ScRNA-seq and spatial atlas of NSCLC samples
To find out the heterogeneity of immune and non-immune mobile states and their spatial panorama in LUAD and LUSC, we collected lung tissue resections from 25 treatment-naive sufferers with both LUAD (n = 13), LUSC (n = 8) or undetermined lung most cancers (LC, n = 4), and two wholesome deceased donors (Fig. 1A, B and Supplementary Information 1). We collected each tumour and matched regular non-tumorigenic tissue (i.e., background), remoted CD45+ immune cells (Supplementary Fig. 1A) in addition to tumour and different non-immune populations (utilizing CD235a column to deplete erythroid cells), and carried out scRNA-seq. As well as, tumour and background tissue sections from eight sufferers (of the aforementioned 25) have been processed for spatial transcriptomics utilizing the 10x Genomics Visium platform (n = 36 sections in complete) (Fig. 1A and Supplementary Information 1).
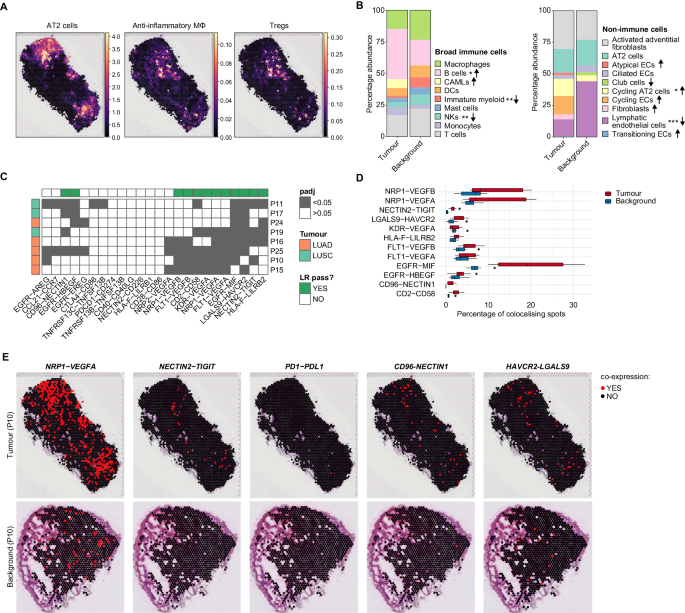
A Examine overview. Single-cell suspensions of resected tumour tissue, adjoining non-involved tissue (background) and wholesome lung from deceased donors have been enriched for CD45+ or CD235− and subjected to scRNA-seq. Cryosections of recent, flash-frozen tumour, background and wholesome tissues have been used for 10x Visium spatial transcriptomics. B Cohort overview. Symbols signify particular person sufferers and carried out analyses. C UMAP projection of tumour and mixed background+wholesome datasets. D Dotplot of consultant genes used for broad cell-type annotations in tumour samples. E Contour plot exhibiting the co-expression of myeloid (LYZ, CD68, MRC1) and epithelial (EPCAM) genes in AT2 cells (44,399 cells), CAMLs (2520 cells) and AIMɸ (16,120 cells). Normalised, scaled and log-transformed gene expression. F Boxplot exhibiting normalised, scaled and log-transformed gene expression of myeloid (LYZ, APOE, CD68, MRC1) and epithelial (EPCAM, KRT8, KRT19) genes in AT2 cells, CAMLs and AIMɸ. Containers: quartiles. Whiskers: 1.5× interquartile vary. G Relative proportion of non-immune cell subsets in tumour and background, calculated inside the CD235− enrichment. Arrows point out enhance (↑) or lower (↓) in tumour versus background. Pairwise comparisons by two-sided Wilcoxon rank check and Bonferroni correction for a number of comparisons. **P < 0.01. Arrows with out asterisks point out that the cell sort was discovered solely in tumour or background. H Relative proportion of broad immune cells in tumour and background, calculated inside all immune cells recognized within the CD235- enrichment. Arrows point out a rise (↑) or lower (↓) in tumour versus background. Pairwise comparisons by two-sided Wilcoxon rank check and Bonferroni correction for a number of comparisons. *P < 0.05, **P < 0.01, ***P < 0.001. Arrows with out asterisks point out that the cell sort was discovered solely in tumour or background. I Relative proportion of NK, DC, B, T and macrophage subsets inside the broad annotations in tumour and background, calculated inside the CD235- enrichment. Arrows point out enhance (↑) or lower (↓) in tumour versus background. Pairwise comparisons by two-sided Wilcoxon rank check and Bonferroni correction for a number of comparisons. ***P < 0.001. Arrows with out asterisks point out that the cell sort was discovered solely in tumour or background.
