Protein selector units (PS) establish affected person teams with distinct scientific outcomes
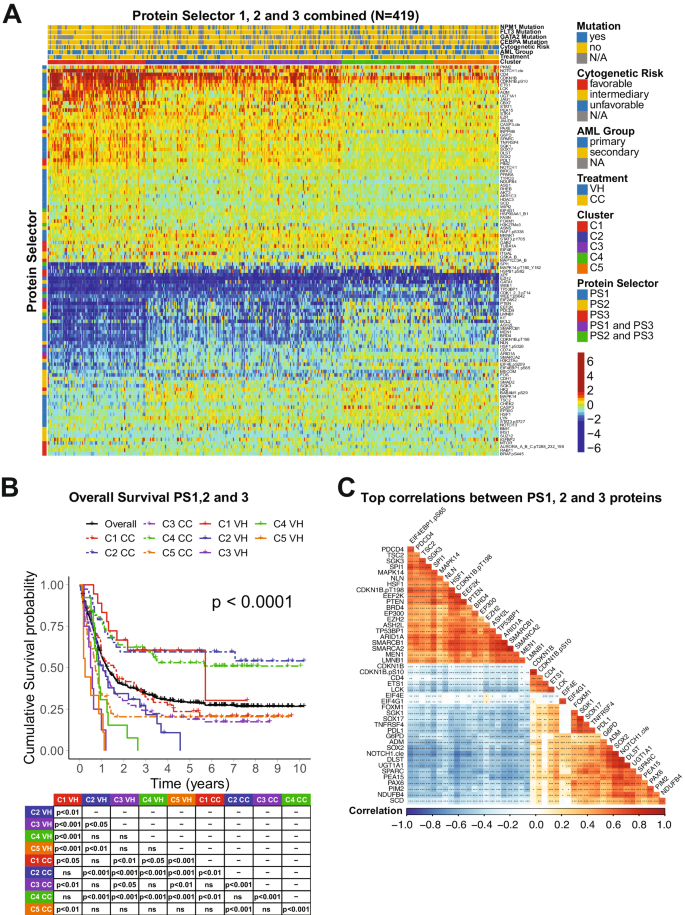
We developed an algorithm to establish probably the most therapeutically discriminating proteins and generated Protein Selector Units (see “Supplies and Strategies” part). The primary one, entitled PS1, was comprised of 55 proteins, which recognized three clusters (C1, C2, and C3) with distinctive expression signatures. Protein ranges throughout the clusters are proven in Fig. 1A. Though the protein signature of every cluster was the identical in each sufferers with VH and CC, their total survival (OS) different tremendously between therapies. As proven in Fig. 1B, sufferers in C1 (pink) handled with VH (stable line) had diametrically completely different and superior responses in comparison with these handled with CC (dashed line), with a Median OS (MS) of 68.5 months (mo.) within the VH group versus (vs.) MS of 19.4 mo. within the CC inhabitants. The alternative was true for C3 (yellow), the place CC sufferers had a MS of 16.8 mo. and the VH inhabitants displayed a really poor MS of 8.7 mo. Nonetheless, PS1 didn’t establish an optimum remedy for sufferers in cluster C2 (mild blue). Due to this fact, to establish the popular remedy for PS1-C2 sufferers (N = 182), we generated PS2, utilizing the identical technique described beforehand. As proven in Fig. 1C, PS2 separated the inhabitants into two clusters with distinct expression profiles. In Fig. 1E, cluster PS2-C1 (blue shade) handled with CC (dashed line) had a markedly higher OS (>120 mo.), in comparison with C1-VH (stable blue), which has a MS of 12.7 mo. The identical was true for cluster PS2-C2 (purple shade), the place CC (dashed line) had a MS 12.2 mo., and VH (stable line) had a MS of 6.4 mo. Furthermore, as proven in Fig. 1B, the very best PS1-C3 curve (dashed yellow, CC-treated) has an OS similar to the worst PS1-C1 group (dashed pink, CC-treated). Due to this fact, we generated a PS3 for PS1-C3 sufferers (N = 146) in an try and establish a bunch with higher OS. Inside PS3, two clusters with contrasting protein expression ranges had been outlined, and separated by therapy (Fig. 1D). As proven in Fig. 1F, sufferers in cluster PS3-C1 (inexperienced shade) had an excellent prognosis when handled with CC (dashed line), with MS > 120 mo., and a really poor final result when handled with VH (stable line), having a MS of 10.4 mo. In distinction, OS of sufferers in PS3-C2 (orange shade) had been equally poor for each therapies.
