1 Introduction
Colorectal most cancers (CRC) is without doubt one of the most prevalent malignant tumors of the digestive system. In keeping with the American Most cancers Society, roughly 81,860 sufferers with CRC have been identified and 28,470 deaths occurred in america of America in 2023, inflicting severe issues for sufferers and public well being (1). Though endoscopic screening has decreased the mortality and morbidity charges of CRC in recent times, and recurrence and metastasis stay main challenges (2). Presently, main therapies for CRC embrace surgical procedure, chemotherapy, and radiotherapy. However, advances in immunotherapies, together with anti-PD-1, anti-PD-L1, and anti-CTLA4 therapies, have offered a brand new and promising therapeutic paradigm for CRC with vital potential efficacy (3). As an illustration, the profitable anti-PD-1 utility in sufferers with CRC and mismatch restore deficiency (dMMR) or excessive microsatellite instability (MSI-H) considerably causes progression-free survival in CRC (4, 5). A brand new mixture of radiotherapy and immunotherapy promotes strong antitumor immune priming (6, 7). Nevertheless, these methodologies face constraints arising from spatiotemporal heterogeneity, average precision, or restricted illustration of inhabitants subsets (8–10). Consequently, within the context of customized therapy paradigms, the identification of sturdy biomarkers is crucial for optimizing prognosis and enhancing the efficacy of drug therapies for CRC.
Ferroptosis, pushed by biochemical and genetic elements, is a programmed cell demise pathway reliant on iron and activated by lipid peroxide buildup on mobile membranes. Its involvement extends to tumor development and therapeutic responses throughout varied malignancies and is commonly intertwined with reactive oxygen species (ROS) that take part in cancer-related pathways (10). Fatty acid metabolism is a vital mobile course of that transforms vitamins into metabolic intermediates used for membrane synthesis, vitality reservation, and signaling molecule manufacturing. This course of has garnered vital consideration as a possible goal for most cancers remedy, significantly as a result of it’s related to regulatory and CD8+ T cells (11–13). Glutathione peroxidase 4 (GPx4) and prolyl hydroxylase-3 (PHD3) symbolize vital regulators of ferroptosis and fatty acid metabolism, respectively, emphasizing the potential position of each in immunotherapy (14, 15). Latest research indicated a major affiliation between fatty acid metabolism and ferroptosis. Microsomal triglyceride switch protein (MTTP)expression will increase within the physique throughout fatty acid metabolism, which inhibits ferroptosis and reduces the density of chemotherapy (16). Related outcomes have been noticed for phospholipids containing a single polyunsaturated fatty acyl tail (PL-PUFA1s), that are additionally strongly correlated with ferroptosis (17). Moreover, cytochrome P450 1B1 (CYP1B1) and cyclin-dependent kinase 1 (CDK1) degrade acyl-CoA synthetase long-chain member of the family 4 (ACSL4), who performs an important position in fatty acid metabolism and inhibits ferroptosis, thereby inducing resistance to anti-PD-1 and oxaliplatin, respectively (18, 19). Thus, the fatty acid metabolism-related genes seem to control ferroptosis and performance as intermediates. The connection between ferroptosis regulators and fatty acid metabolism-related genes, which can considerably affect prognosis and drug resistance in colorectal most cancers, has been much less explored. The tumor microenvironment (TME), which consists of tumor cells, stromal cells, and immune cells, performs an irreplaceable position within the metastasis and tumor development and in addition impacts the efficacy of immune checkpoint blockade (ICB) therapy (20, 21). Contemplating the particular relationship between ferroptosis and fatty acid metabolism regulators, a novel TME could induce novel metabolic pathways in CRC. Thus, the interactions between ferroptosis and fatty acid metabolism regulatory molecules needs to be explored in multicenter cohorts from a multi-omics perspective, together with the TME, immunotherapy, and epigenetic mutations.
On this research, we performed an intensive pan-cancer multi-omics evaluation to look at the molecular correlations between ferroptosis and fatty acid metabolism regulators in 33 most cancers sorts. By performing unsupervised clustering, we recognized three distinct clusters associated to ferroptosis and fatty acid metabolism primarily based on the TME, gene expression, and organic capabilities. Using a sturdy mixture of 117 machine-learning algorithms, we developed the FeFAMscore, which demonstrated superior predictive efficiency in each the coaching cohort and 4 impartial exterior validation cohorts. It additionally successfully reveals potential in forecasting immunotherapy and chemotherapy drug sensitivity in CRC sufferers. Total, the FeFAMscore is promising for the development of novel therapy methods, fostering a nuanced and customized method to medication.