Tumours exhibit a better variety of immune and non-immune cells in comparison with adjoining lung tissue
Following high quality management (QC) on the scRNA-seq dataset, we recognized 895,806 high-quality cells in complete, of which 503,549 have been from tumour and 392,257 from mixed background and wholesome tissue (from right here on known as B/H). After performing normalisation and log1p transformation, highly-variable gene choice, dimensionality discount, batch correction, and Leiden clustering, cells originating from tumour and B/H have been individually annotated into distinct broad cell sorts and visualised by way of Uniform Manifold Approximation and Projection (UMAP) (Fig. 1C, Supplementary Fig. 1B, C, and “Strategies”). We recognized clusters of myeloid cells with transcriptional signatures of monocytes, macrophages, dendritic cells (DCs), in addition to mast cells, pure killer (NK) cells, T cells, B cells and non-immune cells (Fig. 1C, D). We didn’t detect neutrophilic granulocytes, most likely as a consequence of their sensitivity to degradation after assortment and specifically to the freezing-thawing cycle. Lastly, we recognized a cluster characterised by the co-expression of myeloid (LYZ, CD68, CD14, MRC1) and epithelial genes (KRT19, EPCAM) (Fig. 1D–F). These cells have been discovered inside the tumour and exhibited similarities to beforehand described cancer-associated macrophage-like cells (CAMLs)16,17,18. CAMLs signify a definite inhabitants of huge myeloid cells with concomitant epithelial tumour protein expression19. These distinctive cells have been noticed in blood samples of sufferers with varied malignancies, together with NSCLC20. The abundance of CAMLs displays a direct correlation with response to therapeutic interventions, highlighting their practical significance21. Even after additional subclustering, CAMLs maintained their distinct twin myeloid-epithelial signature (Supplementary Fig. 1D). It’s noteworthy that doublet detection software program Scrublet assigned a low doublet rating to CAMLs, suggesting their expression profile is unlikely to be defined as a mixed signature arising from the coincidental sequencing of a tumour cell and a macrophage (Supplementary Fig. 1E). All clusters included cells from a number of sufferers, with the cluster dimension starting from 2520 to 124,459 cells (Supplementary Fig. 1F, G). Moreover, we carried out reference-query mapping utilizing scArches22 to verify the consistency of our annotations within the tumour and B/H dataset (Supplementary Fig. 2A–C and Supplementary Notes).
The composition of the immune and non-immune compartment was markedly completely different between the tumour and background. Within the tumour, we detected fibroblasts and a lower within the fraction of lymphatic endothelial cells (LECs) (Padj = 0.0025, Fig. 1G and Supplementary Information 2). Moreover, the inhabitants of epithelial cells confirmed greater variety, with the presence of alveolar sort II (AT2), atypical epithelial cells which downregulated epithelial markers (KRT19, EPCAM, CDH1), transitioning epithelial cells which upregulated myeloid markers (LYZ), and biking epithelial cells in tumour tissues (Fig. 1G, Supplementary Notes, and Supplementary Fig. 2D, E). These variations are in settlement with the truth that in tumour specimens, epithelial cells are more likely to be a mix of mutant tumour and non-mutant regular cells, and recommend that neoplastic transformation results in additional variety of cell states. We didn’t detect alveolar sort I (AT1) or basal cells, probably as a consequence of their loss throughout dissociations, as beforehand reported by others8.
As beforehand reported, the proportion of monocytes and immature myeloid cells was considerably decreased in tumour samples in comparison with background (Padj = 0.022 and Padj = 0.00001, respectively)7, whereas DCs and B cells have been general expanded7 (Padj = 0.0023 and Padj = 0.0044, respectively; Fig. 1H and Supplementary Information 3). To get additional perception into the mobile composition of tumour versus background tissue, we subclustered every of the broad clusters and recognized 46 cell sorts/states (Supplementary Fig. 2D, E, Supplementary Information 4 and 5, Supplementary Fig. 3, and Supplementary Notes). Within the tumour, we discovered {that a} considerably greater proportion of NK cells had a decrease cytotoxicity phenotype (Supplementary Notes), and that the numerous majority of DCs have been derived from monocytes (i.e., mo-DC2), (Supplementary Notes) in comparison with background (Padj = 0.00002 and Padj = 0.00002, respectively, Fig. 1I and Supplementary Information 6). That is per the monocytic origin of mo-DC2s underneath inflammatory circumstances23. Equally, we discovered an growth of B cells expressing LYZ and TNF, and depletion of NKB cells (Fig. 1I and Supplementary Notes). Amongst T cells, tumour samples confirmed an accumulation of regulatory T cells (Tregs), recognized to hinder the immune surveillance of tumours24 (Fig. 1I). Conversely, there was a discount of exhausted cytotoxic T cells (Padj = 0.00002) within the tumour and absence of (gamma delta) T cells, which have been related to survival in NSCLC25 (Fig. 1I and Supplementary Information 6). (gamma delta) T cells are able to recognising and lysing various ranges of most cancers cells, and thus have been advised for a task in pan-cancer immunotherapy26. Lastly, we noticed a rise in heterogeneity and proportion of anti-inflammatory Mɸ (AIMɸ), with a subset of biking anti-inflammatory Mɸ, STAB1 + Mɸ (Fig. 1I) and CAMLs (Fig. 1H) being abundantly current in tumour tissue. Apparently, we discovered a powerful detrimental correlation between the frequency of STAB1 + Mɸ/AIMɸ and T/NK cells throughout sufferers, highlighting the important thing function of Mɸ in restraining the infiltration of cytotoxic cells within the lung tumour tissue (Fig. 2A). That is according to a latest work describing that monocyte-derived Mɸ in human NSCLC purchase an immunosuppressive phenotype and restrain the infiltration of NK cells27.
