been quantified by LC-MS/MS utilizing a peak space ratio of analyte/inner customary, and the % lack of the mum or dad compound was calculated at every time level to find out the half-life (t1/2, min). Experiments have been carried out in duplicate.
Human and mouse liver cryopreserved hepatocytes
This research was performed at Wuxi (http://www.wuxi.com/). The information collected have been analyzed to calculate a half-life (t1/2, min) for check compounds at a remaining focus of 1 μM. Inventory options of check compound (10 mM; 100% DMSO) and optimistic controls (7-ethoxycoumarin and 7-hydroxycoumarin; 30 mM; 100% DMSO) have been diluted to 1 mM and three mM with DMSO after which diluted to 100 μM and 300 μM with acetonitrile (dosing options). Cryopreserved cells (from Human Donors, Bioreclamation IVT, Cat No. X008001 and from male C57 Mice, Bioreclamation IVT, Cat No. S00463) have been thawed, remoted and suspended in Williams’ Medium E, then diluted with pre-warmed Williams’ Medium E to 0.5 × 106 cells/mL. Cell viability was examined by trypan blue exclusion after thawing. A quantity of two μL of dosing answer and 198 μL of pre-warmed cell suspensions have been added to a 96-well plate and incubated at 37 °C (5% CO2) with fixed shaking at 600 rpm. Following completely different incubation instances (0, 15, 30, 60 and 90 min), 20 μL of every pattern was transferred to a properly containing 80 μL of ice-cold cease answer (acetonitrile containing 200 ng/mL tolbutamide and 200 ng/mL labetalol as inner requirements). Plates have been vortexed at 27 g for 10 min after which centrifuged at 3220 × g for 20 min at 4 °C. An aliquot of supernatant (75 μL) was transferred from every properly and combined with 75 μL of ultra-pure water earlier than submitting to LC-MS/MS evaluation. Concentrations of check and management compounds have been quantified by LC-MS/MS utilizing a peak space ratio of analyte/inner customary, and the % lack of the mum or dad compound was calculated at every time level to find out the half-life (t1/2, min). Experiments have been carried out in duplicate.
Plasma stability assay
This research was performed at Wuxi (http://www.wuxi.com/). Recent feminine C57 BL/6 mouse plasma (6 mL; Wuxi Apptec) was collected on the day of the experiment, and the pooled frozen human plasma (BioreclamationIVT, Cat No. HMPLEDTA2) was thawed in a water tub at 37 °C previous to the experiment. EDTA-K2 was used as an anticoagulant. Plasma was centrifuged at 1520 g for five min, and any clots have been eliminated. The pH was adjusted to 7.4 ± 0.1 if required. Half volumes of mouse and human plasma have been pre-warmed in a water tub at 37 °C, and one other half quantity every of mouse and human plasma have been saved at room temperature previous to the experiment. Intermediate options (1 mM) of examined compounds and optimistic management (propantheline) have been ready by diluting 5 µL of the inventory answer (10 mM; 100% DMSO) with 45 µL DMSO (check compounds) or ultra-pure water (propantheline). Then, working options (100 μM) have been ready by diluting 10 μL of intermediate options with 90 μL of 45% MeOH/H2O. Subsequent, 98 µL of clean plasma was spiked with 2 μL of dosing answer (100 μM) to attain a 2 μM remaining focus in duplicate, and samples have been incubated at 37 °C in a water tub. At every time level (0, 1, 2, 4, 6, and 24 h), the reactions have been stopped by eradicating the plates from a water tub and by addition of 400 μL of ice-cold cease answer (200 ng/mL tolbutamide and 200 ng/mL labetalol in MeOH/acetonitrile (v:v = 1:1)). The samples have been instantly vortexed on a plate shaker at 800 rpm for 10 min. Samples have been centrifuged at 1699 g for 10 min, and an aliquot of supernatant (100 μL) was transferred from every properly and combined with 200 μL ultra-pure water earlier than submission for LC-MS/MS evaluation (Acquity UPLC HSS). Concentrations of check compounds have been quantified by LC-MS/MS utilizing the height space ratio of analyte/inner customary, and the share lack of the mum or dad compound was calculated at every time level to find out the half-life. Experiments have been carried out in duplicate.
