Differential IFNγ exercise in distinct subgroups of AML sufferers
We used single-sample gene set enrichment evaluation (ssGSEA) to look at the transcriptional packages linked to IFNγ signaling in 672 newly recognized grownup AML sufferers from 3 unbiased datasets: TCGA4, MD Anderson Most cancers Middle (MDACC)29, and BEAT-AML3. Affected person traits have been summarized in Supplementary Knowledge 1. By making use of ssGSEA to every pattern individually, we calculated unbiased enrichment scores for every gene set-sample pairing. There was a constructive correlation between the IFNγ signaling rating, and immune activation pathways curated from the Gene Ontology (GO), Hallmark, and Reactome gene set collections of the Molecular Signature Database (MSigDB) (Fig. 1A, Supplementary Knowledge 2). Moreover, IFNγ signaling rating had an roughly regular distribution, suggesting a various extent of signaling amongst people with AML (Supplementary Fig. 1A).
A Single-sample gene set enrichment evaluation of TCGA, MDACC, and BEAT-AML bulk RNA sequencing datasets utilizing pathways curated from Gene Ontology (GO), Hallmark, and Reactome gene set collections of the Molecular Signature Database. B Correlation of blast share within the bulk RNA sequencing cohorts and Hallmark IFNγ response pathway. Error band represents 95% confidence interval. T check was used to judge the importance of Pearson correlation. C Hallmark IFNγ response rating by particular FAB classification in sufferers with diploid cytogenetics (n = 294; NOS = not in any other case specified; Middle line represents the median and decrease and higher bounds of field correspond to the primary and third quartiles). D Hallmark IFNγ response rating by particular cytogenetic teams (n = 378); double deletion signifies sufferers with each a chromosome 5/5q and seven/7q loss. Middle line represents the median and decrease and higher bounds of field correspond to the primary and third quartiles. Two-sided Wilcoxon check was used to match inv(16) with t(8;21). E Hallmark IFNγ response rating evaluating AML samples and wholesome CD34+ sorted bone marrow cells. Middle line represents the median and decrease and higher bounds of field correspond to the primary and third quartiles. Two-sided Wilcoxon check was used. F Pie chart exhibiting the proportion of newly recognized AML sufferers with elevated IFNγ stage in comparison with the traditional vary. G Correlation of IFNγ response pathway with HLA1, HLA2, T-cell exhaustion, T-cell dysfunction, and T-cell senescence scores (see additionally Supplementary Fig. 1B–F). H Correlation of Hallmark IFNγ response with monocytes as decided by means of CIBERSORTx immune deconvolution of bulk RNA profiling information. Error band represents 95% confidence interval. T check was used to judge the importance of Pearson correlation. Supply information are supplied as a Supply Knowledge file.
We subsequent examined medical traits related to IFNγ signaling rating and located a destructive correlation with blast share (r = −0.41, p < 2.2 × 10−16) (Fig. 1B). Moreover, we noticed vital variations in IFNγ signaling rating throughout AML differentiation states, as outlined by the French American British (FAB) classification (Kruskal-Wallis check, p = 1.76 × 10−8) and cytogenetic teams (Kruskal-Wallis check, p = 0.01). Particularly, amongst AML sufferers with diploid cytogenetics, these with monocytic differentiation (FAB M4/M5) had the very best IFNγ signaling scores (Fig. 1C). Sufferers with diploid cytogenetics however with no reported FAB classification, known as diploid, not in any other case specified (NOS), displayed intermediate ranges of IFNγ signaling scores, possible indicating that this group contained a mix of monocytic and non-monocytic sufferers.
Amongst AML sufferers with non-diploid cytogenetics, the IFNγ signaling rating was discovered to be highest in core-binding-factor (CBF) AML with inv(16). This rating was considerably greater than that of t(8;21) CBF AML (p = 5.49 × 10−4), which is noteworthy as a result of inv(16) AML sometimes displays a myelomonocytic (M4) differentiation, whereas t(8;21) CBF leukemia is normally extra myeloid (M2) (Fig. 1D)30. Amongst sufferers with non-CBF, non-diploid AML, these with a deletion in chromosome 7/7q (del7/7q) exhibited highest IFNγ signaling scores (Fig. 1D). Notably, sorted CD34+ cells which marks the wholesome stem cells from 17 wholesome donors3 had markedly decrease ranges of IFNγ signaling scores than did these from AML sufferers (Fig. 1E), suggesting that IFNγ pathway signaling is a predominant function in AML. Additional, we assessed IFNγ concentrations within the sera of 43 consecutively collected, newly recognized AML sufferers that current to our heart and noticed elevated stage of IFNγ in sera of 67.4% of sufferers that exceeded these typical of the wholesome reference group, which fortifies the notion that IFNγ is dysregulated on the time of analysis in AML sufferers and underscores the necessity for additional exploration into its potential medical implications and therapeutic utility (Fig. 1F). Total, these findings point out that IFNγ exercise varies throughout AML subgroups and is related to cell lineage and cytogenetics, each essential predictors31,32.