A Heatmap depicting the protein expression of PS1 sufferers (N = 419). B Kaplan–Meier plots of Total Survival PS1 sufferers (N = 419) separated by cluster and therapy modality (VH = stable line, CC = dashed line; PS1-C1 = pink, PS1-C2 = mild blue, PS1-C3 = yellow). C Heatmap depicting the protein expression of PS2 sufferers (N = 182) and D PS3 sufferers (N = 146). E Kaplan–Meier plots of Total Survival from PS2 sufferers (N = 182) and F PS3 sufferers (N = 146) separated by cluster and therapy modality (VH = stable line, CC = dashed line; PS2-C1 = blue, PS2-C2 = purple, PS3-C1 = orange, PS3-C2 = inexperienced). Annotations above the heatmaps, beginning closest to the heatmap, present the clusters, VH vs. CC therapy modality (second from backside), after which different annotations for a number of beforehand acknowledged prognostic options together with AML group, cytogenetic danger, and presence of complicated karyotype and mutations. Colours for the annotations have the worth proven within the legends alongside the fitting facet. Protein expression starting from above regular (pink) to regular (yellow-green-aqua) to beneath regular (darkish blue) as proven within the shade legend.
The mixture of the PS units led to the technology of 5 clusters separated by the expression ranges of 109 proteins as proven in Fig. 2A. C1 derived from PS1, C2 and C3 from PS2 (former PS2-C1 and PS2-C2), and C4 and C5 from PS3 (former PS3-C1 and PS3-C2). In Fig. 2B, the OS was higher for C1 sufferers (pink) handled with VH (stable) in comparison with CC (dashed) (MS = 68.5 mo. vs. 19.4 mo.). In distinction, each C2-CC (dashed blue) and C4-CC (dashed inexperienced) displayed MS > 120 mo., outperforming each C2-VH (stable blue), with a MS of 12.7 mo., and C4-VH (stable inexperienced), which has a MS of 10.4 mo. Furthermore, though C3-CC (purple dashed) do higher than C3-VH (purple stable) (MS of 12.2 mo. vs. 6.4 mo.), their OS are worse than the C2-CC and C4-CC populations. Lastly, our PS system couldn’t decide which therapy sufferers in cluster C5 (orange) ought to obtain. Contemplating their poor outcomes in each VH (MS = 2.9 mo.) and CC (MS = 8.6 mo), plainly this inhabitants would possibly profit from one other therapy routine (e.g., target-based therapies). Evaluation of CRD for all PS units confirmed an identical final result sample (Supplementary Fig. S1). Comparability of VH vs. CC for every cluster individually is proven in Supplementary Fig. S2.
A Heatmap depicting the protein expression of all sufferers (N = 419). Annotations above the heatmap, beginning closest to the heatmap, present the cluster membership and therapy modality, after which different beforehand acknowledged prognostic options (AML group, cytogenetic danger, and presence of complicated karyotype and mutations). Legends are as described in Fig. 1. B Kaplan–Meier plots of Total Survival and C Prime correlations between all of the PS proteins (N = 45). Squares signify the correlation between every protein are coloured in line with the diploma of the linear correlation, which varies between (−1, 1) and follows a ‘blue’ (−1), ‘white’ (0), and ‘pink’ (1) gradient, as proven within the shade legend. Important correlations are highlighted in line with the next: ***p < 0.001, **p < 0.01, *p < 0.05, and clean = not vital.
To raised assess the organic that means of the PS analyses, we evaluated the correlation of the expression ranges of the 109 prognostic proteins between one another. In Fig. 2C, the highest most correlated proteins, outlined as having a correlation coefficient > 0.60, are proven. Among the many organic processes associated to these proteins, the commonest had been ribosomal and transcriptional exercise (10 proteins), histone modifiers (8 proteins), cell cycle and DNA injury response (7 proteins), cell metabolism (6 proteins). For an expanded view of those protein relationships, the whole correlation plot, along with protein networks of the PS proteins divided by useful group are proven in Supplementary Fig. S3. The correlation coefficients for all proteins, together with p-values of every comparability are proven in Supplementary Desk S4. The stratification of all 109 proteins by organic course of with their respective Protein Selector Set is proven in Supplementary Desk S5.