CM-444 and CM-1758 toxicity assay: hematological and liver parameters
After treating Rag2−/−γc−/− mice with i.p. 10 mg/kg of CM-444 and CM-1758 for five consecutive days adopted by 2 relaxation days over 3 weeks, adopted by a 7-day washout interval. Hematological and liver parameters have been measured.
Hematological parameters, together with white blood cells, crimson blood cells, platelet depend, hemoglobin and hematocrit, have been measured on a Hemavet Hematology Analyzer (Drew Scientific). Liver enzymes (albumin, alkaline phosphatase, aspartate transaminase, alanine transaminase), ldl cholesterol, bilirubin, urea and bile acid ranges have been analyzed from mice serum utilizing a C311 Cobas Analyzer (Roche Diagnostics).
Mice livers have been mounted in paraformaldehyde at 4% for six–8 h, washed twice with saline answer and saved in 70% ethanol. Samples have been included in paraffin, and 3-µm serial sections have been lower, deparaffinated, and stained with hematoxylin–eosin.
Pharmacokinetic research of CM-444 and CM-1758 in plasma samples
Compounds CM-444 and CM-1758 have been measured in plasma utilizing an Acquity UPLC system (Waters, Manchester, UK) coupled to an Xevo-TQ MS triple-quadrupole mass spectrometer with ESI supply.
Compound options have been ready by dissolving the strong in dimethyl sulfoxide (DMSO) and this answer was made as much as a remaining quantity by the addition of a combination of Tween 20 and 0.9% NaCl (1/1/8, v-v:v, DMSO/Tween 20/saline). For the pharmacokinetic experiments, a drug dosage of 10 mg/kg was administered as a single intraperitoneal injection to RAG-2 mice. Blood was collected at predetermined post-injection instances (15 min, and 1, 2, 4, 8 and 24 h) into EDTA-containing tubes, and plasma was obtained through centrifugation (4 °C, 657 g, 5 min) and saved at −80 °C till evaluation.
Chromatographic separation was carried out by gradient elution at 0.6 mL/min utilizing an Acquity UPLC BEH C18 column (50 × 2.1 mm, 1.7 µm; Waters). The cellular part consisted of A: water with 0.1% formic acid, B: methanol with 0.1% formic acid. The autosampler temperature was set at 10 °C and the column temperature at 40 °C. For detection and quantification, the ESI operated within the optimistic mode was arrange for a number of response monitoring. The collision gasoline used was ultra-pure argon at a stream fee of 0.15 mL/min.
Quantification was achieved by exterior calibration utilizing matrix-matched requirements. Concentrations have been calculated utilizing a weighted least-squares linear regression (W = 1/x). Calibration requirements have been ready by including the suitable quantity of diluted compound answer (made in a combination of methanol and water, 50:50, v-v) to aliquots of 20 µL of clean plasma. The calibration customary and pattern preparation have been as follows: 3% formic acid in methanol was added to precipitate the proteins. The combination was then vortex-agitated for 1 min and centrifuged at 16,522 g for 10 min at 4 °C. A 5-µL aliquot of the supernatant answer was injected into the LC-MS/MS system for evaluation. Frozen plasma samples have been thawed at room temperature, vortex combined completely and subjected to the above-described extraction process.
The pharmacokinetic parameters have been obtained by becoming the blood concentration-time knowledge to a non-compartmental mannequin with WinNonlin software program (Pharsight, Mountain View, CA).
In vivo experiments
All animal research had earlier approval from the Animal Care and Ethics Committee of the College of Navarra.
Mice have been housed in an appropriate temperature and humidity setting (25 °C, appropriate humidity (usually 50%), 12 h darkish/mild cycle), and fed with adequate water and meals. All mice have been euthanized through cervical dislocation.
For these research, feminine mice have been used resulting from increased engraftment charges as in comparison with male mice.
