Figuring out transcriptomic adjustments all through the adenoma-carcinoma sequence
To analyze the ACA transcriptomic profiles and uncover the gene expression adjustments by the colorectal adenoma-carcinoma sequence, we carried out and processed RNA-seq knowledge obtained from 40 ACA affected person samples and picked up RNA-seq knowledge for CRC and regular adjoining colon from a public database (SRA ID: SRP029880; CRC: 18; regular adjoining colon: 18). All ACA donors had polyps with diameters higher than 1 cm, and their histological sorts and the situation from which the samples have been obtained have been recorded. Supplementary Desk S1 supplies ACA sample-specific info, whereas Supplementary Fig. S1 depicts colonoscopy photos for every ACA subtype.
With the intention to cut back confounding components, RNA-seq counts have been normalized and batch results between knowledge sources have been adjusted (Supplementary Fig. S2). Earlier than conducting the DEA, potential outliers have been screened with exploratory knowledge evaluation utilizing multidimensional scale (MDS) plotting and principal element evaluation (PCA). In each MDS and PCA plots, just one ACA pattern (ACA_18) confirmed the extraordinary expression sample, which was positioned within the reverse area with out every other samples close by. It was thought-about as an outlier and excluded from the next evaluation to acquire sturdy outcomes (Supplementary Fig. S2).
We carried out DEA to determine genes with steady adjustments in expression worth throughout the adenoma-carcinoma sequence (Supplementary Fig. S3 and Supplementary Information 1–3). Primarily based on the fold-change (FC) between consecutive phases (Regular-ACA and ACA-CRC), there have been 21 up-regulated (log2FCACA-Regular > 1, log2FCCRC-ACA > 1, and false discovery price (FDR) < 0.05) and 79 down-regulated genes (log2FCACA-Regular < − 1, log2FCCRC-ACA < − 1, and FDR < 0.05), which have been recognized as steady differentially expressed genes (DEGs). The complete abstract statistics from DEA for steady DEGs are offered in Supplementary Tables S2 and S3. By making use of the gene-wise scaling, we noticed the gradual adjustments in gene expression ranges on the continuum from regular tissue to CRC (Fig. 2a,b).
DEA for steady DEG identification and their organic capabilities. (a) A heatmap displaying up-regulated steady DEG expression patterns throughout adenoma-carcinoma sequences. The annotation bar signifies pattern kind (darkish inexperienced: regular, inexperienced: ACA, and orange: CRC). The colour of every cell is proportionate to its column-centered relative expression worth. (b) A heatmap of down-regulated steady DEG expression ranges. (c) A bar plot displaying the highest 5 enrichments of up-regulated steady DEGs utilizing GO-BP pathways. The bar colour signifies the mixed rating from enrichR, and the x-axis exhibits the OR of every pathway. (d) A bar plot of the enrichment take a look at outcomes for up-regulated steady DEGs utilizing KEGG pathways. (e) A bar plot displaying the enrichment of down-regulated steady DEGs on GO-BP pathways. (f) A bar plot of the enrichment of down-regulated steady DEGs on KEGG pathways. The pathway enrichment evaluation outcomes are offered as Supplementary Information 4.
To determine the organic mechanisms of steady DEGs, we carried out enrichment checks for gene ontology organic processes (GO-BP) and Kyoto Encyclopedia of Genes and Genomes (KEGG) pathways14,15. Up-regulated steady DEGs have been considerably enriched (P < 0.05) in pathways associated to carcinogenesis, irritation, and immune responses (Fig. 2c,d). Enrichment profiles of down-regulated steady DEGs primarily implied the lack of regular colonic capabilities throughout carcinogenesis (Fig. 2e,f). Reflecting the carcinogenic position of reactive nitrogen species, associated pathways confirmed the 2 highest scores and odds ratios (OR) within the enrichment evaluation utilizing up-regulated steady DEGs in GO-BP classes, whereas down-regulated steady DEGs confirmed the best enrichment of nitrogen metabolism within the KEGG pathway (Fig. 2f). Vital enrichments of immune or inflammatory pathways corresponding to B-cell differentiation, rheumatoid arthritis, and interleukin (IL)-17 pathways in Fig. 2c,d could counsel the significance of immunological landscapes within the adenoma-carcinoma sequence.
