We explored a number of RNA-seq most cancers datasets of bulk (population-based) and single cells which can be out there within the Nationwide Middle for Biotechnology Data (NCBI)’s Gene Expression Omnibus (GEO) database20. We chosen seven most cancers sorts for bulk (breast21, colorectal22, leukemia23, liver24, ovarian25, pores and skin26, and osteosarcoma27) and three sorts for single cells (breast15, ovarian28, and glioblastoma29) which can be appropriate for our analyses (see “Strategies” for particulars). For proteomics meeting information, we looked for information on Proteomic Knowledge Commons (PDC)30 and recognized appropriate datasets for liver and ovarian cancers.
Bulk transcriptome and proteome noise and correlation analyses
Initially, we targeted on bulk datasets. After performing high quality management checks and decrease expressions filtering (“Strategies”), we investigated the extent of worldwide gene expression correlation (and, consequently, the relative quantity of defined and stochastic variability). We in contrast transcriptome- and proteome-wide scatterplots of regular, most cancers, and regular versus most cancers pattern pairs, and evaluated their corresponding Pearson (linear steady), Spearman (monotonic rank-based) correlations, mutual info (MI, nonlinear dependence), and noise (sq. of the coefficient of variation31) (Desk 1, Fig. 1a and Supplementary Fig. 1, grey dots). Basically, as anticipated, the transcriptome-wide variability and noise are decrease (and thus, correlation is greater), between regular samples when in comparison with between most cancers samples or between most cancers and regular samples (Fig. 1a, left panel). Whereas an analogous sample may be noticed within the proteomic information, the distinction between most cancers and regular samples within the expression of most cancers genes is extra pronounced (Fig. 1a, proper panel).
Varied varieties of statistical evaluation had been carried out on bulk datasets to indicate the expression invariance of most cancers genes. The evaluation for transcriptomics samples is on the precise panels, and the evaluation for the proteomics samples is on the left panels. a Scatterplots between regular samples, tumor samples, and regular vs tumor samples for liver most cancers and ovarian most cancers, with the remaining sorts offered in Supplementary Fig. 1. Common genes are represented by grey dots, CGC genes by blue dots, and CSO genes by pink dots. b Pearson correlation for the expression ranges of CGC genes (blue), CSO genes (pink), CGC-sized sampled random genes (purple), CSO-sized sampled random genes (orange). Ovarian most cancers was chosen for instance for each proteome and transcriptome right here, and the remainder of the most cancers sorts are in Supplementary Fig. 2. c PCA plots for entire dataset regular samples (mild blue circles), entire dataset tumor samples (mild orange circles), CGC genes regular samples (darkish blue circles), CGC genes tumor samples (darkish orange circles), CGC-sized random sampling of genes from regular samples (darkish blue stars), CGC-sized random sampling of genes from tumor samples (darkish orange stars), and the remainder of the most cancers sorts are in Supplementary Fig. 4.
Subsequent, we targeted on cancer-associated or most cancers genes (~600 CGC and ~20 cancer-specific oncogenes (CSO), “Strategies”) and in contrast their noise and scatter (Fig. 1a and Supplementary Fig. 1, blue and pink dots, respectively). Notably, for breast, colorectal, liver, ovarian, and pores and skin most cancers transcriptomes, we discover that each CGC and CSO genes have decrease scatter and noise (Desk 2, Fig. 1a (proper panel), Supplementary Fig. 1a, c, e) in comparison with their entire transcriptome, particularly between regular and most cancers. That is opposite to expectations since these genes are typically mutated in cancers and their expressions are anticipated to be considerably altered when in comparison with regular32. Then again, on the proteome stage, CGC and CSO genes present barely greater noise and variability (Desk 2 and Fig. 1a, left panel). To keep away from any statistical biases induced by dimension variation between the entire and subset of most cancers genes, we additionally sampled CGC and CSO dimension random genes/proteins with 100 occasions repeated sampling (Fig. 1b, Supplementary Fig. 2a–g, and Desk 2). For each transcriptome and proteome, the Pearson correlation and noise analyses present greater correlations and decrease noise between regular samples, and the other development between cancers and between regular and most cancers samples for all sampling sizes. Apparently, the correlations between random and most cancers genes (CGC and CSO) in regular and most cancers circumstances, is comparable, and in sure instances such because the liver transcriptome and the ovarian transcriptome, the CSO present better correlations than random samplings. These information recommend that the expression variability and correlations of most cancers genes are typically invariant with respect of the entire genome or randomly chosen genes.