Provided that HLA class 1 and a couple of are main downstream targets of IFNγ signaling33,34, we assessed the correlation of IFNγ signaling with HLA class 1 and a couple of expression. We discovered a major constructive correlation between IFNγ signaling rating and HLA class 1 and a couple of expression (r = 0.56 and 0.52, respectively; p < 2.2 × 10−16) (Fig. 1G, Supplementary Fig. 1B, C). IFNγ signaling rating was additionally related to T cell dysfunction rating, T cell exhaustion rating, and T cell senescence rating, in keeping with the truth that power IFNγ can result in T cell dysfunction24 (Fig. 1G, Supplementary Fig. 1D–F). This discovering steered that an infected immune microenvironment is correlated with IFNγ signaling exercise within the AML bone marrow and the presence of T cell dysfunction.
We subsequent employed immune deconvolution with CIBERSORTx35 to correlate mobile composition with IFNγ signaling exercise. According to the position of IFNγ in selling CD4 T cell activation36,37, we discovered a constructive correlation between IFNγ pathway with helper and regulatory CD4 T cells (Supplementary Fig. 1G). The cell kind most strongly correlated with IFNγ signaling in bulk deconvolution evaluation was with monocytes (r = 0.41; p < 2.2 × 10−16) (Fig. 1H). This was in keeping with prior findings of the enrichment of inflammatory response signaling with monocytic differentiation14. It needs to be famous that deconvolution evaluation can’t differentiate between regular and irregular monocytic cells. We due to this fact hypothesized that this correlation with monocytic cells could also be associated to monocytic differentiation prompting us to look at the respective contributions of AML mobile elements to the expression of IFNγ pathway.
Characterization of IFNγ signaling in newly recognized AML bone marrow aspirates utilizing single cell RNA (scRNA)
To deal with the constraints of bulk RNA profiling in discerning mobile subsets and the relative contributions of cells to IFNγ signaling, we carried out scRNA profiling on 20 bone marrow aspirates from AML sufferers on the time of their analysis. An summary of the demographic, medical, and molecular options of the affected person cohort is supplied in Fig. 2A and Supplementary Desk 1. Briefly, the cohort had a median age of 73 years (vary 52–87), with 4/20 (20%) being feminine and 13/20 (65%) having de novo illness. Seven sufferers had diploid cytogenetics (3/7 non-monocytic, and 4/7 monocytic), 5 sufferers every had del7/7q or deletion in chromosome 5/5q (del5/5q), and three sufferers had each del5/5q and del7/7q (double deletion). Probably the most frequent mutations have been IDH2, NPM1, and DNMT3A in 7/20 (35%) and 6/20 (30%) and 6/20 (30%) of sufferers respectively.
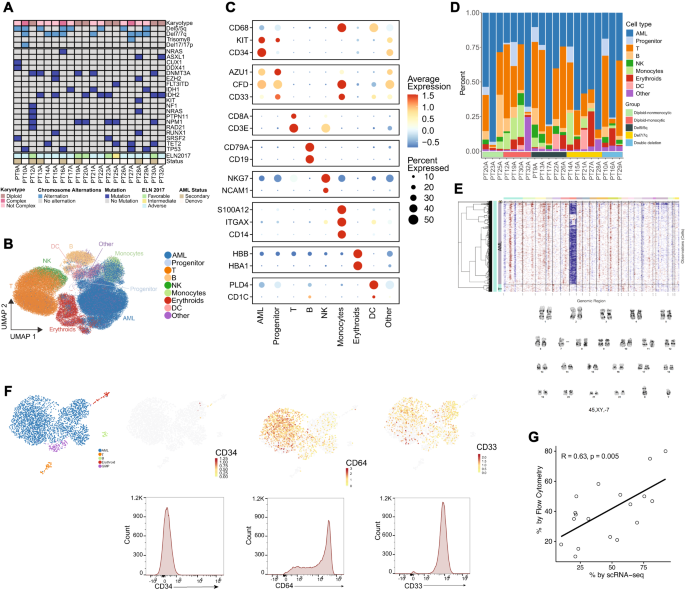
A OncoPrint of sufferers included within the single cell RNA (scRNA) profiling. B UMAP projection of 107,067 cells passing high quality management (see Supplementary Fig. 2 for QC) and cell cluster identities. C Marker gene expression for key marker genes defining cell cluster identities. D Relative proportions of cells by sufferers. E Instance of concordance between inferCNV outcomes and the karyotype of a affected person (PT28A), with a lack of chromosome 7 proven in G-banding karyotype and lack of transcripts akin to chromosome 7 in inferCNV map. F Instance of concordance between gene expression of monocytic AML-defining markers decided by scRNA profiling and protein expression decided by circulation cytometry (CD34 destructive, CD33 constructive, and CD64 constructive) in a consultant affected person with monocytic AML (PT32A). G Concordance of AML blast rely by circulation cytometry and scRNA profiling for all sufferers profiled. T check was used to judge the importance of Pearson correlation. Supply information are supplied as a Supply Knowledge file.