Hub-genes for the adenoma-carcinoma sequence might also be necessary for affected person survival
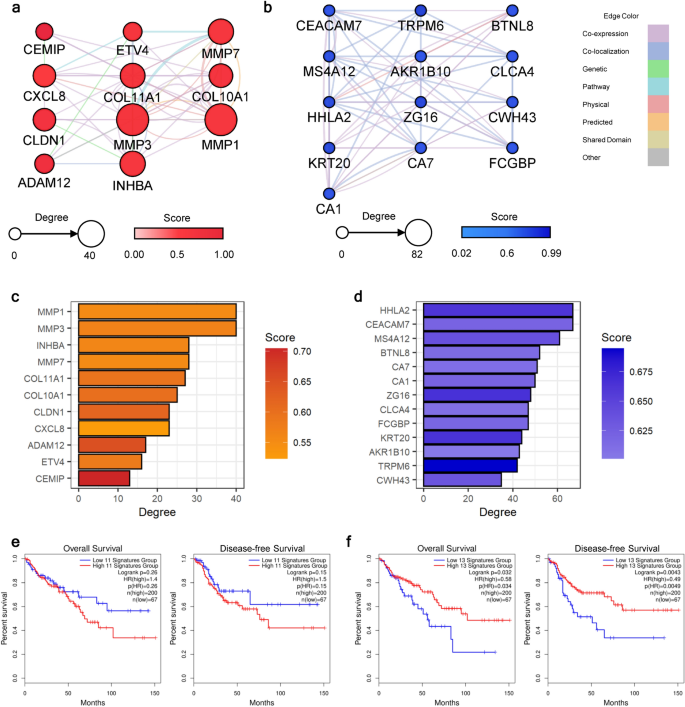
To determine the purposeful or regulatory interconnection inside the up- or down-regulated steady DEGs, we constructed the protein–protein interplay (PPI) networks for the up- and down-regulated steady DEGs individually. Interactions between every gene–gene pair have been searched for his or her co-expression, co-localization, genetic relationship, involvement in the identical pathway, bodily interplay, computationally predicted interplay, shared area, and others (Supplementary Fig. S4). After developing the preliminary community, we computed the community statistics to determine the core-functioning or core-interacting genes amongst every community. We narrowed down the hub-genes by making use of the cut-off of > 50% percentile for 2 standards: diploma and rating from the GeneMania plug-in. Amongst 41 genes within the community of up-regulated steady DEGs, 11 genes handed the cut-off worth (Fig. 3a,c). Implying the transition into the malignant stage, genes associated to cell migration, extracellular matrix (ECM) degradation, or rearrangement, corresponding to matrix metalloproteinase (MMP), collagen (COL) household genes, ADAM metallopeptidase area 12 (ADAM12), and cell migration-inducing hyaluronidase 1 (CEMIP), have been recognized because the community hub-genes. Moreover, CXC motif chemokine ligand 8 (CXCL8) was included within the standards, which is understood for its vital position in tumor microenvironment (TME) alteration16. In a down-regulated community, 13 genes have been recognized as hub-genes (Fig. 3b,d). Genes associated to the carcinogenesis of the gastrointestinal tract, corresponding to carcinoembryonic antigen cell adhesion molecule 7 (CEACAM7), membrane-spanning 4-domains A12 (MS4A12), carbonic anhydrase (CA) 1 and 7, chloride channel accent 4 (CLCA4), Fc gamma binding protein (FCGBP), and aldo–keto reductase household 1 member B10 (AKR1B10), have been concerned within the record of hub-genes. Different genes recognized have been associated to organic capabilities in immunological regulation (HERV-H LTR-associating 2: HHLA2, and zymogen granule protein 16: ZG16) and mobile construction formation (keratin 20: KRT20, and cell wall biogenesis 43 C-terminal homolog). Conforming with the earlier research, this outcomes collectively implied that the overactivated ECM reworking and malfunctioning immune cells are the essential element within the development towards the CRC stage17,18.
