Identification of differential genes associated to bone metastasis in PCa
Determine 1 illustrates the primary circulate of this research. First, we obtained 120 BMRGs by differential expression evaluation as the premise for downstream evaluation. In the meantime, 10 diagnostic markers for PCa metastatic occasions have been screened utilizing PPI community and associated plugins. Cox regression evaluation was used to seek out prognostic genes to assemble the mannequin, which was then used to group sufferers. Purposeful enrichment, immune and mutational profiles, and immunotherapy efficacy have been investigated in several teams to determine the power of the mannequin to discriminate between sufferers and to information scientific apply.
Movement chart of this analysis. DEGs, differentially expressed genes. CTCs, circulating tumor cells. BMRGs, bone metastasis-related genes. PPI, Protein–protein interplay. BMGPI, bone metastasis-related genes prognostic index. KM, Kaplan-Meier survival evaluation. ROC, Receiver Working Attribute Curve. GSEA, gene set enrichment evaluation. GSVA, gene set variation evaluation.
A complete of 1994 DEGs arose between CTCs and first samples within the GSE67980 cohort (Desk S1). Within the GSE32269 cohort, we recognized 624 DEGs between bone metastatic and first samples (Desk S2). Among the many 1994 DEGs, 259 have been up-regulated and 1,735 have been down-regulated in CTCs. The up-regulated genes have been primarily enriched in RNA splicing and protein folding and stabilization (Fig. 2A). Whereas down-regulated genes have been related to cell adhesion and angiogenesis, and have been concerned in PI3K-Akt signaling pathway and Ras signaling pathway (Fig. 2B). In PCa bone metastasis samples, there have been 249 up-regulated and 345 down-regulated genes. The up-regulated genes have been concerned in cell division and cell cycle (Fig. 2C), and the down-regulated genes have been enriched in angiogenesis, growing old, and metabolic pathways (Fig. 2D). The intersection of DEGs from two cohorts was taken to acquire 120 core BMRGs associated to bone metastasis.
Identification of the hub module associated to metastasis by means of the PPI community
We constructed a PPI community primarily based on 120 BMRGs and filtered the highest ten genes with a level higher than 10 utilizing the MCC algorithm of the cytoHubba plugin, together with ACTA2, ACTG2, CNN1, COL1A1, LMOD1, MYH11, MYL9, MYLK, TAGLN and TPM2 (Fig. 3A). Moreover, to make the outcomes extra goal, we additional screened three modules with MCODE plugin, and essentially the most outstanding module, which had a rating of 11.294, 18 nodes, and 96 edges, is proven in Fig. 3B. This module nonetheless consists of the ten genes talked about above. Subsequently, the grasp regulators of ten intersecting genes have been additionally explored by the iRegulon plugin and we discovered that 9 of them have been regulated by SRF (Fig. 3C).
Screening hub genes by Cytoscape software. (A) Cytohubba and (B) MCODE plugins have been utilized to display for hub genes. (C) The gene regulation sample was predicted by iRegulon plugin. Correlation analyses have been carried out for the ten hub genes in (D) GSE32269, (E) GSE6919, (F) and TCGA cohorts, respectively. Expression of those ten genes in (G) GSE32269 and (H) GSE6919 datasets. (I) AUC values demonstrated the predictive efficacy of hub genes for metastasis.
The relevance of those ten hub genes was additional analyzed. Aside from COL1A1, the expression ranges of the opposite 9 genes exhibited a robust constructive correlation (Fig. 3D–F), which was in line with the expected outcomes demonstrated in Fig. 3C. To validate the diagnostic efficacy of those hub genes for tumor metastasis, we recognized their expression patterns and calculated AUC values in each GSE32269 and GSE6919 datasets. These 9 genes have been much less expressed in distant metastatic samples in comparison with major samples, whereas COL1A1 confirmed an reverse development (Fig. 3G,H). As illustrated in Fig. 3I, the AUC values indicated superior diagnostic efficiency of the ten hub genes.