A complete of 107,067 cells handed high quality management and have been additional analyzed (see Strategies and Supplementary Figs. 2, 3). 9 clusters emerged and have been recognized by canonical gene expression: AML cells (52.8%), early progenitors (2.6%), T cells (25.5%), B cells (3.5%), monocytes (3.5%), erythroid cells (5.2%), pure killer (NK) cells (2.2%) and dendritic cells (DC) (1.6%) (Fig. 2B–D, Supplementary Desk 2; Supplementary Knowledge 3). Lower than 3% of cells lacked distinct marker gene signatures and have been labeled along with plasma cells and megakaryocytes as “different”. The non-AML compartment was primarily composed of T cells (53.97% of cells) and confirmed variable distribution throughout cytogenetic teams and sufferers, suggesting the intrinsic heterogeneity of the mobile composition in tumor immune microenvironment of AML sufferers.
To validate the id of AML cells extra particularly, we used infer copy quantity variation (inferCNV) as described in earlier research38. This method efficiently recapitulated the traditional cytogenetic traits of sufferers (Fig. 2E). Moreover, we have been capable of recapitulate monocytic differentiation in AML utilizing gene expression of related marker genes (CD34, CD33, or FCGR1A/CD64) whose protein expression characterization was validated with circulation cytometry (Fig. 2F; Supplementary Desk 3). The proportions of AML cells recognized in scRNA have been much like these decided by histopathologic evaluate and have been positively correlated with the frequency of AML cells decided by circulation cytometry (r = 0.63, p = 0.005) (Fig. 2G). These constant outcomes indicated that scRNA evaluation precisely outlined AML subsets and supplied a dependable foundation for additional downstream evaluation.
Disentangling AML and immune cells on the single-cell stage reveals IFNγ signaling activation in leukemic blasts
Along with AML cells, we targeted our subsequent evaluation on T and NK cells, because the latter cells function the first mediators and effectors for the IFNγ signaling pathway18. We first evaluated the results of IFNγ signaling on the AML cells and tumor microenvironment utilizing AUCell39. We scored each AML and T cells to acquire a single cell-level evaluation of IFNγ signaling exercise. The relative distinction within the expression of IFNγ signaling exercise in T cells among the many cytogenetics teams was smaller than that noticed in AML cells (Fig. 3A). Particularly, we noticed that AML cells in sufferers with diploid monocytic AML had the very best expression of IFNγ signaling scores (ratio of AML to T cells = 0.93), whereas the non-monocytic, diploid AML cells had the bottom (ratio of AML to T cells = 0.56) (Fig. 3A, B, Supplementary Fig. 4A). Amongst nondiploid cytogenetic teams, IFNγ signaling activation was highest amongst these with del7/7q (Fig. 3B). Importantly the commentary of excessive IFNγ signaling in monocytic AML was validated in two unbiased scRNAseq cohorts40,41 (Supplementary Fig. 4B, C). This steered that the variations in IFNγ signaling scores famous in bulk RNA information have been possible pushed by variations inside the AML cells themselves.
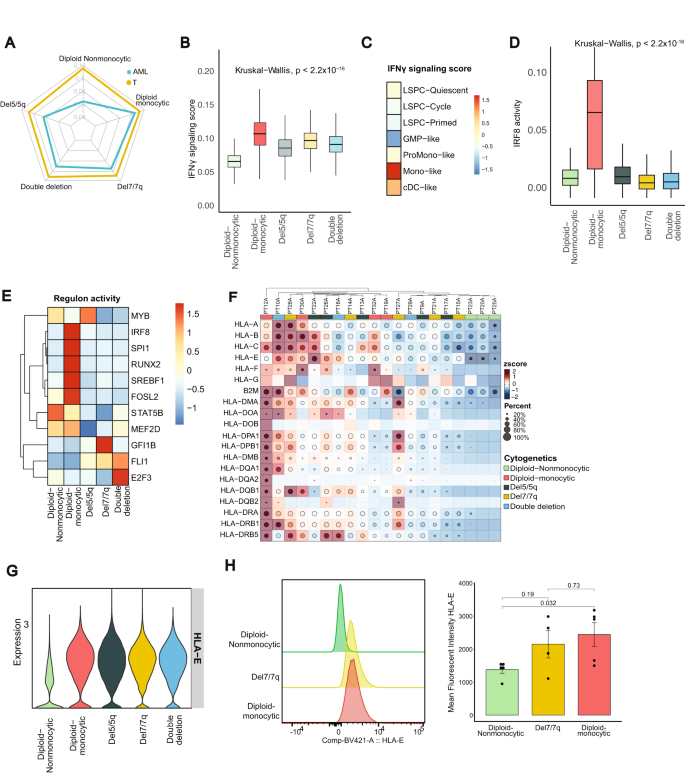
A Radar plot of single cell-level evaluation of IFNγ signaling scored, generated with AUCell, evaluating T cells and AML cells from sufferers with diploid non-monocytic, diploid monocytic, del5/5q, del7/7q, and double deletion (each del5/5q and del7/7q) AML. B IFNγ signaling rating throughout particular AML subgroups (n = 56,168 AML cells; Middle line represents the median and decrease and higher bounds of field correspond to the primary and third quartiles). C IFNγ signaling rating throughout AML differentiation hierarchies as described in Zeng et al. 2022. D Interferon regulator elements 8 regulon exercise throughout AML teams decided by SCENIC (n = 5617; Middle line represents the median and decrease and higher bounds of field correspond to the primary and third quartiles). E Regulon actions of 11 core transcriptional regulators reported by Eagle Ok. et al. visualized by AML teams. F. Heatmap of HLA class 1 and sophistication 2 expression throughout affected person samples. G HLA-E RNA expression in AML subtypes. H Consultant histogram of HLA-E expression in AML blast cells as detected by circulation cytometry (left) and quantification of imply fluorescent depth (MFI) of HLA-E expression between diploid non-monocytic (n = 5), del7/7q (n = 4), and diploid monocytic (n = 5) AML affected person samples (proper; see additionally Supplementary Fig. 4F). Knowledge are offered as imply values ± SD. Supply information are supplied as a Supply Knowledge file.