2 Strategies
2.1 Knowledge acquisition and pre-processing
The workflow is illustrated in Supplementary Determine S1. The Gene Expression Omnibus (GEO) (https://www.ncbi.nlm.nih.gov/geo/) and The Most cancers Genome Atlas (TCGA) (https://portal.gdc.most cancers.gov/databases have been used to acquire the CRC RNA expression profiles, along with the related complete scientific annotations, together with TCGA-COAD, TCGA-READ, GSE17536, GSE17537, GSE29621, GSE38832, and GSE39582. The Meta-cohort and Prepare cohorts (TCGA-COAD, TCGA-READ, and GSE39582) have been established and the batch results have been estimated utilizing the “sva” bundle in R software program. Moreover, three immunotherapy cohorts with completely different immunotherapy efficacies downloaded from the TIGER web site (http://tiger.canceromics.org/), together with IMVigor210 (anti-PD-L1), Braun (anti-PD-1), and PRJNA23709 (anti-PD-1 + anati-CTLA4) have been investigated. The microarray information from the GEO have been normalized and corrected background by the “impute” R bundle. The ferroptosis regulators and fatty acid metabolism-related genes investigated on this research have been extracted from FerrDb (22) and MSigDB (https://www.gsea-msigdb.org/gsea/msigdb/) (Supplementary Desk S1). Lastly, 1448 sufferers with survival data have been acquired from the database. The info of Copy Quantity Variation (CNV) is extracted from TCGA database in 33 cancers and analyzed by the “matfool” packages.
2.2 Unsupervised clustering of ferroptosis regulators and fatty acid metabolism-related genes
The tumor-related FeFAM genes have been obtained from TCGA database utilizing “limma” and “survival” packages within the R software program. Univariate Cox evaluation was used to filter the 50 prognosis genes within the coaching cohort primarily based on p<0.05. Subsequent, the coaching cohort was subjected to unsupervised clustering to establish distinct patterns. The potential groupings have been delineated utilizing Ok-means clustering evaluation with various cluster numbers (ok = 2–9) (23). We then carried out the “Nuclst” bundle to confirm probably the most applicable clusters with 28 standards and repeated 1000 instances on resample charge of 0.8 to validate the classification stability. Principal element evaluation (PCA) was subsequently employed to validate the clustering outcomes utilizing the expression profiles of those genes. This evaluation confirmed the co-expression patterns of ferroptosis regulators and fatty acid metabolism-associated genes.
2.3 Cell infiltration estimation
We evaluated the immune cell microenvironment utilizing the CIBERSORT algorithm, EPIC algorithm, MCPCOUNTER algorithm, TIMER algorithm, quantiseq algorithm, and XCELL algorithm of the “IBOR” and “GSVA” R bundle. Single-sample gene set enrichment evaluation (ssGSEA) algorithm was used to confirm the outcomes. Moreover, the TIDE algorithm (http://tide.dfci.harvard.edu/) was used to guage the tumor immune dysfunction and exclusion (TIDE) rating, CAF, dysfunction and exclusion of immune cells, PD-L1, and cytotoxic T cells (CTL) rating. A excessive TIDE rating could mirror poor ICI efficacy.
2.4 Pathway enrichment evaluation
To investigated the organic distinction between three patterns and cancer-related pathways, we downloaded “h.all.v7.5.1.symbols.gmt” and “c2.cp.kegg.v7.4.symbols” from the MsigDB database (c2.cp.kegg.symbols), and analyzed utilizing the GSVA program. We additional explored the variations in cancer-, immune-, and metabolism-related patterns as reported beforehand (24–27). The pathways with the very best expression among the many three patterns with p<0.05 have been thought-about activated.
2.5 FeFAMscore prognostic mannequin building
To additional discover the biofunction and prognostic worth of FeFAM genes, we first randomly mixed 10 machine studying algorithms, together with random survival forest (RSF), elastic community (Enet), Ridge, Stepwise Cox, Lasso, CoxBoost, partial least squares regression for Cox (plsRcox), generalized boosted regression modelling (GBM), supervised principal elements (SuperPC), and survival help vector machine (survival-SVM), as reported beforehand (28). Then, the coaching cohorts have been enter because the coaching group to the mixed 117 algorithms, and every mannequin was detected in 4 impartial datasets (GSE17536, GSE17537, GSE29621, and GSE38832). Subsequent, Harrell’s concordance index (C-index) was calculated for every mannequin throughout all validation cohorts utilizing the FeFAMscore derived from the coaching cohorts. Primarily based on the common C-index in all validation cohorts, we chosen the optimum mannequin and in contrast the FeFAMscore with these of 69 revealed fashions up to now decade, which proved its dependable and strong predictive energy.