Growth and validation of the BMGPI mannequin primarily based on integrative computational framework
Firstly, univariate Cox evaluation recognized 20 prognostic genes from 120 BMRGs (p < 0.001). These 20 genes have been subjected to 94 mixtures of machine learning-based algorithms so as to assemble a bone metastasis-related genes prognostic index (BMGPI). Afterwards, we additional computed the C-index for the coaching and validation cohorts in every mixed mannequin (Fig. 4A). Curiously, Lasso + RSF and RSF fashions have been equally optimum among the many 94 fashions, each of which led by way of common C-index (0.775). Moreover, each fashions additionally demonstrated good predictive functionality in validation cohorts. However, the Lasso + RSF mannequin contained 11 genes, whereas the RSF mannequin contained solely 6 genes. Due to this fact, given the practicality and translational potential, we thought of the RSF mannequin because the optimum mannequin as the premise for subsequent evaluation.
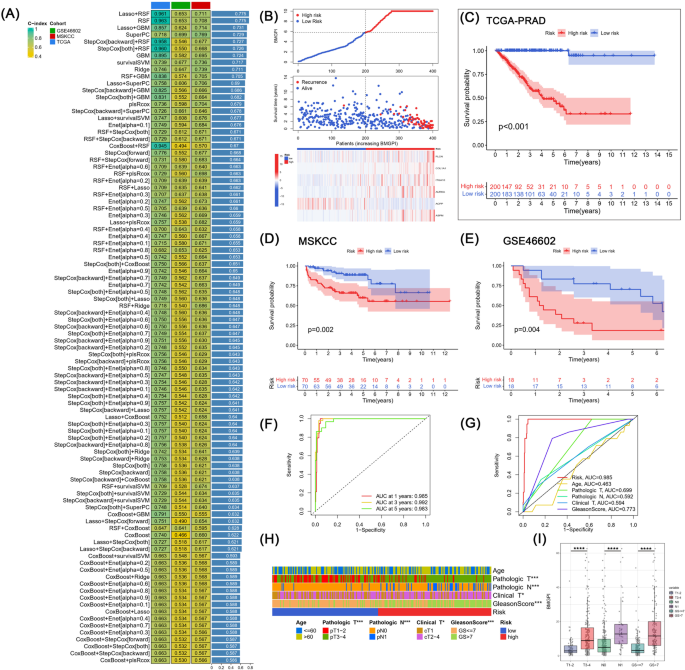
Building and validation of BMGPI by means of the machine learning-based integrative program. (A) Ninety-four predictive fashions have been utilized to the coaching cohort and the validation cohort, and the C-index was calculated for every mannequin throughout all cohorts. (B) The distribution of survival time, survival standing, and the six genes comprising the mannequin within the high- and low-risk teams. KM survival curves for danger teams in (C) TCGA, (D) MSKCC, (E) and GSE46602 cohorts. (F) Time-dependent and (G) scientific characteristic-related ROC curves. (H) Distribution of scientific traits in danger teams. (I) The distinction in BMGPI between sufferers grouped by T stage, N stage, and Gleason rating.
Primarily based on the random survival forest algorithm, every affected person was assigned a danger rating (BMGPI). Sufferers have been stratified into high- and low-risk teams primarily based on median scores. With rising BMGPI, the survival of the sufferers turned progressively worse (Fig. 4B). As well as, all 5 mannequin genes besides ACPP have been extremely expressed within the high-risk group. The survival curve within the coaching group instructed that the high-risk group was related to a poor prognosis, which was validated in each the MSKCC and GSE46602 cohorts (Fig. 4C–E). The AUC values for 1-, 3-, and 5-year survival for the coaching group have been 0.985, 0.992, and 0.983, respectively, as proven in Fig. 4F. Furthermore, in Fig. 4G,danger had superior predictive efficacy in comparison with the Gleason rating, T stage, and N-stage. Subsequently, we additional assessed the distribution of assorted scientific traits in danger teams (Fig. 4H). The proportions of T-stage, N-stage and M-stage have been considerably totally different between the 2 danger teams. BMGPI was additionally discovered to be greater in sufferers with superior phases (Fig. 4I).