Zeng et al.32 not too long ago described a mobile hierarchy of AML leukemic stem cells representing distinct maturation states together with LSPC-Quiescent, LSPC-Primed, LSPC-Cycle, GMP-like, ProMono-like, Mono-like and cDC-like. We investigated IFNγ signaling exercise throughout these AML hierarchies and located once more that it was highest amongst cells within the monocyte-like state (Fig. 3C). Zeng et al. additionally revealed 4 distinct subtypes in bulk cohort primarily based on the composition of their leukemia cell hierarchy. We utilized this technique on our bulk cohort and validated the IFNγ signaling exercise in these sufferers, and it was highest within the mature state (enriched for mature Mono-like and cDC-like blasts) whereas lowest in GMP states (Supplementary Fig. 4D). As a result of IFNγ exercise and signaling are epigenetically regulated, we employed SCENIC, a computational instrument for inferring transcription elements from constructed gene regulatory networks utilizing scRNA-seq information39. Our evaluation revealed excessive regulon actions of interferon regulator elements (IRFs) in AML cells, with elevated ranges of IRF1 and IRF5 regulons in del7/7q and del5/5q, respectively (Supplementary Fig. 4E). Notably, we additionally noticed an elevated IRF8, IRF2, IRF3, IRF7 regulon in diploid AML cells with monocytic differentiation (Fig. 3D; Supplementary Fig. 4E), in keeping with their position as a lineage-determinant issue selling monocytic differentiation42,43,44. Of the not too long ago reported 19 core transcriptional regulators of lineage survival in AML45, 11 have been predicted by SCENIC, all of which demonstrated vital variations in regulon exercise throughout cytogenetic teams (Fig. 3E). These observations point out that the activation ranges of IFNγ signaling in AML cells are related to distinct mobile states and hierarchies, and that disparate regulons of IFNγ signaling characterize numerous affected person subgroups.
The antigen presentation equipment, together with HLA expression, are distinguished downstream targets of IFNγ pathway activation and significant for immune recognition. In our evaluation, the expression of HLA class 1 and a couple of genes was additionally notably completely different throughout AML cells from completely different subgroups, the place diploid-nonmonocytic AML sufferers had the bottom expression of HLA genes (Fig. 3F). To validate these findings, we carried out spectral circulation cytometry on 14 peripheral blood samples from AML sufferers at analysis representing 5 sufferers with diploid-nonmonocytic AML, 5 with diploid-monocytic AML, and 4 sufferers with del7/7q AML (Supplementary Desk 4). Curiously, HLA-E, a non-classical class 1 HLA with regulatory capabilities, was extra extremely expressed at each the RNA and protein stage in diploid-monocytic AML blasts than in diploid nonmonocytic blasts, confirming our scRNA-seq findings (Fig. 3G, H, Supplementary Fig. 4F). Nonetheless, circulation cytometric protein expression of HLA class 2 was extremely expressed on most AML samples and was not considerably completely different throughout AML subtypes (Supplementary Fig. 4G, H). These findings, taken collectively, signifies that the monocytic AML microenvironment is characterised by excessive IFNγ signaling that correlates with excessive HLA-E expression, probably representing an immune evasion technique to restrict CD8 and NK cytotoxic exercise by means of inhibitory interactions with NKG2A.