Identification of the carcinogenesis-stimulating hub-genes. PPI networks have been constructed with (a) up- and (b) down-regulated hub-genes. The node sizes are proportional to their levels. Darker node colours point out increased scores calculated from the GeneMania plug-in, which delineates the purposeful significance of the gene. The sting colours depict interplay sorts between nodes. Bar plots depicting the community statistics of recognized hub-genes for (c) up- and (d) down-regulated networks. The bar lengths point out the diploma of every gene, and the darker colour signifies the next rating from the GeneMania plug-in. The survival evaluation outcomes utilizing the (e) up- and (f) down-regulated hub-genes. Each the general and disease-free survival charges have been analyzed, and their outcomes are introduced on the left and proper sides of every panel, respectively. For every enter gene set, survival curves for sufferers with excessive or low expression ranges are marked with pink or blue strains, respectively.
With the intention to validate the population-specificity and generalizability of the hub-genes, we in contrast the route of gene expression adjustments towards exterior datasets. We collected one consultant dataset for every inhabitants: Northeast Asian (NEA), European (EUR), and blended (multi-study meta-analysis) (Desk 1)5,6,19. By calculating the Pearson’s correlation coefficients (PCC) of the log2FC values between our outcomes and exterior research, we noticed constructive correlations aside from the CRC-ACA distinction in EUR dataset (Supplementary Desk S4) and our outcomes confirmed excessive similarity with the DEA leads to the NEA dataset (0.522 < PCC < 0.924). Since there have been solely few overlapping genes with multi-study meta-analysis (6, 6, and a pair of genes for ACA-normal, CRC-normal, and CRC-ACA, respectively), we discovered that all the instructions of differential expression have been equivalent. These outcomes could suggest the underlying population-specific mechanisms in ACA to CRC development; nonetheless, the distinction in gene expression measurement platform in EUR dataset (microarray) in itself could have brought on the technical variation.
Subsequently, we validated the significance of the hub-genes by inspecting the affiliation between gene expression ranges and affected person survival. Within the survival evaluation, we examined the general and disease-free survival charges utilizing colorectal adenocarcinoma (COAD) knowledge from The Most cancers Genome Atlas (TCGA). As proven in Fig. 3e, sufferers with excessive expression ranges of up-regulated hub-genes confirmed a decrease survival price with nominal statistical significance (P total survival = 0.26 and P disease-free survival = 0.15). Survival charges considerably dropped (P < 0.05) in each total and disease-free survival when the gene expression ranges of down-regulated hub-genes have been within the decrease 25% vary (Fig. 3f). Collectively, we discovered that the hub-genes from our analyses could crucially have an effect on all phases of illness prognosis, from illness development to relapse to the survival price of sufferers.
Steady adjustments have been estimated within the panorama of innate immunity cells
Contemplating the rising position of immune cell composition in TME growth, we carried out single-cell deconvolution evaluation to estimate the immune cell kind fraction throughout the adenoma-carcinoma sequence. Inspecting the immune cell sorts concerned in innate immunity, there have been three cell sorts that confirmed a sure development from a standard state to CRC.