A Heatmap exhibiting the Pearson correlation between the relative cell-type abundance for every immune cell sort (calculated inside the CD235− enrichment). Color signifies the Pearson correlation worth, asterisks point out the extent of significance of the two-sided affiliation check computed on Pearson’s product-moment correlation coefficients (*P < 0.05, **P < 0.01, ***P < 0.001). B Heatmap exhibiting the variety of LR interactions between all cell sorts summarised by broad cell annotations in LUAD (left) and LUSC (proper). Rows have been hierarchically clustered utilizing the entire linkage methodology on euclidean distances. C Sankey diagram exhibiting the tumour-specific interactions in LUAD and LUSC for chosen ICIs detected by cellphoneDB. Line color identifies whether or not the LR interplay between every cell sort was present in LUAD solely (orange), in LUSC solely (inexperienced) or in each tumour sorts (blue). D Dotplot for the ICI genes and cell sorts highlighted in (C), cut up by tumour sort. The dimensions of every dot represents the proportion of cells within the cluster expressing the gene, whereas the color represents the imply normalised scaled log-transformed expression of every gene in every group. E Sankey diagram exhibiting the tumour-specific interactions in LUAD and LUSC for VEGFA/B interactors detected by cellphoneDB. Line color identifies whether or not the LR interplay between every cell sort was present in LUAD solely (orange), in LUSC solely (inexperienced) or in each tumours (blue). F Sankey diagram exhibiting the tumour-specific interactions in LUAD and LUSC for EGFR interactors detected by cellphoneDB. Line color identifies whether or not the LR interplay between every cell sort was present in LUAD solely (orange), in LUSC solely (inexperienced) or in each tumours (blue).
LUAD and LUSC have comparable mobile composition however utilise completely different cell–cell interplay networks
LUAD and LUSC have very completely different prognoses and are sometimes thought of as completely different scientific entities28. To look at if variations in scientific options stem from distinct mobile composition, we in contrast the frequency of immune and non-immune cell subsets inside CD235- samples from LUAD versus LUSC sufferers. We noticed minor variations in cell frequency that didn’t attain statistical significance after P worth correction (Supplementary Fig. 4A and Supplementary Information 7 and 8). Moreover, there was no clear affiliation between the frequency of immune and non-immune cells noticed in sufferers and the most cancers subtype, most cancers stage or intercourse (Supplementary Fig. 4B, C), suggesting that the TME composition is relatively comparable in LUAD and LUSC. Whereas LUAD and LUSC shared comparable mobile compositions, the noticed scientific distinctions might come up from various intercellular interactions. Subsequently, we examined whether or not completely different cell–cell interplay networks have been employed inside the TME in LUAD versus LUSC. To this finish, we recognized a putative record of cell–cell interactions completely noticed in every tumour sort surroundings by inferring statistically important ligand–receptor pairs (L–Rs) that weren’t detected in background or wholesome and their corresponding cell sorts, utilizing CellPhoneDB29. Though the 2 tumour subtypes confirmed an analogous interplay community that largely concerned interactions between non-immune cells, AIMɸ and T cells (Fig. 2B), there have been additionally some notable variations.
First, we recognized general a better variety of L–Rs within the LUAD dataset (Supplementary Fig. 4D and Supplementary Information 9–12), which was not pushed by a distinction within the variety of cells within the LUAD (n = 105,749 cells) vs LUSC (n = 230,066 cells) dataset. Secondly, a number of pairs of immune checkpoint inhibitors (ICI) and their respective inhibitory molecules have been differentially co-expressed in LUAD versus LUSC (Fig. 2C, D). For instance, LGALS9-HAVCR2 (TIM3), NECTIN2-CD226 (DNAM1) and NECTIN2/NECTIN3-TIGIT have been ceaselessly recognized in LUAD, and the putative ICI CD96-NECTIN1 was discovered preferentially in LUSC (Fig. 2C, D). In distinction, CD80/CD86-CTLA4 and HLAF-LILRB1/2 have been present in each tumour subtypes (Fig. 2C, D). LILRBs (leucocyte Ig-like receptors) are rising as potential targets for next-generation immunotherapeutics as their blocking can potentiate immune responses30. Probably the most generally used immunotherapies for lung most cancers block the interplay between PD1 and PDL1, and up to date scientific trials advised that anti-CTLA4 and anti-PD1 mixture remedy improved the survival of sufferers unbiased of tumour PD1 expression31,32. Inside our dataset, we didn’t observe PD1-PDL1 interactions in both of the tumour subtypes (Fig. 2C, D). Our preliminary evaluation means that different ICIs (akin to CTLA4, TIGIT, LILRB1/2 and TIM3) is likely to be promising targets within the therapy of NSCLC.