The human AML HL-60 and ML-2 cell strains have been first pretreated in vitro each day with 270 and 260 nM of CM-444 or 300 and 210 nM of CM-1758, respectively, for 4 days. Previous to injection in mice, the cell differentiation induction by CM-444 and CM-1758 compounds was verified by measuring CD11b by stream cytometry, as described earlier. Then, 5 × 106 cells of cells, pretreated with CM-444, CM-175, or automobile (80% saline answer, 10% DMSO and 10% Tween 20) and diluted in 100 µL of saline answer, have been implanted subcutaneously into the flank of feminine BALB/cA Rag2−/−γc−/− mice between 6 and eight weeks of age (n = 8). Tumor measurement was analyzed each 2–3 days utilizing the next technique: V¼ D_d2/2, the place D and d corresponded to the longest and shortest diameter, respectively. The maximal tumor measurement/burden permitted by the Animal Care and Ethics Committee of the College of Navarra is 2000 mm3. In some instances, this restrict was reached on the final day of measurement and the mice have been instantly euthanized.
The subcutaneous xenograft fashions of ML-2 and MV4-11 derived cells, 5 × 106 and 10 × 106 cells, respectively, diluted in 100 µL of saline answer have been subcutaneously inoculated within the again left flank of feminine BALB/cA Rag2−/−γc−/− mice. When tumors turned palpable, the mice have been randomized into three teams: automobile, CM-444 and CM-1758, (eight animals/group). Remedy with 10 mg/kg of CM-444 or CM-1758 i.p. was began 4 days after injection when all mice offered subcutaneous tumors and was administered for five consecutive days adopted by 2 resting days over 3 weeks for the ML-2 mannequin or each day for two weeks for the MV4-11 mannequin. The management group acquired solely 80% saline answer, 10% DMSO or 10% Tween 20 (diluents of CM-444 and CM-1758). Tumor measurement was analyzed each 2–3 days utilizing the next technique: V¼ D_d2/2, the place D and d corresponded to the longest and shortest diameter, respectively. The maximal tumor measurement/burden permitted by the Animal Care and Ethics Committee of the College of Navarra is 2000 mm3. This maximal tumor measurement/burden was not exceeded in any case. ML-2 and MV4-11 tumors have been explanted for differentiation and international histone acetylation evaluation. Histones have been extracted, and a western blot of H3Ac was carried out as described above. For the differentiation research, RNA was extracted, and q-PCR of CD11b was carried out as described above (CD11b sequence primer will be present in Supplementary Desk 10).
The human AML MV4-11 and MOLM-13 xenograft mice fashions have been generated by intravenous (i.v.) injection of 10 × 106 (MV4-11) or 1 × 104 (MOLM-13) cells diluted in 100 µL of saline answer within the tail vein of a 6-8-week-old feminine BALB/cA Rag2−/−γc−/− mice (n = 10). Remedy began 7 or 1 day after MV4-11 or MOLM-13 cells injection, respectively. MV4-11 and MOLM-13 mice have been handled with i.p. 10 mg/kg of CM-444 and CM-1758 throughout 5 consecutive days adopted by 2 resting days for 7 and a pair of weeks, respectively. The management group acquired solely 80% saline answer, 10% DMSO and 10% Tween 20 (diluents of CM-444 and CM-1758). The mice weight was managed and remained secure in the course of the therapy. Statistical outcomes have been calculated utilizing the statistical software program Medcalc. CD11b ranges in blood samples from an intravenous MV4-11 mice mannequin have been measured by stream cytometry, as described above.