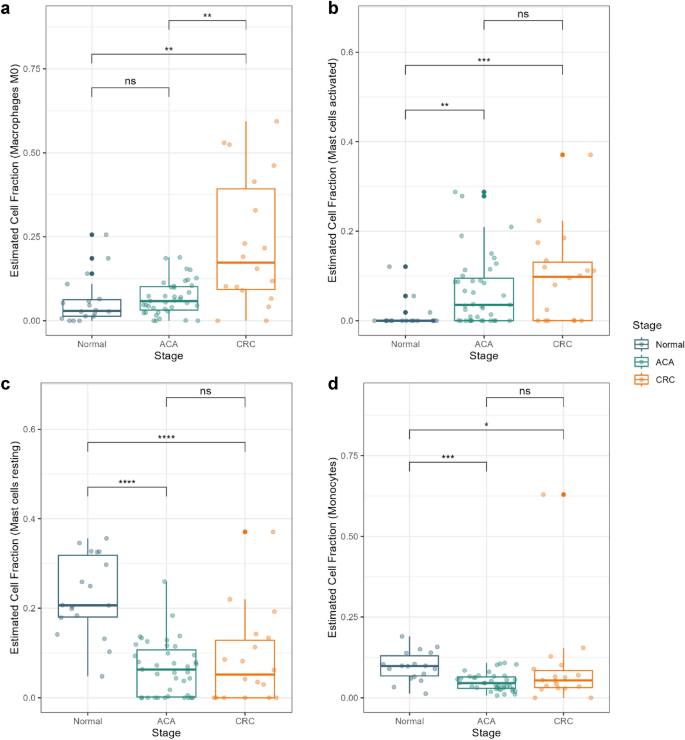
In keeping with earlier findings suggesting the recruitment of M0 macrophages in TME, our outcomes confirmed a considerably excessive cell fraction of M0 macrophages in CRC samples (Fig. 4a)20. Recruited macrophages appear to be actively polarized into each M1 and M2 states in CRC, which distinguishes it from ACA and regular tissues (Supplementary Fig. S5). Macrophages are identified for his or her position in TME regulation by secreting cytokines that promote tumor cell proliferation. Our outcomes display that this vital transition of macrophage proportion doesn’t seem on the pre-malignant stage, however a speedy shift can happen when the foremost proportion of the tissue turns malignant.
Cell kind fraction estimation for innate immunity cells. Field plots presenting the estimated cell fractions calculated with CIBERSORT X software program. Cell fractions for (a) macrophage M0, (b) activated mast cells, (c) resting mast cells, and (d) monocytes are introduced. Regular, ACA, and CRC pattern sorts are marked in blue, inexperienced, and yellow colours, respectively. **** P < 0.0001, *** P < 0.001, ** P < 0.01, * P < 0.05, and ns > 0.05.
Mast cells have been one other type of innate immune cell that demonstrated a big distinction. Primarily based on a earlier research by Yu et al., cross-talk between tumor-resident mast cells and surrounding most cancers cells promotes tumor development by releasing protumorigenic indicators21. Our outcomes counsel that in ACA and CRC tissues, tumor-resident mast cells have been extra more likely to be in an energetic state, whereas mast cells in regular tissue have been largely in a resting state (Fig. 4b,c). Moreover, a big depletion of the monocyte inhabitants was estimated in ACA and CRC samples (Fig. 4d). Whereas earlier research had primarily mentioned the adverse correlation between monocyte abundance and CRC prognosis, we recommend that this modification may begin from premalignant state of CRC22.
We then constructed the multinomial logistic regression fashions for classifying the tissue stage utilizing the estimated cell fractions for macrophages and monocytes (see Strategies). By inspecting the accuracy of the constructed fashions, we discovered that these predictors can effectively classify the tissue phases as much as 88% of accuracy (Supplementary Determine S6). We additionally examined the regression fashions with single cell kind. Whereas macrophage confirmed its potential to be a helpful predictor (70% < accuracy), monocyte alone confirmed the poor property because the prognostic marker (accuracy = 56%). This outcomes could suggest that the cell fraction of macrophage can function the outstanding prognostic/diagnostic marker for CRC continuum.