Of the numerous L–Rs detected in each LUAD and LUSC we famous a number of pairs concerned in angiogenic signalling in several populations of myeloid cells akin to VEGFA/B-FLT1, VEGFA-KDR and VEGFA-NRP1/2. Though VEGFA and VEGFB have been discovered to be expressed in each LUAD and LUSC, their receptors have been extra ceaselessly present in LUAD, particularly in fibroblasts (Fig. 2E and Supplementary Fig. 4E). Equally, we noticed important expression of EGFR ligands signalling in AT2 and biking epithelial cells, akin to EGFR-EREG, EGFR-AREG, EGFR-HBEGF and EGFR-MIF, though MIF expression was discovered extra ceaselessly in cells from LUSC (Fig. 2F and Supplementary Fig. 4F). Lastly, we noticed key co-stimulatory indicators required to assist lymphoid cell activation, akin to CD40-CD40LG, CD2-CD58, CD28-CD86, CCL21-CCR7, and TNFRSF13B/C-TNFSF13B (TACI/BAFFR-BAFF) (Supplementary Fig. 4G), which are sometimes related to the presence of ectopic lymphoid organs primarily consisting of B cells, T cells, and DCs i.e., tertiary lymphoid constructions (TLS). TLS are normally correlated with the longer relapse-free survival in NSCLC33.
Integration of scRNA-seq and spatial transcriptomics validates L–R interactions in situ
The numerous L–Rs and their interacting cell sorts have been calculated based mostly on the co-expression of genes in several cell-type clusters from the scRNA-seq dataset utilizing CellPhoneDB. Nonetheless, to be able to discern biologically important interactions, it’s important to determine whether or not the cell sorts recognized as interacting are certainly bodily co-located. To attain this, we thought of how the scRNA-seq-identified cell sorts are spatially organized on tissue sections. We utilized an integrative strategy which mixes the scRNA-seq of the tumour and background samples with the spatial transcriptomic (STx) profile of the recent frozen tumour and background tissue sections. We carried out 10× Visium on two consecutive, 10-µm sections, from eight sufferers, seven of which matched the samples used for the scRNA-seq. We analysed 36 sections in complete (ntumour = 20, nbackground = 16) with a median UMI depend of 6894/spot in tumour and 3350/spot within the background. Subsequent, we used cell2location34 and cell-type particular expression profiles from our scRNA-seq dataset to deconvolute cell-type abundances on the tissue (Fig. 3A, see “Strategies”).
A Spatial photographs depicting the cell abundance estimated by cell2location for AT2 cells, AIMɸ and Tregs on a consultant tumour part. B Relative proportion of immune (left) and non-immune (proper) cell sorts calculated on the cell abundance estimations by cell2location in tumour and background sections. Immune cells have been grouped in response to their broad annotations. Arrows point out a rise (↑) or a lower (↓) within the tumour, in comparison with the background. Pairwise comparisons have been carried out with a two-sided Wilcoxon rank check and Bonferroni correction for a number of comparisons. *P < 0.05, **P < 0.01, ***P < 0.001. Arrows with out asterisks point out that the cell sort was discovered solely within the tumour or background. Please consult with Supplementary Information 13 and 14 for the precise P values. C Heatmap of spatial LR colocalization. LR gene pair co-expression was estimated in every spot for all sections, and the frequency of colocalising vs. non-colocalising spots within the tumour and background was in contrast utilizing a χ2 check adopted by Bonferroni a number of comparability correction. Darkish-grey tiles point out that the frequency of colocalising gene pairs was considerably completely different in tumour and background sections. Inexperienced column annotations point out the LR pairs which have been important in no less than 4 out of eight sufferers. Row annotations point out tumour sort. D Boxplot exhibiting the frequency of colocalising LR pairs considerably completely different in tumour vs background in every part analysed. N = 8 sufferers. Containers are plotted with default settings within the Python Seaborn package deal, i.e., bins present quartiles with whisker size being 1.5 instances the interquartile vary. Supply information is supplied as a Supply Information file. E Spatial photographs depicting the placement of spots by which the LR pair was discovered co-expressed in tumour (high) and background (backside), for NRP1-VEGFA, NECTIN2-TIGIT, PD1-PDL1, CD96-NECTIN1 and HAVCR2-LGALS9. Consultant sections from one affected person.
As soon as the cell sorts have been resolved on the tissue sections, we examined the frequency of various cell sorts throughout all sections from tumour and background tissue. The cell-type abundance in tumour and background have been computed by summing up the posterior 5% quantile (q05) worth of estimated cell abundance by cell2location, throughout spots that handed QC (“Strategies”). Our evaluation confirmed that the variations within the frequency of cell sorts throughout all sections in tumour versus background was according to the outcomes obtained within the scRNA-seq information (Fig. 3B). For instance, in tumours we discovered a rise within the proportion of B cells (Padj = 0.0372) and biking AT2 cells (Padj = 0.0147) in comparison with the background tissue, and a lower within the proportion of immature cells (Padj = 0.0012), NK cells (Padj = 0.0012), and LECs (Padj = 0.00077, Supplementary Information 13 and 14). Nonetheless, the proportions of different cell sorts estimated from the scRNA-seq information or the STx information inside the tumour or background confirmed some discrepancies (Supplementary Fig. 4H, I). This was notably evident inside the non-immune populations, the place STx estimated greater proportions of LECs, activated adventitial fibroblasts and biking subsets, in comparison with scRNA-seq. Disparities in cell proportions between completely different methodologies have been beforehand proven by others35,36, underscoring the potential affect of distinct sampling biases inherent to scRNA-seq and STx methods like Visium. Within the case of scRNA-seq, variations in cell digestion sensitivity can result in differential illustration of cell sorts. In the meantime, with Visium, discrepancies would possibly come up from variations within the location of tumour resections in addition to variations in pattern sizes in comparison with scRNA-seq research. However, the general concordance within the outcomes obtained by scRNA-seq and Visium means that our spatial “map” of various cell sorts faithfully represents their distribution within the tissue.