Acetylome and proteome evaluation
Pattern preparation
Mobile pellets have been lysed for 15 min at 95 °C in 4% SDS, 100 mM Hepes/NaOH pH 8.0, containing 5 mM sodium butyrate and 5 mM Nicotinamide. After cooling down, lysates have been incubated at 25 °C with 10 items of DNAse (Benzonase, Merk) and sonicated for 15 min in a Bioruptor for DNA shearing. Protein focus was decided utilizing microBCA, utilizing BSA as customary. Samples have been digested utilizing on-bead protein aggregation seize (PAC) with ReSyn Biosciences MagReSyn® Hydoxyl microparticles (ratio Protein/Beads 1:5) in an automatic King Fisher instrument (Thermo). Proteins have been digested for 16 h at 37 C with LysC/Trypsin combine at a protein: enzyme ratio of 1:100 in 50 mM TEAB pH 8.0 (Trypsin Sequence grade, Sigma; LysC, Wako). The ensuing peptides have been speed-vac dried and dissolved in 100 µL of 200 mM HEPES pH 8.5.
Peptides (500 µg) have been labeled with Thermo Scientific TMTpro 18plex™ Isobaric reagents for 1 h at 25 C. Response was quenched by including 5% hydroxylamine for 15 min. Samples have been pooled at a 1:1 ratio based mostly on the entire peptide quantity, decided by evaluating total sign intensities on a daily LC-MS/MS run. The ultimate combination was desalted utilizing a 500 mg Sep-Pak C18t cartridge (Waters) and dried previous to excessive pH reverse part HPLC pre-fractionation.
Excessive pH reverse part chromatography
Labeled peptides have been fractionated offline by a excessive pH reverse part chromatography utilizing an Final 3000 HPLC system geared up with a pattern collector. Briefly, peptides have been dissolved in 1200 µL of Buffer A (10 mM NH4OH) and loaded onto a XBridge BEH130 C18 column (3.5 µm, 250 mm size and three.5 mm ID, Waters). Buffer B was composed of 10 mM NH4OH in 90% CH3CN. The next gradient was used at a stream fee of 500 µL/min: 0-50 min 0-25% B, 50-56 min 25-60% B, 56-57 min 60-90% B. One-minute fractions (500 µL) from minute 15 to 65 have been collected and neutralized with 10% formic acid. Based mostly on the UV absorbance at 280 nm, 95% of the amount of 40 fractions was pooled in 5 fractions for Acetyl-Lysine enrichment. The 5% of those fractions was stored for proteome evaluation. Pooled fractions have been snap-frozen and lyophilized.
Immuno-Purification of Lysine Acetylated Peptides
Lyophilized peptides (1.5 mg) from every pool have been immuno-affinity purified with the PTMScan HS Acetyl-Lysine Motif (Ac−Okay) equipment (Cell Signaling Know-how). For every pool, three consecutive immuno-purifications (IPs) have been carried out utilizing the producer’s protocol in an automatic King Fisher instrument (Thermo). Elution of purified peptides was carried out in two consecutive steps of fifty μl of 0.15% TFA. Samples have been desalted with C18 stage suggestions and eluates have been vacuum dried and resuspended in 0.1% formic acid for subsequent evaluation by LC−MS/MS.
Mass spectrometry
LC-MS/MS was carried out by coupling an UltiMate 3000 RSLCnano LC system to an Orbitrap Exploris 480 mass spectrometer (Thermo Fisher Scientific). Samples have been loaded right into a entice column (Acclaim™ PepMap™ 100 C18 LC Columns 5 µm, 20 mm size) for 3 min at a stream fee of 10 µL/min in 0.1% FA. Peptides have been then transferred to an EASY-Spray PepMap RSLC C18 column (Thermo) (2 µm, 75 µm x 50 cm) operated at 45 °C and separated utilizing a 90 min efficient gradient (buffer A: 0.1% FA; buffer B: 100% ACN, 0.1% FA) at a stream fee of 250 nL/min. The gradient used was, from 4% to six% of buffer B the primary 2 min, from 6% to 33% B the following 58 min, plus 10 extra minutes at 98% B.