2.6 Cell tradition
Regular colon mucosal epithelial cells (NCM460) and HCT116, DLD-1, and CACO2 cell traces have been obtained from the Chinese language Academy of Sciences (Shanghai, China), cultured in DMEM supplemented with 10% fetal bovine serum (each from Thermo Fisher Scientific, Waltham, MA, USA), and maintained underneath customary cell tradition situations (37°C, 5% CO2) in a cell incubator.
2.7 RNA extraction and RT-qPCR
Mobile and tissue RNA was extracted utilizing TRIzol reagent (R411-01, Vazyme, Nanjing, China), adopted by reverse transcription utilizing HiScript III RT SuperMix (R323, Vazyme). Quantitative PCR evaluation was carried out utilizing the Common SYBR Inexperienced Quick qPCR Combine (ABclonal, Hong Kong, China, RK21203). The info have been analyzed utilizing the two(−ΔΔCt) technique, with GADPH serving as the interior management. The primer sequences are offered in Supplementary Desk S1.
2.8 siRNA transfection
siRNA-ACAA2-1 or siRNA-ACAA2 (GenePharma, Shanghai, China) was used to silence the ACAA2 gene. The siRNA sequences have been as follows: si-ACAA2-1 (sense: 5′-UGCUGAGACAGUGAUUGUATT-3′; antisense: 5′-UACAAUCACUGUCUCUCATT-3′), and si-ACAA2-2 (sense: 5′-GGGCACTGAAGAAAGCAGGA-3′; antisense: 5′-CGTGAACCAGGTGTGCAGTA-3′). Transfection was carried out utilizing Lipofectamine 3000 (Thermo Fisher Scientific) following the producer’s directions.
2.9 Cell viability assay
CRC cell viability was evaluated utilizing the Cell Counting Equipment 8 (CCK-8, Dojindo, Japan). Briefly, 3000–5000 cells have been seeded per effectively in 96-well plates. Subsequently, 100 μL medium containing 10 μL CCK-8 resolution was added to every effectively and incubated at 37°C for 3 h. The absorbance at 450 nm was measured.
2.10 Transwell assay
HCT116 and CACO2 cell traces transfected with siRNAs concentrating on ACAA2 (si-ACAA2-1 and si-ACAA2-2) or non-targeting management siRNA (si-NC) have been harvested, washed twice with PBS, and resuspended in DMEM. The suspended cells have been then positioned within the higher chamber of 24-well chambers outfitted with 8 μm pore inserts.
2.11 Colony formation assay
To guage colony formation within the monolayer tradition, 1000 cells have been seeded in 6-well plates. Following two weeks of tradition, the colonies have been mounted and stained with 4% paraformaldehyde and 0.1% crystal violet for 30 min at room temperature.
2.12 Western blot
Protein focus was decided utilizing the BCA Protein Assay Equipment (Thermo Fisher Scientific, USA). Samples containing 30 μg of protein have been separated on a 12% SDS-PAGE gel and transferred onto a PVDF membrane. The membrane was blocked with 5% BSA for two hours after which incubated in a single day at 4°C with the first antibody. Afterward, the membranes have been incubated for 1 hour with the secondary antibody and washed thrice with TBST buffer. Antibody alerts have been detected utilizing the ECL system (Bio-Rad, California, USA).
2.13 Immunotherapeutic response prediction
We predicted the immunotherapy response of the FeFAMscore by analyzing the expression of tumor mutational burden (TMB), TIDE rating, and variations in pathway enrichment. Primarily based on these outcomes, we calculated the FeFAMscore of sufferers within the coaching cohort to discover the perform of the FeFAMscore in immunotherapy. Subsequently, we used Subclass Mapping (Submap) to find out the connection between excessive or low FeFAMscore teams and anti-PD-1 and anti-CTLA4 antibodies. As well as, we utilized immunotherapy cohorts with scientific response data to validate the immunotherapy response. The IMVigor210CoreBiologies R bundle was used to acquire transcriptome, survival, and immunotherapy efficacy information for the IMVigor210 cohort (29). The anti-PD-1 and anti-CTLA4 cohorts have been validated utilizing Braun and PRJNA23709.
2.14 Chemotherapeutic sensitivity prediction
The correlation with FeFAMscore and drug sensitivity was predicted by the GDSC and CTRP datasets with “oncoPredict” packages. The relation with gene expression and drug sensitivity was measured by the “Hmsic” bundle in R software program.