Subsequent, we examined the spatial co-localisation of the L–Rs recognized by cellphoneDB. The L–Rs have been thought of to co-localise if each genes have been expressed in the identical spot and above median worth for the given genes throughout the part spots. We then in contrast the frequency of spots by which L–R genes have been colocalising versus non-colocalising within the matched tumour versus background sections, utilizing a χ2 check (“Strategies”). Because of the low variety of tissue blocks collected from LUSC and LUAD sufferers (NLUSC = 3, NLUAD = 5), the statistical energy was not enough to carry out a comparative evaluation between spatial localisation of LUAD/LUSC-specific L–Rs. However, we confirmed that a number of of the aforementioned tumour-specific L–Rs colocalized considerably extra in tumour than in background sections, together with NRP1-VEGFA and the ICIs NECTIN2-TIGIT, LGALS9-HAVCR2, and CD96-NECTIN1 (Fig. 3C–E and Supplementary Information 15). Per the cellphoneDB outcomes, we discovered no important colocalization of PD1-PDL1 within the tumour sections.
CAMLs share comparable copy quantity aberrations (CNAs) with tumour cells
Tumour samples obtained from surgical resection include each malignant and residual regular epithelial cells. A big problem in scRNA-seq of human tumours lies within the differentiation of most cancers cells from non-malignant counterparts. Subsequently, we utilized Copynumber Karyotyping of Tumors (CopyKAT37) to discern genome-wide aneuploidy inside particular person cells. The precept driving the computation of DNA copy quantity occasions from scRNA-seq information is rooted within the notion that the expression ranges of neighbouring genes can present helpful info to deduce genomic copy numbers inside that particular genomic phase. Since aneuploidy is frequent in human cancers, cells with genome-wide CNAs are thought of as tumour cells.
Evaluation utilizing CopyKAT revealed in depth, patient-specific CNAs in tumour tissue (Fig. 4A and Supplementary Fig. 5A) however not within the background. Inside particular person tumour samples, the CNAs have been detected in AT2 and biking AT2 cells, and in some sufferers these genetic alterations have been shared between AT2/biking AT2 cells and atypical epithelial cells, suggesting an in depth lineage relationship between completely different epithelial subpopulations (Fig. 4A and Supplementary Fig. 5A). We confirmed this discovering by inferring the trajectory of non-blood cell populations in tumour utilizing Partition-Primarily based Graph Abstraction (PAGA)38. PAGA confirmed differentiation continuity between AT2 cells, biking AT2/epithelial cells, and atypical epithelial cells on one aspect and ciliated epithelial cells and transitioning epithelial cells on the opposite (Fig. 4B). Moreover, blinded histological analysis confirmed the overlap between pathologist-defined tumour websites and AT2 and biking AT2 cells predicted by cell2location, suggesting their tumour cells standing (Fig. 4C). Much less overlap was noticed for atypical epithelial cells (Fig. 4C). The differential expression evaluation (DEA) of AT2 cells from tumours in comparison with background confirmed upregulation of genes concerned in hypoxia, TP53 pathways, and metabolic rewiring in tumours. AT2 cells in tumour-upregulated genes concerned each in glycolysis and oxidative phosphorylation (Fig. 4D and Supplementary Information 16). Whereas the significance of glycolysis in tumour cells is well-established39, it was not too long ago reported that human NSCLC use glucose and lactate to gasoline the tricarboxylic acid (TCA) cycle40. As well as, the tumour AT2 cells have been famous to specific extra LYPD3 in comparison with background AT2 cells (log2FC = 2.04, Padj = 0.039, Supplementary Information 16), an adhesion protein which has beforehand been linked to poor prognosis in NSCLC and is at the moment being focused in preclinical and scientific research41,42.
A CNA evaluation. The plot exhibits chromosomal beneficial properties (crimson strains) and losses (blue strains) estimated by CopyKat in every chromosome arm for various cell sorts and sufferers within the tumour dataset. All immune cell sorts have been grouped collectively for plotting functions. B PAGA graph overlaid on the diffusion maps (force-directed format—FLE embedding) computed for non-immune cell sorts in tumour. C First three panels—Consultant blind annotations from a professional pathologist, indicating the areas of tumour infiltration (left), binning of the tumour space on the Visium spots (centre) and the spots that handed QC (proper). The final three panels—cell2location estimation for AT2 cells (left), Biking AT2 cells (centre) and Atypical epithelial cells (proper) on the identical sections, overlaid with the pathologist’s annotation for the tumour infiltration (inexperienced contour). D Overrepresentation evaluation on gene ontology—organic processes (GO:BP) and REACTOME database by clusterProfiler R package deal, utilizing DEGs upregulated by AT2 cells in tumour vs background. Supply information is supplied as a Supply Information file. E Detailed overview of CNAs in AT2 and CAMLs from the tumour of 1 consultant affected person. Bars point out the frequency of cells harbouring chromosomal beneficial properties (crimson bar) or losses (blue bars) in particular chromosomal areas. F Scatterplot of the KL divergence for losses (x axis) and beneficial properties (y axis) between every cell sort within the tumour dataset calculated utilizing their acquire and loss distribution. All immune cell sorts have been grouped collectively for plotting functions. G Spatial photographs depicting the cell abundance estimated by cell2location for AT2 cells and CAMLs on three consultant tumour sections. H Hierarchical clustering of the correlation distance calculated on cell-type composition (as estimated by cell2location) throughout spots that handed QC in all tumour sections. I Non-negative matrix factorisation constructed on the q05 estimation of cell-type abundance throughout spots that handed QC (as estimated by cell2location) in all tumour sections.