Clusters associations with demographic, scientific, and molecular options
We examined how the clusters differed contemplating demographic (age, gender, race), scientific (AML group and laboratory parameters), and molecular options (cytogenetics and mutation profiles), as proven in Desk 1. There have been vital variations in age distribution, in addition to the frequency of many scientific variables (main vs. secondary AML, white blood cell depend, proportion of blasts and platelets quantity), cytogenetics (by danger group, easy vs. complicated karyotype, or for particular occasions, resembling −5/5q-, −7/7q- and inv16), and for a number of particular person mutations (ASXL1, CEBPA, DNMT3A, EZH2, FLT3 [individually for ITD and D835, and in combination], NPM1, and TP53). An expanded desk with all variables assessed is proven in Supplementary Desk S6.
Since many of those options with unbalanced distributions among the many clusters are recognized to be prognostic, we questioned whether or not the cluster prognostic impression was only a reflection of those imbalances or if the clusters had been independently predictive. Right here, we generated KM plots to confirm whether or not cluster membership is prognostic for OS and CRD when the inhabitants is filtered for particular variables (e.g., males solely, secondary AML solely, and so forth.). KM plots with p-values are proven in Supplementary Figs. S4 and S5. The prognostic impression of the 5 clusters was sustained for nearly all of the variables, together with gender, all three age teams, all races, each main and secondary AML, and main cytogenetic groupings (whether or not divided into three prognostic teams or for complicated karyotypes). Since most particular person cytogenetic and mutation occasions happen at a low frequency when the 5 clusters are subdivided by therapy modality (ten teams in whole), the small pattern sizes usually preclude reaching statistical thresholds. Nonetheless, comparable developments (C1, C2, and C4, higher than C3 and C5) had been maintained for almost all, with exceptions famous for FLT3, IDH1, IDH2, JAK2, MLL, PTPN11, and TP53 mutations.
Subsequent, we measured the prognostic worth of the clusters and different variables utilizing univariate (UV) and multivariate (MV) Cox proportional-hazards fashions (CoxPH) for each OS and CRD. In each analyses, clusters had been condensed into three teams to keep away from a lot of ranges in a single variable, which could negatively affect the CoxPH fashions. Due to this fact, clusters with good prognosis (C1-VH, C2-CC, and C4-CC) had been joined and renamed Group1; those with intermediate OS and CRD (C1-CC, C2-VH, C3-CC) had been compacted into Group2; and at last, the remaining clusters, with poor prognosis, (C3-VH, C4-VH, C5-VH and C5-CC) had been merged into Group3. As demonstrated in Desk 2, all cluster teams had been predictive of survival and remission in each the UV and MV fashions, reinforcing their prognostic worth. Furthermore, a number of demographic (age, white race, and Asian race), scientific (secondary AML, blasts, Hbg, and serum B2M), cytogenetic (complicated karyotype, −5/5q-, −7/7q-, t(8;21), Inv16, and Del12), and mutational (ASLX1, CEBPA, FLT3 [individually for ITD and D835, and in combination], IDH2, JAK2, MLL, NPM1, PTPN11, and TP53 mutations) options had been additionally prognostic within the UV mannequin for OS. Nonetheless, solely clusters, secondary AML, complicated karyotype, Inv16, and IDH2 and PTPN11 mutations remained vital within the MV evaluation. Concerning CRD, within the UV evaluation clusters remained extremely vital together with different traits (age, black race, AML group, complicated karyotype, −5/5q-, Inv16, and FLT3, RUNX1, and TP53 mutations), with solely clusters, black race, and sophisticated karyotype, which remained vital within the MV mannequin. Taken collectively, these findings corroborate the unbiased prognostic worth of the PS protein signatures. An expanded desk containing all variables evaluates within the UV mannequin for each OS and CRD is proven in Supplementary Desk S7.