Excessive IFNγ pathway activation in AML cells is correlated with T cell infected microenvironment and distinct regulatory pathway activation
To establish the supply of IFNγ within the AML microenvironment, we assessed IFNG expression in AML and effector immune cell subsets. IFNG was most prominently expressed in CD8 and NK cells with a smaller contribution from CD4 T cells and nearly no expression of IFNG in AML cells (Supplementary Fig. 5A). To comprehensively quantify IFNγ manufacturing, we ran gene set enrichment evaluation on AML, CD4, CD8, and NK cells with the IFNγ manufacturing gene set from GO. Once more, we discovered that CD8 T cells confirmed the very best manufacturing exercise adopted by NK cells whereas the AML cells had the bottom exercise (Fig. 4A). To additional elucidate the immune microenvironmental variations amongst cytogenetic and phenotypic AML teams, we assessed the T and NK cell composition inside every group. Curiously, diploid non-monocytic AML had the bottom infiltration of T cells and had the bottom proportion of NK cells with expression of the inhibitory receptor NKG2A which binds to HLA-E to suppress the cytolytic exercise of NK cells (Supplementary Fig. 5B–F)46.
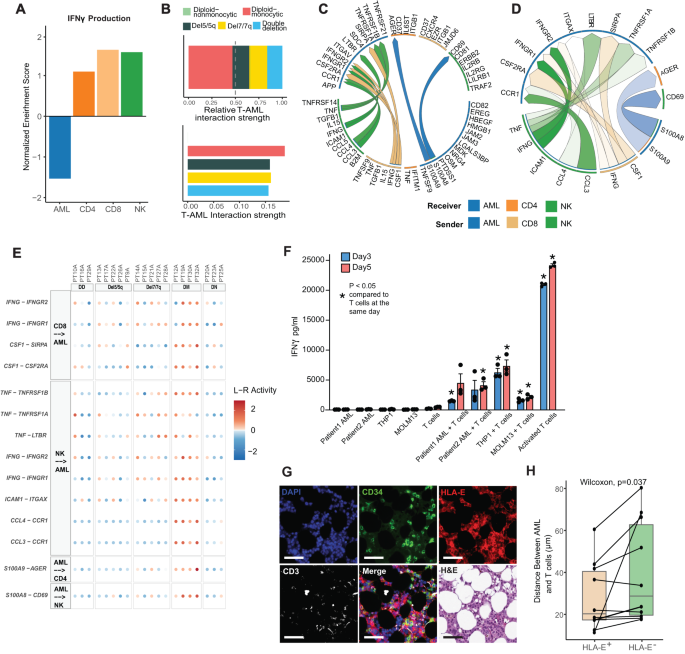
A Normalized enrichment rating of GSEA of IFNγ manufacturing pathway for AML, CD4, CD8, and NK cells. B T cell-AML interactions and interplay power amongst AML teams predicted by CellChat. C Circos plot of the highest predicted non-AML-to-AML ligand-receptor interactions inside the diploid monocytic subset among the many prime 100 ligand-receptor interactions predicted by MultiNicheNet. D Circos plot of the highest 50 ligand-receptor interactions inside the diploid monocytic subset solely. E Dot plot of prime 50 ligand-receptor interactions inside the diploid monocytic subset solely, excluding AML-to-AML interactions. DD double deletion, DM diploid-monocytic, DN diploid-nonmonocytic. F IFNγ ranges in tradition media have been assessed by ELISA after a 72 and 120-h co-culture of CD14+, CD34 + AML blasts, THP1 cells or MOLM13 cells with wholesome donor T cells. Management teams included blasts alone, T cells alone, and T cells co-cultured with DynaBeads T cell activator. Co-culture information is from 1 affected person with CD14+ blasts, 1 affected person with CD34+ blasts, THP1 cells or MOLM13 cells with every situation having 3 replicates. Knowledge are offered as imply values ± SEM. Two-sided t check was used (*p < 0.01). G Consultant multiplex IF panel from 2 affected person bone marrow biopsy samples imaged utilizing Lunaphore exhibiting a consultant picture enriched for CD34+ AML cells, serial part stained with DAPI, CD34, HLA-E, and CD3, H&E, and merged part proven for comparability with scale bar indicating 50 µm. The experiment was repeated 10 instances. H Boxplot of distance between CD3 T cells and AML cells by whether or not AML cells specific HLA-E or not. (n = 10; Middle line represents the median and decrease and higher bounds of field correspond to the primary and third quartiles). Two-sided Wilcoxon check was used. Supply information are supplied as a Supply Knowledge file.