BMGPI is an impartial predictor for survival of PCa sufferers
Univariate and multivariate Cox analyses have been utilized to judge the affiliation of BMGPI and different scientific traits with prognosis of sufferers. BMGPI was recognized as an impartial prognostic issue for PRAD sufferers in each the TCGA and MSKCC cohorts (Fig. 5A). To increase the worth of the mannequin for scientific software, on the premise of danger stage, pathologic T stage and Gleason rating, we constructed a nomogram to foretell 1-, 3- and 5-year survival (Fig. 5B). As demonstrated in Fig. 5C, the expected survival of the nomogram was properly in line with the noticed survival. The AUC and C-index additionally indicated that the constructed nomogram had extra correct and sturdy predictive potential in contrast with different variables (Fig. 5D,E).
Building of the nomogram counting on danger stage and scientific parameters. (A) Univariate and multivariate Cox regression evaluation for DFS in TCGA and MSKCC cohorts. (B) The nomogram primarily based on T stage, Gleason rating, and danger stage. (C) Calibration plot for 1-, 3-, and 5-year survival. The comparability of (D) the AUC values and (E) C-index between the nomogram and different scientific traits.
Purposeful enrichment in danger teams
In accordance with GSEA, the pathways related to the high-risk group primarily included cell cycle, base excision restore, DNA replication, P53 signaling pathway and homologous recombination, whereas a number of amino acid metabolic pathways have been associated to the low BMGPI (Fig. 6A). Then, we utilized GSVA to additional take a look at the affiliation of BMGPI with mobile pathways. The development of the correlation between the BMGPI and enrichment scores remained in line with the GSEA outcomes (Fig. 6B). With a view to decide whether or not these pathways affect the prognosis of sufferers, we carried out a survival evaluation primarily based on the enrichment scores (Fig. 6C). Poor prognosis was correlated with pathways positively correlated with BMGPI, reminiscent of cell cycle, base excision restore, spliceosome, DNA replication, homologous recombination, mismatch restore and NOD like receptor signaling pathway. Whereas pathways positively correlated with BMGPI was correlated with good prognosis, reminiscent of beta alanine metabolism, propanoate metabolism and valine leucine and isoleucine degradation.
Perform enrichment evaluation. (A) The outcomes of GSEA in danger teams. (B) Correlation evaluation between BMGPI and immune enrichment scores. The colours of the heatmap signify the immune rating calculated by GSVA for every affected person. (C) Survival evaluation revealed that GSVA scores for sure purposeful pathways have been considerably related to survival of sufferers.
Immune and genomic variation panorama in danger teams
BMGPI was positively related to immune scores (r = 0.2399, p < 0.0001) (Fig. 7A). Moreover, the high-risk group had greater immune exercise in comparison with the low-risk group (Fig. 7B). The bubble plot demonstrated the correlation coefficients between BMGPI and totally different immune cell infiltration (Fig. 7C). The panorama of immune cells primarily based on totally different strategies in danger teams was introduced within the warmth map (Fig. 7D). The outcomes of ssGSEA revealed that the TME of the high-risk group contained extra dendritic cells (DCs), plasmacytoid dendritic cells (pDCs), macrophages, tumor infiltrating lymphocytes (TIL) and T helper cells, whereas the TME of the low-risk group had the next content material of mast cells (Fig. 7E). We additionally discovered that the high-risk group additionally confirmed extra energetic in some immune capabilities, reminiscent of cytolytic exercise and T cell co-stimulation (Fig. 7F). Correlation evaluation of BMGPI with immune cells and immune operate was visualized in Fig. 7G. Moreover, quite a lot of immune checkpoints have been enriched within the high-risk group (Fig. 7H).