To examine mutual and nonlinear dependence between the samples, we investigated mutual info (MI, “Strategies”) based mostly nonlinear correlations for each most cancers and random genes (Supplementary Fig. 3). Once more, the outcomes are inconclusive in that we couldn’t generalize throughout the completely different most cancers sorts whether or not random or most cancers genes show a special diploma of affiliation. Whereas these outcomes can’t be adjusted for the tumor purity of most cancers samples since not all of the datasets offered this info, the purity of the tumor samples within the pores and skin26 and liver24 most cancers transcriptomic datasets had been established to be enough (>30% tumor cell fraction, and 0.821–0.905 purity index respectively). Notably, the invariance of most cancers genes from random genes is even stronger for liver most cancers samples.
Thus, by finding out scatterplots, noise, linear, and nonlinear correlations, we couldn’t conclude whether or not CGC or CSO, collectively, show completely different statistical properties with respect to equally sized random samples of genes in each transcriptomic and proteomic information. To probe this outcome additional, we subsequent carried out dimensional discount utilizing Principal Part Evaluation (PCA) (“Strategies”).
Dimensionality discount by PCA
Determine 1c and Supplementary Fig. 4 present the PCA resolution within the first two dimensions accounting for the biggest variance quantity. For each transcriptome and proteome, we noticed that ordinary samples (mild blue circles) are situated nearer to at least one one other, whereas their most cancers counterparts (mild orange circles) are extra dispersed throughout the x–y house (Fig. 1c and Supplementary Fig. 4). This outcome signifies that the entire organic datasets (Fig. 1c, transcriptome-left panel, proteome-right panel) of regular replicates is much less noisy than that for cancers, confirming the outcomes of the bivariate correlation analyses above. Nonetheless, after we analyzed the identical metrics for most cancers genes (CGC and CSO), they’re much less variable and nearer to the metrics between regular (darkish blue circles) and most cancers (darkish orange circles). To check whether or not that is as a result of dimension or gene quantity results, as above, we examined random gene samples of the identical dimension within the regular tissue (darkish blue stars) and the tumors (darkish orange stars). Notably, the places of the randomly chosen samples are nearly invariant to these of the most cancers genes, in each omics’ ranges. Moreover, the imply Euclidian distance between the samples’ places in entire datasets between regular and most cancers is the best in comparison with that of most cancers genes or random genes alone (Desk 3).
Taken collectively, these outcomes point out that though regular and most cancers counterparts have vastly completely different quantities of heterogeneity on the entire transcriptome and proteome scale, they’re, nevertheless, very comparable when solely most cancers and randomly sampled genes are considered.
Pattern similarity evaluation
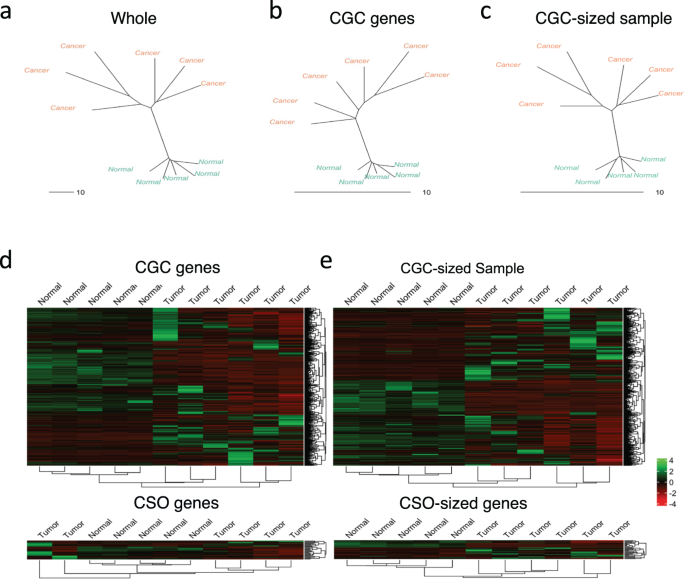
To additional examine the noticed invariance between CGC and random genes, we employed two varieties of pattern similarity analyses on the transcriptomic datasets: Neighbor-Becoming a member of (NJ) and hierarchical clustering. We first used the entire transcriptomes to mission samples from every most cancers sort into NJ dendrograms (Fig. 2a and Supplementary Fig. 5, left panels). For breast, glioma, ovarian, and osteosarcoma, the dendrograms present clear separations and clusters differentiating most cancers (orange), and regular samples (blue). For the remaining leukemia, liver, and pores and skin, the dendrograms, nevertheless, appeared randomly clustered for a subset of most cancers and regular samples. This might not be stunning as some most cancers information are extremely heterogenous even between their replicates33.