To additional elucidate the cell-cell interactions within the immune microenvironment we employed the CellChat47 and MultiNicheNet instruments48. CellChat quantifies interplay power by aggregating the communication possibilities throughout all cell group pairs. We noticed a considerably greater communication chance between T cell-AML interactions in diploid monocytic AML in comparison with different teams (P < 0.0001) and no predicted T cell-AML interactions in diploid non-monocytic AML (Fig. 4B, prime). This commentary held when the power was scaled to account for confounding elements (Fig. 4B, backside). We then employed MultiNicheNet to foretell the highest 100 receptor-ligand interactions amongst AML, CD8 T, CD4 T, and NK cells (Supplementary Knowledge 4). Curiously, IFNγ from CD8 T cells and NK cells was predicted as a prime interplay with AML cells just for sufferers with diploid monocytic AML (Fig. 4C, Supplementary Fig. 5G–J). Of be aware, del7/7q AML additionally confirmed indicators of each a pro- and anti inflammatory microenvironment with distinguished predicted interactions involving TNFα and the TGFβ pathway (Supplementary Fig. 5I). Given these information suggesting an infected microenvironment as a distinguished function of diploid monocytic AML, we additional analyzed receptor-ligand interactions within the diploid monocytic group solely (Supplementary Knowledge 5). This confirmed that IFNγ from CD8 T and NK cells performing on AML cells as a distinguished function on this group of sufferers (Fig. 4D, E). Curiously, this evaluation additionally predicted different distinguished inflammation-related interactions together with TNFα, CCL3, CCL4, and the SIRPα pathway which is an rising drug goal in AML49 (Fig. 4E). To evaluate the capability of AML cells to instantly induce INFγ secretion, we carried out co-culture assays of AML cells with T cells in main cells of two AML sufferers and two AML cell strains, MOLM13 and THP1. Our evaluation revealed that in isolation, AML cells didn’t secrete INFγ, whereas T cells had a low basal stage of INFγ secretion (Fig. 4F). In distinction, when AML cells have been co-cultured with T cells, there was a constant and vital elevation within the ranges of INFγ detected within the supernatant at each assessed time factors relative to T cells in isolation (Fig. 4F). This improve in INFγ ranges upon co-culture means that AML-T cell interactions is enough for INFγ manufacturing. Of be aware, sturdy T cell activation with agonistic anti-CD3/CD28 stimulation was related to a significantly higher improve in INFγ secretion from T cells in comparison with the co-culture assay (Fig. 4F). This substantial improve means that, whereas direct AML-T cell contact is able to initiating INFγ manufacturing, different elements might contribute to amplifying or dampening T cell activation. To additional validate T cell-AML interactions, we employed COMET primarily based multiplex immune-fluorescence (IF) panel to spatially quantify the bone marrow microenvironment in 2 AML bone marrow samples. Utilizing this expertise we confirmed HLA-E+CD34+ AML cells are in nearer proximity to CD3+ T cells than that of HLA-E– AML cells, suggesting an immunosuppressive neighborhood between HLA-E expressing AML cells and T cells (Fig. 4G, H; Supplementary Desk 5). Taken collectively, these findings recommend that IFNγ signaling in AML which is probably going instigated from neighboring T/NK cells confers immune-evasion by way of upregulation of HLA-E.
Excessive IFITM3 expression predicts worse survival in AML
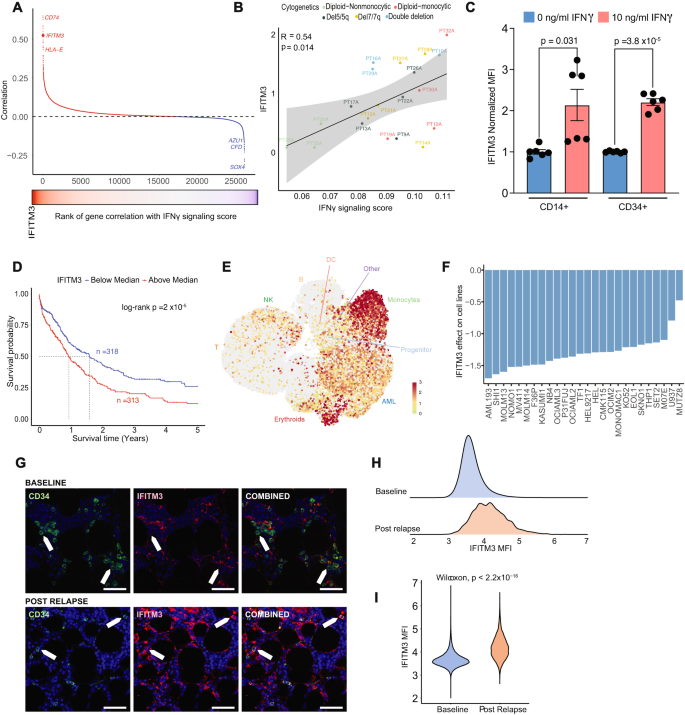
To delineate the results of IFNγ signaling in AML and its dependency genes, we calculated correlations of IFNγ signaling rating with all expressed genes within the single cell information. The highest genes positively correlated with IFNγ signaling rating included CD74, IFITM3, and HLA-E (Fig. 5A, Supplementary Knowledge 6). According to our findings on mobile hierarchy (Fig. 3C), markers for GMP cells (AZU1, CFD) had sturdy destructive correlation with IFNγ signaling rating (Fig. 5A). The typical rating at a affected person stage additionally confirmed a major affiliation between IFNγ and IFITM3 (Fig. 5B). IFITM3 encodes an IFNγ-induced protein which has been steered to play a job in tumor development of a number of cancers together with B-ALL, mantle cell lymphoma, colorectal, prostate, and hepatocellular carcinoma50,51. The excessive correlation of IFITM3 expression with IFNγ signaling prompted us to judge whether or not IFNγ instantly stimulates the expression of IFITM3 in AML cells. We remoted CD14+ and CD34+ AML blasts from two sufferers every and carried out IFNγ stimulation for twenty-four h, adopted by IFITM3 protein stage measurement by way of circulation cytometry. Certainly, IFNγ induced IFITM3 expression in each CD14+ and CD34+ leukemic blasts by 2.1 (p = 0.0312) and a couple of.2 (p < 0.0001) fold improve, in comparison with unstimulated CD14+ and CD34+ cells, respectively (Fig. 5C). This establishes a direct hyperlink between IFNγ and IFITM3 expression in AML blasts. Cox proportional hazard and Kaplan-Meier fashions of AML sufferers built-in from the TCGA, BeatAML and MDACC cohorts revealed that sufferers with excessive IFITM3 (>median) expression had considerably worse general survival (Fig. 5D; Supplementary Fig. 6A–D). Excessive IFITM3 expression remained considerably related to worse general survival with changes for age, blast share, and cytogenetic danger in a multivariable cox mannequin (Supplementary Fig. 6E). Of be aware, IFITM3 was nearly solely expressed in AML cells, with the very best expression in monocytes and restricted expression in T cells and different microenvironmental cells (Fig. 5E). This implies that IFITM3 is a downstream goal of IFNγ, and elevated expression provides priceless prognostic data to stratify newly recognized AML affected person outcomes.