Apparently, the inhabitants of CAMLs additionally confirmed substantial CNAs that have been much like these of AT2 cells and biking AT2 cells from the identical affected person (Fig. 4A, E and Supplementary Fig. 5A, B). To measure the distinction of the distribution of genomic acquire and loss between cell sorts in a statistically sturdy method, we calculated the Kullback–Leibler (KL) divergence (Fig. 4F and Supplementary Fig. 5C). CAMLs had KL divergence values corresponding to CNA-harbouring tumour cells, thus confirming the similarity of their CNA profiles (Fig. 4F and Supplementary Fig. 5C). As CAMLs co-expressed a wide selection of myeloid genes in addition to typical epithelial genes (Fig. 1D–F and Supplementary Fig. 1D), had a low doublet rating and shared the identical CNA signature as tumour cells, we hypothesised that these cells would possibly signify a subset of Mɸ tightly connected to a most cancers cell. It’s doable that these Mɸ have been present process phagocytosis or fusion.
CAMLs have been beforehand remoted from peripheral blood of most cancers sufferers and described to facilitate circulating tumour cells seeding of distant metastases16. Our evaluation advised that CAMLs may also be remoted from tumour tissue. To validate that CAMLs are in bodily proximity to tumour cells in situ we examined our STx sections. We calculated throughout all sections (8 sufferers, nsections = 20) the Pearson correlation between the relative abundance of the cell sorts that reside in the identical spot and are subsequently co-localised. Our evaluation confirmed that CAMLs certainly co-localised with AT2 cells (Fig. 4G, H). We confirmed this discovering utilizing non-negative matrix factorisation (NMF) on absolutely the cell-type abundances estimated by cell2location that outlined components of co-occurring cell states (Fig. 4I).
To find out the precise Mɸ inhabitants from which CAMLs seemingly originate, we employed PAGA to elucidate the differentiation path of the myeloid cell inhabitants in our tumour dataset (Supplementary Fig. 5D). The evaluation revealed continuity of the differentiation transitions between various populations of myeloid cells43. Inside the PAGA trajectory, alveolar Mɸ (AMɸ) and AIMɸ confirmed excessive PAGA connectivity indicating their excessive transcriptional similarity. Each AIMɸ and AMɸ confirmed the strongest connectivity on the PAGA trajectory with STAB1 + Mɸ which, in flip, have been linked with CAMLs. Consistent with trajectory evaluation, CAMLs co-expressed most of the genes particular to STAB1 + Mɸ (Supplementary Fig. 2A), supporting the speculation that CAMLs are seemingly derived from STAB1 + Mɸ following their shut interplay with tumour cells. Lastly, DEA evaluation between CAMLs from LUSC versus LUAD sufferers, confirmed upregulation of KRT17, KRT5 and KRT6A in LUSC samples (Supplementary Information 17). These KRT genes have been beforehand recognized as markers of LUSC in a number of research44,45, which helps speculation that CAMLs come up from the interplay between Mɸ and tumour cell.
TAMs promote ldl cholesterol and iron efflux in tumour
Mɸ, historically categorised into distinct M1 (classically activated) and M2 (alternatively activated) phenotypes, at the moment are understood to exist alongside a dynamic spectrum of practical states46. This idea of Mɸ plasticity underscores their skill to seamlessly transition between pro-inflammatory and anti inflammatory roles in response to intricate cues from their microenvironment (Supplementary Fig. 5D). To raised perceive the transcriptional modifications that completely different Mɸ populations endure within the TME, we carried out DEA. In tumours, each AMɸ and AIMɸ upregulated genes concerned in ldl cholesterol and lipid transport and metabolism (akin to ABCA1, APOC1, APOE, FABP3 and FABP5) in comparison with the background tissue (Fig. 5A, B and Supplementary Information 18 and 19). Ldl cholesterol performs an important function in tumour progress as a result of excessive demand of newly synthesised mobile membranes throughout most cancers cell proliferation. Hypoxia-related genes have been upregulated in AT2 cells in tumour in comparison with the background (Fig. 4D), which may promote ldl cholesterol auxotrophy in tumour cells by suppressing ldl cholesterol synthesis, thereby forcing them to depend on exogenous ldl cholesterol uptake47. In our dataset, we detected greater expression of the ldl cholesterol exporter ABCA1 and no expression of low-density lipoprotein receptor (LDLR) in AMɸ and AIMɸ, the latter gene being liable for the uptake of cholesterol-carrying lipoprotein particles into cells, suggesting preferential export of ldl cholesterol from TAMs to the TME (Fig. 5A). Apparently, we additionally famous a excessive expression of TREM2 in each AMɸ and AIMɸ (Fig. 5A), which performs a outstanding function in efflux of ldl cholesterol in microglia48,49,50. To validate the elevated ranges of ldl cholesterol within the TME, we stained matched tumour and background tissue sections with BODIPY™ 493/503, a stain focusing on ldl cholesterol and different impartial lipids. We discovered a big enhance within the BODIPY sign within the tumour sections, in comparison with background tissue (Fig. 5C, D), confirming an elevated availability of impartial lipids within the tumour, probably on account of an elevated export by TAMs.