The mass spectrometer was operated in a data-dependent mode, with an automated swap between MS and MS/MS scans utilizing a prime 15 technique. (Depth threshold ≥ 1e4, dynamic exclusion of 20 sec and excluding fees unassigned, +1 and ≥ +7). MS spectra have been acquired from 350 to 1500 m/z with a decision of 60,000 FMHW (200 m/z). Ion peptides have been remoted utilizing a 0.7 Th window and fragmented utilizing higher-energy collisional dissociation (HCD) with a normalized collision power NCE of 36. MS/MS spectra have been acquired with a set first mass of 120 m/z and a decision of 45,000 FMHW (200 m/z). The ion goal values have been 3e6 for MS (most IT 25 ms) and 1e4 for MS/MS (most IT, auto).
Information evaluation
For acetyl-lysine enriched TMT-labeled samples, uncooked information have been processed with MSFragger (Yu, F et al, Nat. Comm. 2020) towards a human protein database (UniProtKB/TrEMBL, 20,593 sequences, UP000005640_9606, Could 2023) supplemented with contaminants. Carbamidomethylation of cysteines, N-term and lysine TMTpro-label have been set as mounted modifications, whereas oxidation of methionine, protein N-term acetylation, N/Q de-amidation and acetylation of lysines have been set as variable modifications. Minimal peptide size was set to 7 amino acids and a most of 4 tryptic missed-cleavages have been allowed. Outcomes have been filtered at 0.01 FDR (peptide and protein degree).
For whole protein evaluation, uncooked information have been processed with MaxQuant (v 2.4.9.0) utilizing the usual settings towards the identical protein database.
The acetyl-Okay websites and protein intensities information have been loaded in Prostar (v1.32.3) (Wieczorek et al, Bioinformatics 2017) utilizing the depth values for additional statistical evaluation. Briefly, proteins/websites with lower than eighteen legitimate values have been filtered out. Then, a world normalization of log2-transformed intensities throughout samples was carried out utilizing the LOESS perform. Differential evaluation was carried out utilizing the empirical Bayes statistics Limma. Proteins with a p-value < 0.01 and a log2 ratio >0.3 or <-0.3 have been outlined as regulated (Martinez-Val et al, Proteome Res, 2016). The FDR was estimated to be beneath 2% by Benjamini-Hochberg.
Practical evaluation, hierarchical clustering, PCA, motif annotation and Fisher precise assessments have been carried out utilizing the Perseus software program platform55.
Evaluation of the acetylation motifs
Acetyl-lysine motif evaluation was carried out and offered utilizing the iceLogo motif builder56 utilizing a p-value of 0.01.
We constructed the reference a number of sequence alignment set through the use of the 15 amino acids surrounding the Okay-acetylation web site of the quantified Okay-acetylation websites from the experiment. For the optimistic set, we used the identical 15-amino-acid surrounding sequence window comparable to the histones. The usual deviation (σ) is calculated utilizing the pattern measurement (N) and the frequency (f%) of an amino acid within the reference set, with the system:
$$sigma=sqrt{left(left(fpercentright)/Nright)}$$
For each place, the amino acid frequencies within the optimistic set might be in contrast with the frequencies within the reference set. An amino acid might be regulated if the Z-score isn’t part of the arrogance interval. For the calculation of the arrogance interval the Wichura algorithm was used (Supplementary Desk 11) The Z-score is calculated with the system: Z-score = X − μ σ. The system will calculate what number of instances the frequency (X) is deviated from the imply (μ, the frequency of a particular amino acid on a particular place within the reference set) when it comes to the usual deviation (σ).
For visualization the share distinction, utilizing the distinction in frequency for an amino acid within the experimental set and the reference set as a measure of the peak of a letter within the amino acid stack, was used57.
STRING database
The protein–protein interactions community was constructed from the web STRING database (https://string-db.org/, model 11.5). The interactions (the default threshold > 0.4 within the STRING database) have been chosen to create the PPI community and additional clustered by the Okay-means technique.