Improvement of a protein classifier (PC) for therapy suggestion
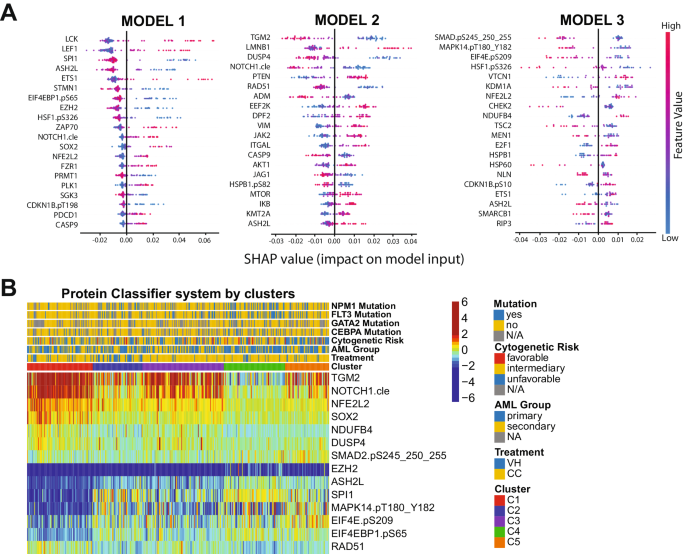
Though the PS system can effectively separate sufferers who ought to obtain VH from those that would do higher with CC, it isn’t possible to measure greater than 100 completely different proteins within the scientific setting. The variety of proteins required to be assessed is extreme and poses a serious cost-benefit problem for the appliance of the tactic. As an alternative, the identification of some proteins that may be measured utilizing a Scientific Laboratory Enchancment Amendments (CLIA)-certified check to precisely assign a person affected person to a particular protein expression profile is sensible. Due to this fact, we designed a classification algorithm utilizing the random forest machine studying approach entitled Protein Classifier (PC). The system can establish probably the most predictive proteins for therapy suggestion, primarily based on beforehand developed cluster memberships and protein expression information. In different phrases, we really helpful VH therapy for sufferers belonging to cluster C1 (N = 91); CC remedy for sufferers in clusters C2, C3, and C4 (N = 267); and neither VH nor CC for the C5 affected person inhabitants (N = 61). The system was developed with the aim of defining clusters utilizing three completely different fashions sequentially:(1) Outline C1 sufferers (N = 91); (2) Distinguish C2 and C4 teams (N = 154) from the C3 and C5 populations (N = 174); and (3) Separate C3 (N = 113) from C5 (N = 61) sufferers. In Fig. 3A, the highest predictive proteins are visualized along with their respective SHAP values. Step one of the PC system recognized the six most predictive proteins for C1: SPI1, ASH2L, EIF4EBP1.pS65, EZH2, NFE2L2 and SOX2 (C-index: 0.951). Thus, in line with our earlier OS and CRD analyses, sufferers with this protein signature ought to obtain VH remedy. Within the second step of the PC system, TGM2, NOTCH1.cle, DUSP4, and RAD51 had been the very best proteins to distinguish C2 + C4 from C3 + C5 (C-index: 0.903). Of observe, distinguishing C3 from C2 and C4 is critical, as a result of though each affected person teams ought to obtain CC, the OS and CRD for C3 is far decrease, so this affected person group might profit from further remedy (e.g., CC and stem cell transplant in first remission), whereas C2 and C4 appear to do nicely with CC alone. Lastly, SMAD2.pS245_250_255, MAPK14.pT180_Y182, EIF4E.pS209, and NDUFB4 had been recognized as the very best proteins to segregate C3 and C5, defining the final step of our system (C-index:0.923). The expression of all proteins within the PC system by cluster is proven in Fig. 3B. Importantly, the C-index, a measure of particular person affected person discriminatory energy, of all fashions in our PC system is above 0.90, demonstrating that it robustly predicts optimum remedy alternative (a C-index larger than 0.7 is taken into account predictive, whereas a measure of 1 would point out perfection). Furthermore, by contemplating all three fashions working collectively, we predicted that 87.3% of sufferers would obtain the proper remedy, and solely a small fraction of 5.5% can be misassigned. The proportion of sufferers within the C5 group who could possibly be assigned to both CC or VH, as an alternative of being outlined as ‘undetermined’, was 7.1%. Total sensitivity, specificity, and accuracy had been 84.2%, 79.6%, and 82.8%, respectively. The predictive calculations for the PC mannequin are introduced in Supplementary Desk S8. Due to this fact, the event of a package that determines the expression of the aforementioned 14 proteins can be helpful and financially possible for triaging sufferers and guiding the advice for VH or CC.
A Prime predictive proteins for every of the three fashions developed (y-axis), for all test-set sufferers, in line with every protein’s calculated SHAP worth (x-axis). Colour legend signifies the worth of a given prognostic protein’s expression relative to different expression values of that protein amongst test-set sufferers. (pink = excessive predictive worth; blue = low predictive worth). B Heatmap displaying the expression ranges of the proteins chosen for the protein classifier by cluster and therapy modality. Annotations and legends are as described in Figs. 1–3.