A Volcano plot of DEGs (crimson) for AIMɸ in tumour vs background, extracted utilizing the py_DESeq2 package deal. B Overrepresentation evaluation on gene ontology—organic processes database by clusterProfiler R package deal, utilizing the DEGs upregulated by Alveolar Mɸ and AIMɸ in tumour vs background. Supply information is supplied as a Supply Information file. C IHC for CD68 and impartial lipids (BODIPY 493/503) on tumour and background tissue sections. Most depth projection of Z-stacks. Scale bar 50 µm. D Space lined by the BODIPY sign in tumour and background part. The distinction in BODIPY space protection was decided with a paired, two-sided t check, matching tumour and background sections from the identical sufferers. N = 5 sufferers. Supply information is supplied as a Supply Information file. E IHC for CD68 and STAB1 on tumour (left) and background (proper) tissue sections. Most depth projection of Z-stacks. Inlets present an in depth magnification on a single cell. Scale bar 20 µm. F Quantification of STAB1+ cells inside the CD68+ macrophage inhabitants. The fraction of the STAB1 + CD68+ space is proven as a proportion of the full CD68+ space. Information are introduced as imply worth and commonplace deviation (n = 3 organic replicates). Supply information is supplied as a Supply Information file. G Staining for CD68, STAB1 and PanCK on tumour tissue sections. Most depth projection of Z-stacks. Inlets present an in depth magnification on a single cell. Scale bar 20 µm. H Quantification of STAB1 + CD68+ cells inside the CD68+ macrophage inhabitants in NSCLC. Information are introduced as imply worth and particular person information factors (n = 2 organic replicates). Supply information is supplied as a Supply Information file. I Dotplot exhibiting the expression of the “STAB1 signature genes” throughout all macrophage subsets and CAMLs in tumour. J Volcano plot of DEGs recognized by py_DESeq2 (crimson) for Alveolar Mɸ vs STAB1 Mɸ in tumour. Ok Overrepresentation evaluation on gene ontology— organic processes database by clusterProfiler R package deal, utilizing the DEGs from Alveolar Mɸ vs STAB1 Mɸ (high) and AIMɸ vs STAB1 Mɸ (backside) in tumour (left—upregulated by STAB1 Mɸ; proper—upregulated by Alveolar Mɸ or AIMɸ). Supply information is supplied as a Supply Information file.
STAB1 + Mɸ have been recognized within the tumour resections (Fig. 5E–H, Supplementary Fig. 2 and Supplementary Notes), so we used DEA to establish a set of genes that have been particular for STAB1 + Mɸ in comparison with tumour AIMɸ or AMɸ. We recognized 20 genes, from right here on known as “STAB1 signature genes” (Fig. 5I). Apparently, STAB1 + Mɸ uniquely expressed SLC40A1, which encodes for the ferroportin, the one recognized protein that exports ferrous iron from the cytoplasm throughout the plasma membrane and is vital for the iron-releasing exercise of macrophages (Fig. 5I, J and Supplementary Information 20 and 21)51. Ferroportin-mediated launch of free iron by M2 Mɸ was reported to advertise the proliferation of renal carcinoma cells in vitro, probably by supporting the excessive iron requirement as a consequence of elevated DNA synthesis52. Moreover, in comparison with AMɸ, STAB1 + Mɸ expressed decrease ranges of ferritin heavy chain 1 (FTH1) and ferritin gentle chain (FTL) encoding for the iron storer ferritin (Fig. 5J and Supplementary Information 20). Per the speculation of their sustained export of free iron to the extracellular milieu, STAB1 + Mɸ downregulated genes concerned in iron sequestration (Fig. 5K). Taken collectively, our evaluation means that macrophages endure “reprogramming” inside the TME and undertake a transcriptional signature that facilitates ldl cholesterol efflux and iron export, thus supporting tumour development.