A Correlation of IFNγ signaling rating with gene expressions within the single cell information. Genes in crimson have a constructive correlation and genes in blue have a destructive correlation. B Correlation of IFITM3 expression with IFNγ signaling rating at a person affected person stage. Error band represents 95% confidence interval. T check was used to judge the importance of Pearson correlation. C IFITM3 expression in AML blasts was assessed by circulation cytometry after a 24-h stimulation with 10 ng/ml IFNγ. Knowledge represents outcomes from 2 sufferers’ CD14+ blasts and a couple of sufferers’ CD34+ blasts, with every situation having 3 replicates. The error confirmed normal error of imply. Two-sided t check was used. D Kaplan-Meier survival curve of the AML sufferers from TCGA, Beat-AML and MDACC by median IFITM3 expression within the bulk RNA profiling information. E UMAP projection of all sufferers’ cells scored by IFITM3 expression in cells. F Change in cell line health following genetic deletion of IFITM3 within the DepMap CRISPR knockout display screen information, exhibiting leads to the 26 AML cell strains examined. G Consultant multiplex IF panel. Baseline AML pattern with CD34 constructive blasts present low quantities of IFITM3(Prime). Put up relapse AML pattern with CD34 constructive blasts present excessive quantities of IFITM3 (Backside). Inexperienced is CD34, crimson is IFITM3, blue is DAPI. Scale bar 50 μm. The experiment was repeated 2 instances. H Density plot of IFITM3 fluorescence depth on AML blasts at analysis and publish relapse. I Violin plot of fluorescence depth on AML blasts at analysis and publish relapse. Put up relapse AML pattern present considerably greater quantities of IFITM3. Two-sided Wilcoxon check was used. Supply information are supplied as a Supply Knowledge file.
We subsequent sought to validate the potential affect of IFITM3 loss in AML cell strains. From the DepMap Public Dataset, which consists of 1,086 cell strains and 17,386 genes, we targeted our evaluation on 26 AML cell strains and 188 genes that overlapped with the IFNγ response signature within the HALLMARK gene set52. Unsupervised clustering utilized to the gene impact recognized a cluster of seven genes (WARS1, IFITM3, NAMPT, PSMA2, NUP93, PSMB2, and PSMA3) that persistently decreased the health of AML cells after knockout (Supplementary Fig. 6F, G). Specifically, the knockout of IFITM3 precipitated the cell demise in all AML cell strains examined (Fig. 5F). The importance of this variation was verified by a two-sided t check evaluating the health rating of every gene to these of the remaining 187 genes. To additional consider its impact on AML cells, we used multiplex immunofluorescence to measure the fluorescence depth of IFITM3 on blasts in baseline and publish relapse samples. A complete of 15 TMA cores have been stained for CD34, CD56, CD45, CD4, CD14, and IFITM3 primarily based on markers decided by circulation to establish blasts (Fig. 5G; Supplementary Desk 5). The fluorescence depth in publish relapse samples was considerably greater than in baseline samples (p < 2.2 × 10−16) (Fig. 5H, I), supporting the affiliation of IFITM3 with illness resistance. These information recommend IFITM3 as a possible dependency in AML cells and may very well be related to therapeutic resistance. Nonetheless, extra experimental information shall be additional wanted to validate the position of IFITM3 as a mediator for drug resistance.