NJ pattern tree of ovarian most cancers (remainder of most cancers sorts proven in Supplementary Fig. 5) used to spotlight the variations between tumor samples (orange) and regular samples (blue). The bushes had been generated from a entire transcriptome, b CGC genes, c CGC-sized random sampling of genes. Hierarchical clustering was used to indicate the variations between tumor samples and regular samples of ovarian most cancers utilizing d CGC genes and CSO genes, and e CGC-sized random sampling of genes and CSO-sized random sampling of genes.
Investigating the precise impact of CGC genes on NJ dendrograms, we noticed that the general distance between all samples decreased considerably, in an analogous method to our dimension discount evaluation (Fig. 2b and Supplementary Fig. 5, center panels). Nonetheless, it’s value noting the truth that for some cancers small native modifications occurred the place tumor samples had been rearranged. To keep away from the statistical bias induced by pattern dimension variation, we as soon as once more carried out the pattern similarity evaluation on a CGC-sized random sampling of genes. As anticipated, the gap between samples decreased for the random sampling of genes as properly (Fig. 2c and Supplementary Fig. 5, proper panels), with the general imply sum of department lengths between the samples being comparable. Not like the CGC genes, nevertheless, the dendrograms generated from the random sampling of genes had been extremely much like the entire transcriptome dendrograms (Fig. 2a, c and Supplementary Fig. 5, proper and left panels). Unexpectedly, the dendrograms generated utilizing the CGC genes had the tumor samples mildly rearranged. Whereas these outcomes additionally present invariance between CGC and randomly sampled genes with regards to differentiating between tumor and regular samples, CGC genes do seem to play some function in amongst tumor pattern variations.
Subsequent, we carried out hierarchical clustering for CGC, CSO and randomly sampled genes (Fig. second, e and Supplementary Fig. 6). Notably, solely in some cancers CGC and random genes had been capable of accurately cluster most cancers and regular samples (ovarian and osteosarcoma), whereas in the remainder the clustering was not exact. However, the general clustering based mostly on CGC and random genes had been extremely comparable for all cancers with minor rearrangements solely. Within the case of CSO, the separation between regular and tumor samples decreased when in comparison with the CGC clustering in all most cancers sorts. Moreover, after we in contrast this end result with the outcomes of the CSO-sized sampling of random genes, we noticed that the random genes carried out poorly in separating tumor samples and regular samples as a result of small variety of genes.
General, these outcomes level to the same clustering of samples between CGC and CGC-sized random samplings of genes, in addition to CSO and CSO-sized random samplings of genes, suggesting invariance as noticed for the earlier sections. To date, the analyses are unable to spotlight any important impact of CGC genes in comparison with the remainder of transcriptome.
PPI community evaluation
For the reason that definition of oncogenes solely permits for protein-coding genes, their interplay properties may be investigated to make clear whether or not oncogenes are really like another protein-coding genes within the transcriptome. Notably, organic PPI networks are inclined to show energy legislation distribution34 with few nodes (hubs) having 1000’s of interactions (edges), whereas most nodes (leaves) may have a couple of or only a single interplay.
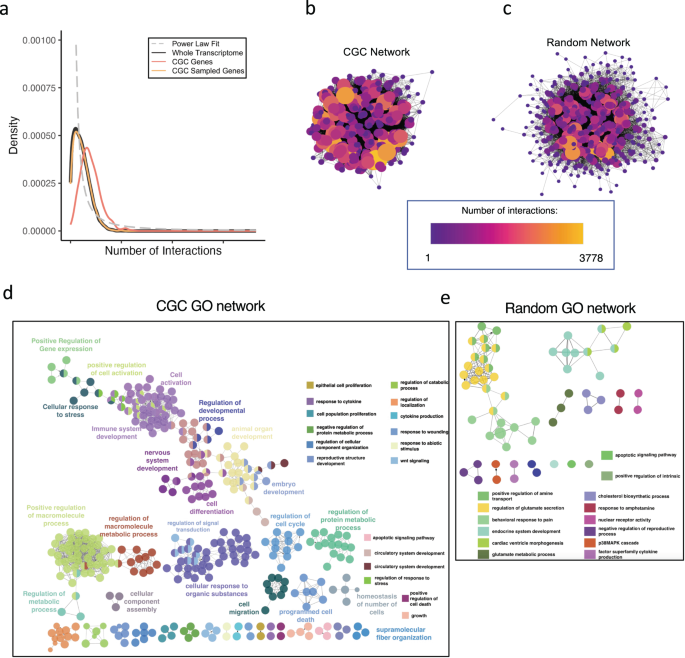