2.15 Statistical evaluation
Knowledge processing and visualization have been carried out utilizing R software program (model 4.3.2) and GraphPad Prism 8.0, respectively. Group comparisons have been carried out utilizing the Wilcoxon check for pairwise comparisons, whereas ANOVA and Kruskal–Wallis checks have been used to evaluate variable distributions amongst a number of teams, contemplating normality assumptions. Categorical variables have been analyzed utilizing the chi-square and Fisher’s actual checks. Correlations have been decided utilizing the Spearman and Pearson methods. Survival disparities have been evaluated utilizing the Kaplan–Meier technique and log-rank check. Statistical significance was set at p<0.05, and all p-values have been two-tailed.
3 Outcome
3.1 Identification of novel correlations between ferroptosis and fatty acid metabolism regulators
To discover the connection between cell demise and metabolism, we investigated the crosstalk between ferroptosis-associated regulators and fatty acid metabolism-regulating genes. The 486 ferroptosis-associated genes derived from the FerrDb included markers, suppressors, and drivers (Supplementary Desk S1). In the meantime, 272 FAM genes have been retained from the MSigDB database. Genome-wide omics information for 33 most cancers sorts have been retrieved from the TCGA database for evaluation. The frequency of mutations in these genes was considerably correlated between ferroptosis and fatty acid metabolism within the tumors (Determine 1A). PCA was carried out to measure the degrees of ferroptosis-associated and FAM genes within the 33 most cancers sorts database, then the Spearman’s evaluation additional demonstrated a major correlation between them (Supplementary Determine S1A). Apparently, the COREAD database, with the most important variety of sufferers, exhibiting a distinguished affiliation between them (R = -0.93) (Figures 1B, C). Consequently, to additional discover the colorectal most cancers, the highest 10 mutations in ferroptosis-associated regulators and fatty acid genes have been recognized in 480 (95.98%) of 497 sufferers with COREAD. The very best mutation frequencies have been detected in TP53 (67%), KRAS (44%), and PIK3CA (26%) (Supplementary Determine S2B). The exploration of copy quantity variation (CNV) alteration frequency confirmed a excessive incidence of CNV positive aspects within the TCGA cohort, demonstrating the potential for remedy in CRC (Supplementary Determine S2C). The areas of FeFAM genes with CNVs on the chromosomes are marked within the Circle Map (Supplementary Determine S2C). Moreover, co-mutations have been widespread amongst these genes (Supplementary Figures S2E, F).
Determine 1 Panorama of genetic and relation of FeFAM regulators and discovery of novel FeFAM patterns. (A) Mutation frequency of FeFAM regulators in 33 sorts cancers in TCGA. (B) PCA of FeFAM regulators within the CRC. (C) Scatter plot confirmed the spearmen correlation of FeFAM regulators. (D) Volcano plot confirmed the differential FeFAM regulators in CRC. (E) KEGG and GO analyze of fifty OS-related FeFAM regulators. (F) Community confirmed the interactions amongst FeFAM regulators in CRC. (G) A The consensus rating matrix of all samples when ok = 3. (H) The CDF curves of consensus matrix for every ok (indicated by colours). (I) Principal element (PC) evaluation revealed exceptional distinction between three FeFAM patterns from practice cohort (n = 1029). (J) Kaplan-Meier curves of survival for 3 FeFAM patterns primarily based on CRC sufferers from practice cohort. (Ok) This boxplot demonstrates the expression variations within the FeFAM-related genes amongst three FeFAM patterns. The highest portion represented Fisher’s exact check. The decrease portion indicated the Wilcoxon rank-sum check. ***p < 0.001. FeFAM, ferroptosis and fatty acid metabolism regulators.
Primarily based on the evaluation of outcomes, we in contrast the CRC and regular samples from COREAD databases in TCGA, which lastly recognized 159 ferroptosis-associated and fatty acid metabolism-related genes in accordance with logFC>1 and FDR <0.05 (Determine 1D). To counterbalance the implications between TCGA and GEO database, we enrolled COREAD database and GSE39582 and adopted the “sva” bundle to take away batch results and extract related genes because the coaching cohort. A complete of fifty genes have been subsequently screened utilizing univariate Cox regression evaluation (P <0.05) of the FeFAM genes within the mixed database (Supplementary Determine S2G). The KEGG evaluation, depicted utilizing a barplot, revealed enrichment of those genes in pathways similar to “PPAR signaling pathway”, “fatty acid degradation”, “fatty acid metabolism”, “propanoate metabolism”, “tyrosine metabolism”, and “p53 signaling pathway”. Moreover, GO evaluation of molecular capabilities (MF), organic processes (BP), and mobile elements (CC) highlighted their relevance in fatty acid metabolism, response to oxygen ranges, and most cancers pathways. (Determine 1E). Community evaluation provided a holistic view of the prognostic implications and molecular interactions throughout the FeFAM framework (Determine 1F). Contemplating the discernible variations within the transcriptional profiles and the distinctive interaction between these molecules, dysregulation throughout the FeFAM community considerably contributes to CRC initiation and development.