Composition of adaptive immune cells and immune repertoires are related to CRC growth
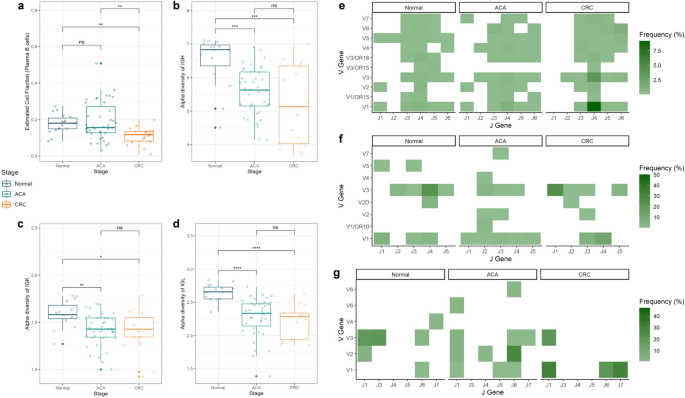
Tumor-infiltrating adaptive immune cells, together with B- and T-cells, are concerned in TME formation and immune evasion mechanisms23. Single-cell deconvolution evaluation confirmed a big depletion of the plasma B-cell inhabitants, which is a B-cell subtype that largely secretes Igs (Fig. 5a). Moreover, we discovered elevated cell abundance in reminiscence B-cells in ACA samples and a nominally lowering development in naive B-cells in ACA and CRC samples (Supplementary Fig. S7).
Cell fraction and immune repertoire evaluation for adaptive immunity elements throughout the CRC continuum. (a) Field plots presenting the plasma B-cell fraction throughout the CRC continuum. The graph colours point out the pattern sorts (regular: blue; ACA: inexperienced; CRC: yellow). The field plots of the immune repertoire range evaluation are: (b) IGH, (c) IGK, and (d) IGL chains. The heatmaps delineate the VJ recombination frequencies of (e) IGH, (f) IGK, and (g) IGL chains throughout the adenoma-carcinoma sequences. The colour of every cell is proportional to the relative frequency of VJ recombination in sure pattern sorts.
To research the heterogeneity of Ig sequences among the many samples, we extracted the unmapped reads matching Ig sequences utilizing the ImReP software program. On this process, 4 out of 40 samples (SRR975551, SRR975573, SRR975574, and SRR975577) exhibited ambiguous Ig sequences, which will be thought-about poor-quality reads and have been excluded from subsequent evaluation. We calculated the alpha range index, which describes sequence range inside the samples, to guage the heterogeneity of Ig repertoires. Whereas the lower in plasma cell inhabitants doesn’t develop into apparent in CRC samples, we discovered that the Ig repertoire began to lower from the premalignant stage of CRC. This development was noticed for complementarity-determining area 3 (CDR3) sequences of all three varieties of Ig chains: IgH, IgK, and IgL (Fig. 5b–d).
Together with the numerous lower in Ig range, we might determine the everyday recombination patterns of the V and J segments of Igs (Fig. 5e–g). Particularly, the mixture of V3-J2 and V6-J6 segments in IgH, V2-J3 segments in IgK, and V1-J1 segments within the IgL chain have been solely noticed in samples from ACA and CRC phases, suggesting a possible affiliation between the VJ recombination patterns and CRC development. Analyzing the range of Ig repertoires between the samples with beta range calculation, we discovered that the IgH chains between samples from regular phases have been comparatively related (Sørensen-Cube index: 0.409; Supplementary Fig. S8). We additionally recognized a big lower within the CD8 T-cell inhabitants; nonetheless, we have been unable to detect conforming patterns for TCR range (Supplementary Figs. S9–S11) as a result of we have been unable to seize enough sequences comparable to the T-cell receptor (TCR) area in regular or CRC samples.
Estimating tumor-associated HLA allele typing and expression evaluation
Whereas Igs or TCRs typically counteract the non-self antigens beforehand encountered by the host, HLA genes are additionally concerned in self-antigen recognition, whose malfunctioning is carefully associated to the immune evasion of most cancers cells24. Thus, we extracted the reads aligned to the HLA alleles with seq2HLA software program25. To evaluate the mutation occurring by carcinogenesis, we profiled the HLA sorts throughout the sufferers. We discovered the adjustments between regular adjoining and CRC tissues for HLA-A (02:01) and HLA-C (07:02) amongst class I alleles, and DPA1 (01:03, 02:01, 02:03, and 03:02), DPB1 (02:01, 04:01, 04:02, 05:01, and 104:01), DQA1 (01:01 and 01:02), DQB1 (03:01, 03:03, 04:01, 05:01, 05:03, and 06:02), and DRB1 (13:02) on class II alleles (Supplementary Fig. S12). Contemplating that the traditional adjoining tissue and CRC tissue have been collected from paired sufferers, these adjustments can suggest the somatic mutation of the HLA allele through the carcinogenesis.