We plotted the distribution density of the variety of interactions per gene for all recognized protein-coding genes in STRING and GeneMania databases mixed35,36 (black), and located it to comply with the final development of the ability legislation distribution (dotted grey line), the place solely a strict minority of genes have a number of thousand of interactions adopted by a speedy lower in connectivity for the opposite genes (Fig. 3a). Subsequent, we investigated the PPI distributions of CGC and CGC-sized random genes. First, for CGC-sized random genes (yellow), it may be noticed that the distribution of interactions appears to be like much like that of the entire, aside from the height density, which is barely decrease. Second, for the distribution of PPI per CGC gene (orange), we observe that its imply is greater, and the density plot is shifted farther from the ability legislation curve (grey). When computing the common variety of interactions per gene, we noticed that the entire transcriptome and the random sampling have a virtually equivalent common, whereas the CGC genes have a considerably greater imply variety of interactions per gene (Desk 4). This outcome isn’t a surprise since most cancers genes are far more extensively studied with respect to all different genes, so their variety of connections (stemming from literature information) is prone to be greater. Moreover, some most cancers genes are recognized to be transcription components (TFs) as properly, which might clarify their connectivity. However, solely 20% of CGC genes (Supplementary Fig. 7) are discovered to be TFs37, and upon eradicating these 20%, the common variety of PPI doesn’t drop considerably (Desk 4). Thus, the outcomes spotlight CGC oncogenes as a particular subset of genes with above-average connectivity and far more homogeneous (with respect to the entire gene set) when it comes to their physiological roles.
a Utilizing the literature-known connectivity properties of protein-coding genes we generated a density plot for the variety of PPI per gene for all human protein-coding genes (black), CGC genes (pink), CGC-sized random sampling of genes (yellow) and the fitted energy legislation distribution (grey dashed line). We then used Cytoscape to visualise PPI interplay networks for b CGC genes and c CGC-sized random sampling of genes. We additionally explored the GO networks (“Strategies”) generated from d CGC genes and e a set of randomly sampled genes of the identical dimension because the CGC set.
To additional illustrate this level, we generated PPI community plots for CGC and random genes (Fig. 3b, c). A colour and dimension gradient had been used to spotlight the variety of interactions per node, the place bigger connectivity reveals greater dimension and lighter colour. When in comparison with the randomly sampled genes, the CGC community possesses extra extremely related nodes, much like community hubs, and fewer nodes with single connections. Moreover, the CGC community appeared to have a considerably greater variety of edges connecting the nodes, indicative of CGC genes being genes associated to one another. The random community, however, regardless of containing genes with a comparatively excessive variety of international connections, seemed to be significantly much less regionally related.
Lastly, to be able to examine the organic significance of CGC genes, we generated GO networks utilizing ClueGO38 in Cytoscape (Fig. 3d). We noticed that the GO community of CGC genes is dense and composed of varied essential organic processes that may have an effect on cell destiny. Then again, after we generated GO networks from the identical variety of random genes as CGC genes (Fig. 3e), we noticed that the community is sparser with fewer organic phrases which can be capable of cross the minimal filtering threshold. Lastly, in contrast to within the CGC GO community, the place every node was composed of tens to tons of of genes, within the random GO community the utmost variety of genes per node is 6 genes. This reveals that CGC genes cowl a variety of interconnected organic processes which have the flexibility to have an effect on cell destiny.
Collectively, these outcomes emphasize the distinctiveness or significance of oncogenes because the extra extremely related “hub genes” and recommend that the insignificant conduct of oncogene expression ranges in transcriptomic information may not be reflective of their true significance.