Immune panorama related to BMGPI. (A) Correlation evaluation between BMGPI and immune scores. (B) ESTIMATE rating, immune rating, and stromal rating for danger teams. (C) The bubble plot confirmed the correlation coefficients between varied sorts of immune cells and BMGPI. (D) The heatmap of immune cell abundance primarily based on totally different software program platforms. SsGSEA of (E) immune cells and (F) immune operate. (G) The heatmap illustrated the correlation between the BMGPI and immune cell or operate. (H) Extra immune checkpoints have been activated within the high-risk group.
We additionally depicted the mutation profiles of the high- and low-risk teams (Fig. 8A,B). P53, a widely known oncogene, was mutated considerably extra continuously within the high-risk group (18%) than within the low-risk group (5%). Moreover, as a key regulator of the androgen receptor (AR), mutations in FOXA1 might trigger alterations within the exercise of transcription components, selling epithelial-mesenchymal transition (EMT) and most cancers metastasis26. Its mutation frequency was 7% within the high-risk group, whereas it was solely 4% within the low-risk group. The distribution of base mutation sorts was exhibited in Fig. 8C,D. BMGPI is positively correlated with tumor mutation burden (TMB) (r = 0.2739, p < 0.0001) (Fig. 8E). Furthermore, we explored the copy quantity variation (CNV) of the differential genes between the 2 danger teams (|logFC| > 1, p. adj < 0.05). Aside from ALOX15B, CHRNA2 and PEBP4, the primary alteration in different genes was CNV achieve (Fig. 8F). Lastly, we analyzed the phenomenon of co-occurrence mutation within the high- and low-risk teams (Fig. 8G,H), which appeared extra continuously within the high-risk group.
Genetic alterations in danger teams. Waterfall plots displaying the clusters of genes with the best frequency of somatic mutations in (A) high- and (B) low-risk teams. Abstract of mutation patterns in (C) high- and (D) low-risk teams. (E) Scatter plot displaying a constructive correlation between the BMGPI and TMB. (F) CNV of DEGs between high- and low-risk teams. Heatmaps demonstrating the collinearity of mutations within the prime 25 mutated genes of (G) high- and (H) low-risk teams.
Prediction of response to immunotherapy remedy
A TIDE rating was calculated for every affected person within the TCGA cohort to foretell response to immunotherapy. TIDE scores elevated with rising BMGPI, indicating that the high-risk group might not reply properly to immunotherapy (Fig. 9A). The chi-squared take a look at instructed that the high-risk group responded to immunotherapy at a decrease charge than the low-risk group (Fig. 9B,C). We additional noticed elevated BMGPI within the subgroup of sufferers who didn’t reply to immunotherapy (p = 0.002) (Fig. 9D). Curiously, within the IMvigor210 cohort, we famous that BMGPI was greater within the CR group than within the PR group (p = 0.026), whereas there was no vital distinction in BMGPI between the responding and non-responding teams (Fig. 9E). To additional assess the efficacy of PD1 inhibitor and CTLA4 inhibitor in subgroups, we resorted to the IPS rating from the TCIA database. In each CTLA4 (+) / PD1 (−) and CTLA4 (−) / PD1 (−) therapies, IPS scores have been greater within the low-risk group, implying higher efficacy (Fig. 9F,G). Lastly, we screened 64 medication with considerably totally different IC50s within the excessive and low danger teams, 10 of which have been visualized in Fig. 9H, reminiscent of Bicalutamide and Paclitaxel.
Immunotherapy sensitivity. (A) Scatter plot displaying a constructive correlation between the BMGPI and TIDE rating. (B) The distribution of TIDE rating in NR and R teams. (C) A graph plot depicting share of sufferers receiving immunotherapy who responded or didn’t reply in danger teams. (D) The distinction in BMGPI between NR and R teams. (E) A field plot presenting the BMGPI of sufferers with CR, PR, SD, and PD within the IMvigor210 cohort. The IPS of the (F) CTLA4+/PD1- or (G) CTLA4-/PD1- remedy in danger teams. (H) Drug sensitivity evaluation.