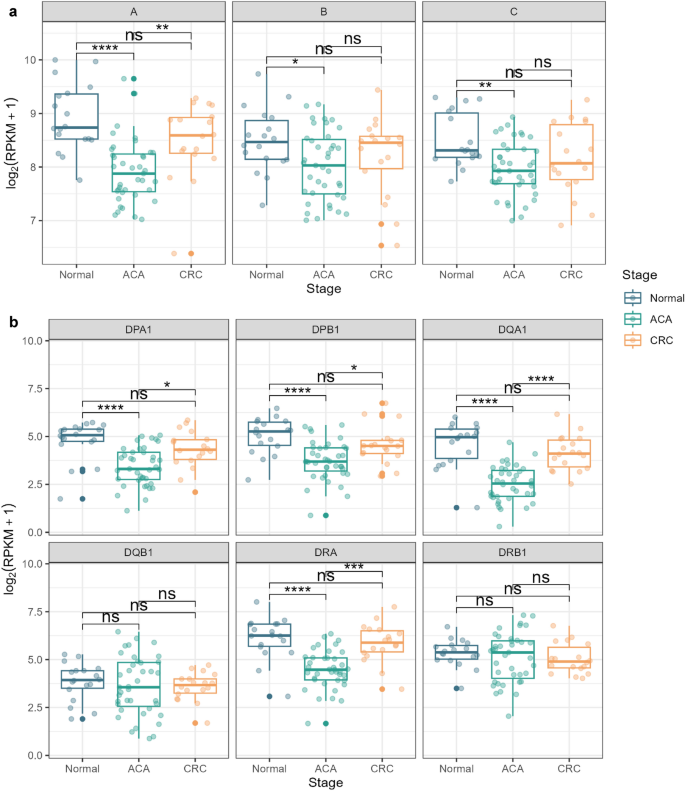
Subsequently, we analyzed the expression of HLA genes throughout samples utilizing a non-parametric Kruskal–Wallis take a look at, adopted by Dunn’s take a look at as a post-hoc take a look at. We detected considerably low HLA class I gene expression ranges in ACA samples (Fig. 6a). This development was constant in all three HLA class I alleles: HLA-A, HLA-B, and HLA-C. Though the distinction between regular and CRC tissue was not statistically vital, we be aware that there’s a particular development towards decreased CRC expression ranges in contrast with regular tissue, with Z-scores of − 1.30 (P = 0.19), − 1.06 (P = 0.28), and − 1.75 (P = 0.07) for HLA-A, B, and C alleles, respectively. Considerably decreased expressions of HLA genes in ACA samples have been equally noticed for HLA class II genes, aside from HLA-DQB1 and DRB1 alleles (Fig. 6b). Together with HLA class I genes, the median expression degree of HLA class II genes was highest in regular tissue. Primarily based on these outcomes, we suspect that immune evasion mechanisms mediated by HLA gene expression could start on the ACA stage and be related to somatic mutations at HLA loci.
HLA gene expression evaluation throughout the CRC continuum. (a) Field plots presenting the expression ranges of kind I HLA genes (HLA-A, HLA-B, and HLA-C) throughout adenoma-carcinoma sequences. (b) Field plots of the expression degree of kind II HLA genes (HLA-DPA1, HLA-DPB1, HLA-DQA1, HLA-DQB1, HLA-DRA, and HLA-DRB1). The graph colour signifies the pattern stage (regular: blue; ACA: inexperienced; CRC: yellow).