CUT&RUN
CUT&RUN was carried out with the equipment CUTANA ChIC/CUT&RUN (Epicypher, 14-1048) following the producer’s directions. Briefly, 0.5 million of HL-60 cells have been harvested and resuspended in 100ul wash buffer [20 mM HEPES pH 7.5, 150 mM NaCl, 0.5 mM Spermidine, supplemented with Protease Inhibitor EDTA-Free tablet (Roche 11836170001)]. Activated Concanavalin A was incubated with cells at room temperature for 10 min to let the cells bind to the beads. For every goal protein issue, 0.5 μg of BRD4 CUTANA CUT&RUN antibody (Cat No 13-2003, Epicypher), Acetyl-Histone H3 (Lys 9) (C5B11) rabbit monoclonal antibody diluted 1:50 (Cat No 9649 T, Cell Signaling) or Acetyl-Histone H3 (Lys 27) (D5E4) XP rabbit monoclonal antibody diluted 1:100 (Cat No 8173T, Cell Signaling) was added to every pattern and incubated within the antibody buffer (wash buffer +0.01% Digitonin and a pair of mM EDTA) in a single day at 4 °C. Isotype management IgG was used as a destructive management. The beads have been then washed twice with digitonin buffer [wash buffer +0.01% Digitonin] and a pair of.5 μL pAG-MNase was added to every pattern. After ten minutes of incubation at room temperature, extreme pAG-MNase was washed out by a two-time digitonin buffer wash. Then focused chromatin was digested and launched from cells by 2 h of incubation with the presence of two mM CaCl2 at 4 °C. After incubation with cease buffer [340 mM NaCl, 20 mM EDTA, 4 mM EGTA, 50 μg/mL RNase A, 50 μg/mL Glycogen] for 10 min at 37 °C the supernatant containing CUT&RUN-enriched DNA was purified utilizing the CUTANA DNA Purification equipment (Epicypher). Lastly, Illumina sequencing libraries have been ready from 6 ng of purified CUT&RUN DNA utilizing NEBNext UltraTM II DNA Library Prep Package for Illumina, following the producer’s directions. For BRD4, some modifications have been added following the protocol of Nan Liu (Harvard College; https://doi.org/10.17504/protocols.io.wvgfe3w). Libraries have been sequenced utilizing NEXTseq500 (Illumina).
The Lower&Run samples have been analyzed utilizing the CUT&RUN instruments pipeline with default settings and no filtering for 120 bp fragments58. Peak calling was carried out on deduplicated reads with macs2 model 2.2.6 utilizing the respective IgG samples as management, slim peak calling and paired-end settings, and a threshold of q = 0.01.
To research the chromatin state distribution within the three circumstances (Management, CM-444 therapy and Panobinostat therapy) we overlapped the referred to as areas of every pattern of the research with the chromatin states of every AML pattern. Particularly, 12 chromatin states, based mostly on the mixture of 6 histone marks, for 38 AML samples have been used59. The overwhelming majority of the areas could possibly be solely annotated to a single chromatin state whereas some could possibly be solely annotated to a single chromatin state after the substitute of consecutive chromatin states with related traits with one among them (i.e mixture of Transcription Transition / Transcription Elongation / Weak Transcription to Transcription Transition or mixture of Heterochromatin; Represesed/ Heterochromatin; Low Sign to Heterochromatin; Repressed and so on). The areas that couldn’t be solely annotated to a single chromatin state or simplified chromatin state have been eliminated. Utilizing this method, we achieved to annotate between 82.2%-92.2% of the areas referred to as within the 9 samples. We then calculated the share of every chromatin state making an allowance for the width of the areas. As background, we used all of the areas of every AML pattern and we calculated the share of the chromatin states equally. Lastly, we calculated the fold change of every chromatin state as a proportion within the pattern to a proportion within the background AML pattern.
Statistical evaluation
Comparisons have been made with the Mann–Whitney U-test (for unpaired samples with no regular distribution) and Scholar’s t-test (for paired samples displaying Gaussian distribution). Survival analyses (recurrence-free) in keeping with varied variables have been carried out utilizing the Kaplan–Meier technique, and variations between the completely different teams of sufferers or mice have been examined with the log-rank check. GraphPad Prism 6.0 was used. P < 0.05 was thought of to be indicative of statistically vital variations.
Reporting abstract
Additional info on analysis design is obtainable within the Nature Portfolio Reporting Abstract linked to this text.