IFNγ signaling confers venetoclax resistance
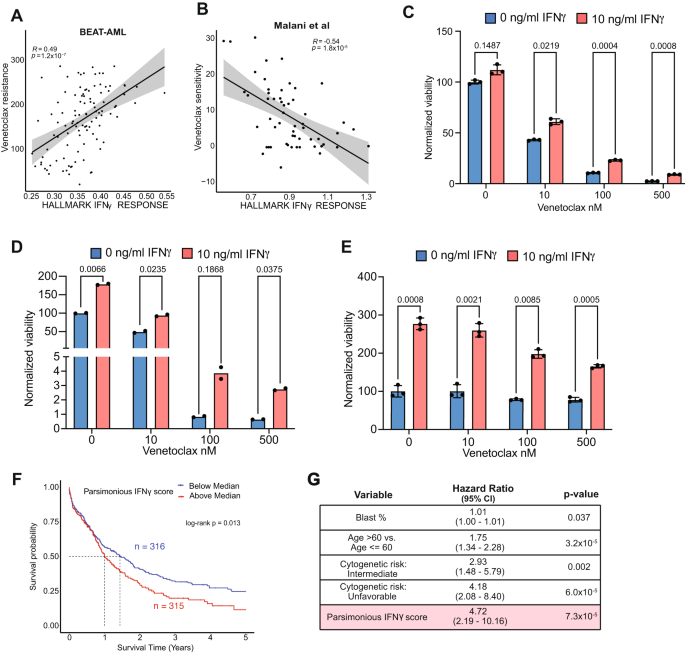
We subsequent assessed the correlation of IFNγ with drug response. We leveraged the BEAT-AML ex vivo drug sensitivity information which supplied detailed matched medical, genomic, and transcriptomic analyses3 (Supplementary Fig. 7A). Provided that AML sufferers with monocytic differentiation and del7/7q are reported to have resistance to venetoclax-based remedy10,53, we evaluated the correlation of IFNγ signaling rating with drug sensitivity. We discovered a robust constructive correlation between IFNγ signaling rating and venetoclax resistance, indicating that IFNγ signaling confers venetoclax resistance (Fig. 6A). This discovering was validated in an unbiased cohort54 of ex vivo drug screening in AML, the place sufferers with excessive IFITM3 expression or IFNγ signaling rating have been much less delicate to venetoclax-based remedy (i.e. extra proof against venetoclax) (Fig. 6B). To additional validate the position of IFNγ signaling in AML cell survival and drug resistance, we remoted leukemic blasts from main affected person samples (n = 3 sufferers) and cultured them within the absence or presence of IFN-γ, and with growing concentrations of venetoclax then assessed the AML cell viability. IFNγ induced proliferation and better resistance to venetoclax, confirming that IFNγ promotes survival and venetoclax resistance of AML blasts (Fig. 6C–E).
A Correlation of IFNγ signaling rating and venetoclax resistance in BEAT-AML information. Error band represents 95% confidence interval. T check was used to judge the importance of Pearson correlation. B Correlation of IFNγ signaling rating and venetoclax resistance in Malani et al. information. Error band represents 95% confidence interval. T check was used to judge the importance of Pearson correlation. C–E Viability evaluation of AML blasts after stimulation with 10 ng/ml IFNγ and incubation with venetoclax utilizing the CellTiter-Glo luminescent cell viability assay. Knowledge represents outcomes from 3 AML sufferers’ blasts, with every situation having 3 replicates, aside from (D), which has two replicates. For (C, E), information are offered as imply values ± SEM. Two-sided t check was used. F Kaplan-Meier survival curves of the AML sufferers from TCGA, Beat-AML, and MDACC by parsimonious IFNγ rating in bulk RNA profiling cohort. G Multivariable adjusted Cox regression mannequin for general survival among the many sufferers within the mixed bulk RNA profiling cohort adjusting for parsimonious IFNγ rating, age, blast share, and cytogenetics. Wald check was used to measure the importance of things. Supply information are supplied as a Supply Knowledge file.
Regardless of its correlation with venetoclax resistance, the IFNγ signaling rating didn’t predict survival outcomes in our bulk cohort (Supplementary Fig. 7B), suggesting that the entire gene set might include genes which are much less delicate for predicting outcomes. Subsequently, to enhance the prognostic sensitivity of the IFNγ signaling rating, we outlined a lightweight weighted IFNγ signature utilizing the least absolute shrinkage and choice operator (LASSO) mannequin55 (Supplementary Fig. 7C). After LASSO regression, 47 genes associated to survival have been retained, forming a parsimonious IFNγ signature (Supplementary Desk 6). We then scored the majority AML sufferers information with the brand new parsimonious IFNγ rating and revealed a good constructive correlation with HLA class 1 and a couple of scores (Supplementary Fig. 7D, E). Importantly, this parsimonious IFNγ rating was capable of predict affected person outcomes in our bulk cohort, whereby the next rating predicted worse survival (Fig. 6F, Supplementary Fig. 7F). This discovering of worse general survival was additional validated in an unbiased dataset54 (Supplementary Fig. 7G). The parsimonious IFNγ rating remained a major and powerful predictor of survival in a multivariate mannequin once we accounted for age, blast share, and cytogenetics, with the very best hazard ratio of all these predictors (Fig. 6G). Finally, these outcomes recommend that the IFNγ pathway is related to resistance to venetoclax-based remedy and might predict affected person outcomes independently of recognized danger elements, making it a promising goal for therapeutic intervention.