Sufferers with the worst outcomes have a novel and targetable protein signature
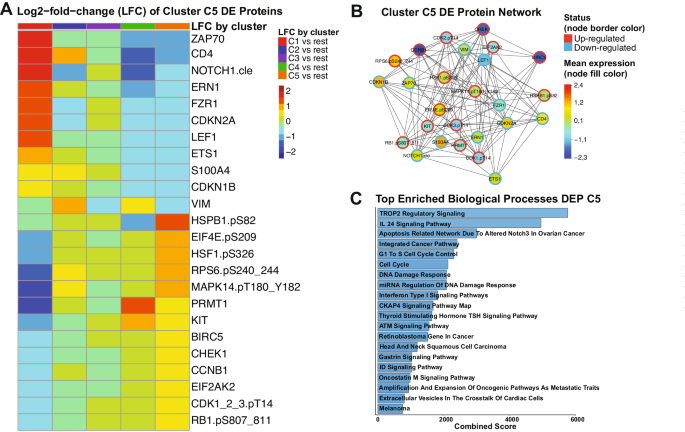
Since our PS system was unable to suggest both VH or CC for cluster C5 sufferers, we determined to find out probably the most related signaling pathways inside this inhabitants. We recognized 24 proteins among the many 411 in our database which together type a novel expression profile in C5 sufferers, in comparison with all the opposite clusters. In Fig. 4A, the Log2-fold-change (LFC) values of every every cluster towards all of the others is proven for every differentially expressed (DE) protein of cluster C5. Proteins from ZAP70 till VIM have decrease LFC values and, thus, had been thought-about down-regulated in C5, whereas the proteins from HSPB1.pS82 to RB1.pS807_811 had been categorized as up-regulated since their LFC values are larger in C5 in comparison with the others. A desk with FDR-adjusted p-values and LFC values evaluating every cluster towards all of the others is proven in Supplementary Desk S9. To raised visualize connections of the C5 DE proteins with one another, we generated a protein community, annotating the imply expression values of every one in comparison with regular bone marrow (node fill shade), and whether or not the protein is up- or down-regulated (node border). Importantly, though a number of proteins are up-regulated in comparison with the opposite clusters, their imply expression is beneath the degrees of regular bone marrow (e.g., CHEK1, BIRC5, CCNB1). A desk with all of the DE proteins and their directionality (up- or down-regulated), stratified by cluster is in Supplementary Desk S10. Volcano plots highlighting the directionality of DE proteins for each cluster are proven in Supplementary Fig. S6.
A Heatmap demonstrating the imply Log2-fold change (LFC) values of each assessed comparability, concerning the 24 proteins which are differentially expressed in cluster C5 sufferers. Heatmap annotation check with the comparability as established by the legend on the fitting. The imply LFC ranges in line with the values and shade legend on the fitting B Protein community of DE proteins of Cluster C5. Community nodes are coloured in line with the imply expression worth, starting from above regular (pink) to regular (yellow-green-aqua) to beneath regular (darkish blue) as proven within the shade legend (proper). Node borders are coloured in line with the Differential Expression standing of protein (up-regulated or down-regulated), following the colours proven within the legend (proper). C Prime twenty enriched organic processes associated to the differentially expressed proteins of cluster C5. Y-axis reveals the title of every ontology and x-axis reveals the mixed scores of every course of. Bargraphs are coloured in line with a blue gradient, the place darker blue corresponds to decrease values and lighter blue to larger values.
To achieve insights concerning the organic that means of our information, we carried out pathway enrichment evaluation of the 24 DE proteins. As proven in Fig. 4C, processes with the best mixed scores (i.e., lowest p-value and highest odds-ratio) had been most importantly correlated to those proteins. Most of these had been associated to cell cycle regulation and the DNA injury response (DDR), however particular pathways had been additionally enriched (e.g., TROP2, IL-24, and CKAP4 signaling). The whole desk with all of the processes and their mixed scores, together with adjusted p-values and odds ratios could be present in Supplementary Desk S11. Altogether, though we had been unable to suggest a particular therapy for C5 sufferers, our DE evaluation revealed potential druggable signaling pathways that could possibly be helpful for growing target-based therapies.