STAB1 + Mɸ in tumour tissue endure oncofoetal reprogramming
Embryonic growth shares many traits with tumour tissue, together with speedy cell division, mobile flexibility, and a extremely vascular microenvironment. It has been not too long ago reported that in tumorigenesis, Mɸ can endure oncofoetal reprogramming53 and purchase a foetal-like transcriptional identification that helps tumour progress and metastasis53. Contemplating that a number of the STAB1 signature genes are usually expressed by foetal Mɸ (akin to STAB1, FOLR2, SLC40A1, MERTK, GPR34 and F13A1)54, we wished to discover if additional transcriptional commonalities exist between tumour-originating STAB1 + Mɸ and Mɸ remoted from human foetal lung. To this finish, we mixed tumour- and background-originating myeloid cells from our dataset (n = 347,364 cells) with myeloid and progenitor cells from a publicly accessible foetal lung scRNA-seq dataset55 (n = 6,947 cells) utilizing Concord. Subsequent, we carried out Leiden clustering on the neighbourhood graph and examined how cell sorts are distributed inside the clusters (Supplementary Fig. 6A, B). To look at similarity of their gene expression profile, we utilized hierarchical clustering and constructed a dendrogram by estimating the correlation distance between cell sorts on the harmonised PC embedding house, underneath the entire linkage criterion of hierarchical clustering (Fig. 6A).
A Hierarchical clustering of the correlation distance calculated on every cell within the harmonised (tumour myeloid + background myeloid + foetal lung myeloid) PC house. B Violin plot exhibiting the expression stage of the “STAB1 gene signature” throughout myeloid cell and progenitor populations recognized in a publicly accessible human foetal lung atlas. C Dotplot of the expression of every gene within the “STAB1 gene signature” in chosen foetal lung macrophage populations. The dimensions of every dot represents the proportion of cells within the cluster expressing the gene, whereas the color represents the imply expression of every gene in every cluster. D Violin plot exhibiting the expression stage of the “STAB1 gene signature” throughout the clusters recognized within the publicly accessible MoMac-VERSE dataset. E Dotplot of the expression of every gene within the “STAB1 gene signature” in chosen macrophage populations from the MoMac-VERSE. The dimensions of every dot represents the proportion of cells within the cluster expressing the gene, whereas the color represents the imply expression of every gene in every cluster. F Violin plot exhibiting the expression stage of the “AMɸ gene signature” throughout myeloid cell and progenitor populations recognized within the publicly accessible “MoMac-VERSE” dataset. G Violin plot exhibiting the expression stage of the “AMɸ gene signature” throughout myeloid cell and progenitor populations recognized in a publicly accessible human foetal lung atlas. H Dotplot of the expression of every gene within the “AMɸ gene signature” in chosen macrophages populations recognized within the “MoMac-VERSE” dataset. The dimensions of every dot represents the proportion of cells within the cluster expressing the gene, whereas the color represents the imply expression of every gene in every cluster. I Dotplot of the expression of every gene within the “AMɸ gene signature” in chosen foetal lung macrophage populations. The dimensions of every dot represents the proportion of cells within the cluster expressing the gene, whereas the color represents the imply expression of every gene in every cluster.
We noticed that tumour cDC2 exhibited the strongest correlation with background cDC2, whereas tumour mo-DC2 displayed the very best correlation with foetal DC2 and, in a broader context, with background mo-DC2. The inhabitants of pDC from tumour, background and foetal lung have been carefully correlated. Equally, tumour monocytes have been correlated with foetal classical monocytes and background monocytes. In distinction, macrophage populations in tumour, and specifically STAB1 + Mɸ, have been correlated with foetal macrophages. STAB1 + Mɸ clustered predominantly with foetal SPP1 + Mɸ (Fig. 6A), which accounted for over 80% of all foetal lung macrophages reported in ref. 55. Per this discovering, SPP1 + Mɸ had a excessive expression of the “STAB1 signature genes” in comparison with different haematopoietic populations (Fig. 6B, C). Our evaluation substantiates the concept monocytes inside the tumour surroundings, as they endure differentiation into anti-inflammatory macrophages, purchase a transcriptional signature akin to that of foetal macrophages. This distinctive transcriptional signature was not noticed within the macrophages from surrounding regular tissue.
To additional study the prevalence of STAB1 + Mɸ in different pathologies, together with different cancers, we examined the expression of “STAB1 signature genes” throughout a various group of myeloid cells utilizing a broadcast atlas of human monocytes and Mɸ collected from 12 completely different wholesome and pathologic tissues (n = 140,327 cells), referred to as MoMac-VERSE56. The cluster of “HES1+ macrophages” recognized in MoMac-VERSE confirmed the very best expression of the “STAB1 signature genes” (Fig. 6D, E). Much like STAB1 + Mɸ, HES1+ macrophages amassed in tumours of lung most cancers sufferers but additionally liver most cancers sufferers57 and have been advised to signify a cluster of “long-term resident-like” Mɸ with foetal-like transcriptional signature56. In distinction, “C1Q” Mɸ from MoMac-VERSE, which have been described as lung alveolar Mɸ, had a excessive expression of genes distinctive to our tumour alveolar AMɸ (from right here on referred as “AMɸ signature genes”, Fig. 6F, H). Within the context of foetal lung, a uncommon inhabitants of APOE + Mɸ, which accounted for lower than 1% of all foetal lung macrophages reported in ref. 55, had a excessive AMɸ signature genes rating (Supplementary Notes and Fig. 6G, I, see “Strategies”).
Taken collectively, our evaluation means that tumour macrophages, particularly STAB1 + Mɸ, exhibited a transcriptional signature harking back to Mɸ throughout foetal lung growth, suggesting that they’ve undergone oncofoetal reprogramming inside the NSCLC tumour surroundings.