scRNA-seq transcriptome evaluation
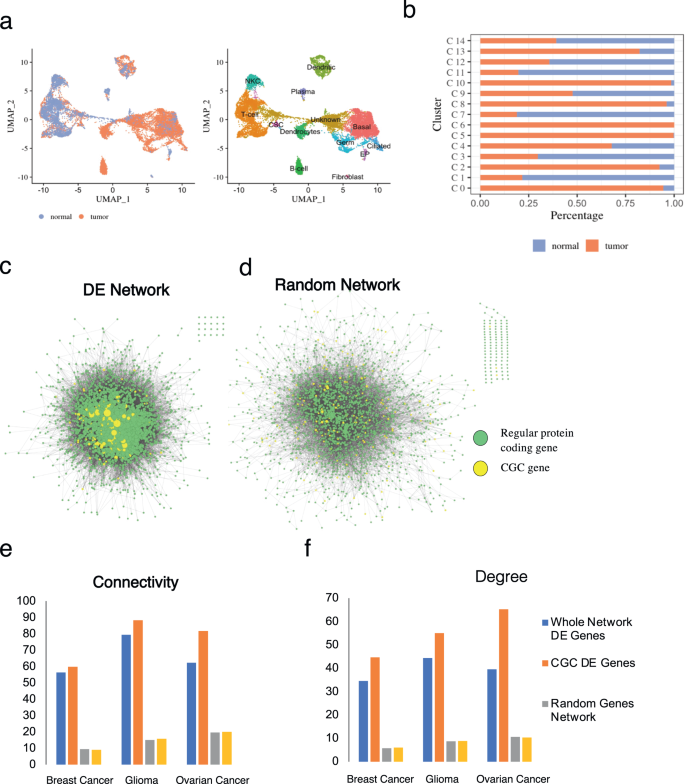
The benefit of single-cell sequencing is the truth that it provides a extra in-depth overview of particular person cell expression ranges inside numerous subpopulations, which is important contemplating the advanced nature of tumor microenvironments that make use of a wide selection of cell sorts throughout tumor development. Subsequently, to research the expression patterns of oncogenes inside numerous cell populations we searched the GEO database for scRNA-seq datasets that contained affected person tumor and regular samples. We chosen three scRNA-seq datasets from the GEO database (“Strategies”) composed of paired regular and tumor affected person tissue samples for breast most cancers, ovarian most cancers, and glioma. We first carried out high quality management and normalization, after which we proceeded to combine regular and tumor samples for concurrent evaluation of most cancers gene expression (“Strategies”). We noticed a various microenvironment (Supplementary Tables 3–5) in all three most cancers sorts (Fig. 4a and Supplementary Fig. 8a, c, g), with every cluster comprising completely different proportions of most cancers and regular subpopulations (Fig. 4b and Supplementary Fig. 8b, d).
a uMAP dimensional discount for built-in tumor and regular samples of ovarian most cancers the place completely different colours symbolize completely different cell populations. b Cell situation composition breakdown per cluster in ovarian most cancers. DE evaluation recognized DE genes between regular and tumor cells in every cluster. These DE genes had been utilized in Cytoscape to generate PPI networks for c DE community and random community of the identical dimension from d random sampling of genes for comparability. Within the PPI networks, node dimension scaled to the diploma of the gene, and highlighted the CGC genes (yellow). The typical e connectivity and f diploma properties of the entire DE community (blue), and CGC DE genes (orange) are greater when in comparison with the random community (grey), and the random subset of genes (yellow).
To confirm whether or not there exist cancer-specific variations in gene expressions at every cluster, we carried out Differential Expression (DE) evaluation between tumor and regular cells of the preserved clusters throughout the 2 circumstances (e.g., cluster 9, Fig. 4b). Our analyses present that 5–10% of all of the recognized DE genes in each cluster and throughout all three most cancers sorts are CGC genes (Desk 5). Amongst the differentially expressed CGC genes (Desk 6), we noticed and visualized a number of oncogenes which have been beforehand studied for his or her function in numerous cancers (e.g., TMSB4X and TNFAIP3, Supplementary Fig. 9). Moreover, regardless of representing solely a small fraction of the community, CGC genes additionally exhibit particular connectivity properties (“Strategies”) throughout the community of DE genes (Fig. 4c, e, f). To keep away from pattern dimension bias in our outcomes, we equally generated random networks composed of the equivalent variety of nodes because the DE community (Fig. 4d). For these random units of genes of equivalent dimension, the differentially expressed DE genes exhibit common connectivity properties much like that of the bigger networks, that’s, of decrease connectivity in comparison with CGC genes. (Fig. 4e, f). This result’s in line with our earlier evaluation, and with the truth that CGC genes are usually higher related as a result of them being higher studied within the literature.
Taken collectively, these outcomes present certainly that there exist notable variations between regular and tumor tissues that emerge on a single-cell stage from the heterogeneous tumor microenvironment (mirrored by cell clusters). Differential expression between regular and tumor cell sorts in cell clusters, in addition to the improved connectivity properties of most cancers genes spotlight the truth that oncogenes may be distinguished from different genes as a particular subset.
Collectively, the evaluation of single-cell transcriptomic information revealed not solely a heterogeneous tumor microenvironment (mirrored by cell clusters), but in addition heterogeneous expression patterns for oncogenes that can not be generalized on a world inhabitants stage. Moreover, differential expression evaluation between comparable cell clusters in tumor and regular samples additional highlighted the truth that oncogenes may be distinguished from different genes as a particular subset with distinct community connectivity properties.