3.2 Discovery of novel FeFAM patterns by unsupervised clustering evaluation
To elucidate the potential FeFAM phenotypes in CRC, we utilized Ok-means-based unsupervised clustering within the coaching cohort. The R bundle “ConsensusClusterPlus” was used to initially categorize the sufferers with CRC into ok (ok=2–9) FeFAM clusters (Determine 1G; Supplementary Figures S3A-Ok). The cumulative distribution perform (CDF) curves, derived from the consensus rating matrix and PAC statistics, elucidated the optimum variety of clusters (ok=3) throughout the complete coaching affected person cohort. These clusters, denoted A, B, and C, exhibited discernible segregation patterns (Determine 1H). Nbclust testing, which included 28 standards, yielded the identical outcomes (Supplementary Determine S3L). The PCA demonstrated a transparent distinction between the three clusters (Determine 1I). The Kaplan–Meier curve confirmed that cluster C had higher survival prognosis than clusters A and B (p<0.001) (Determine 1J). The expression of FeFAM genes additionally indicated the flexibility to distinguish between the three subtypes (Determine 1K).
3.3 TME characterization in numerous FeFAM patterns
The ssGSEA technique, which simulates the complete tumor immune course of, was first used to calculate tumor immune cell infiltration within the coaching cohorts to analyze the variations within the TME (Determine 2A). Subsequent, six completely different algorithms, similar to CIBERSORT, EPIC, MCPCOUNTER, TIMER, QuantiSeq, and XCELL, obtained the identical outcomes, verifying the essential results of FeFAM genes within the immune system (Supplementary Determine S4A). In the meantime, in accordance with the Spearman’s correlation evaluation, nearly all FeFAM genes have been considerably implicated within the immune microenvironment composition (Determine 2B). Ferroptosis-associated regulators, together with ENPP2, CAV1, FABP4, PDK4, ADIPOQ, NOX4, COKN2A, CDO1, WWTR1, DDR2, CPEB1, and TIMP1, are preferentially related to probably the most immunosuppressive cells, whereas FAM genes correlated with immune microenvironment activation. Moreover, Spearman’s evaluation demonstrated vital co-expression of the Fe and FAM genes (Supplementary Determine S4B). In comparison with FeFAM clusters B and C, FeFAM cluster A had a considerably worse prognosis. Evaluation of gene signatures revealed an elevated presence of immune cells exhibiting notable immunosuppressive performance, macrophages, regulatory T cells (Tregs), and kind 2 T helper cells, together with inside FeFAMA cluster A throughout all cohorts. Remarkably, CD4+ T cells, CD8+ T cells, neutrophils, dendritic cells, and pure killer (NK) cells have been considerable within the FeFAM cluster A throughout practically all of the algorithms, suggesting that the immune cells inside FeFAM cluster A could concurrently govern immune evasion and anti-tumor actions. Regardless of having related prognoses, FeFAM clusters B and C exhibited contrasting ranges of immune infiltration, implying that they might have completely different immunotherapeutic potentials. The XCELL and ESTIMATE algorithms additionally demonstrated excessive immune, stroma, and microenvironment scores in FeFAM clusters A and C (Supplementary Determine S4C). Consequently, FeFAM clusters A, B, and C have been thought-about immune-excluded, immune-desert, and immune-activated clusters, respectively. To make sure stability of the outcomes, the TIDE algorithm, which is often adopted to measure immune escape ranges and ICD therapy efficacy, was used. We found that FeFAM cluster A confirmed the very best CAF, IFN, TIDE, PD-L1 and exclusion among the many three clusters, according to the above evaluation outcomes and presumably attributed to immune escape (Figures 2C–J). Apparently, though FeFAM cluster A had extra MDSCs than FeFAM cluster C, their CTL.scores and dysfunction have been related, suggesting that activated ferroptosis and fatty acid pathways could inhibit intratumoral CD8+ T cell effector perform and impair their anti-tumor potential, which was just like beforehand reported outcomes (12, 30–32). Above all, ferroptosis regulators could cooperate with fatty acid metabolism-associated genes to contribute to a specific immune microenvironment, thereby presenting potential targets for immunotherapy.
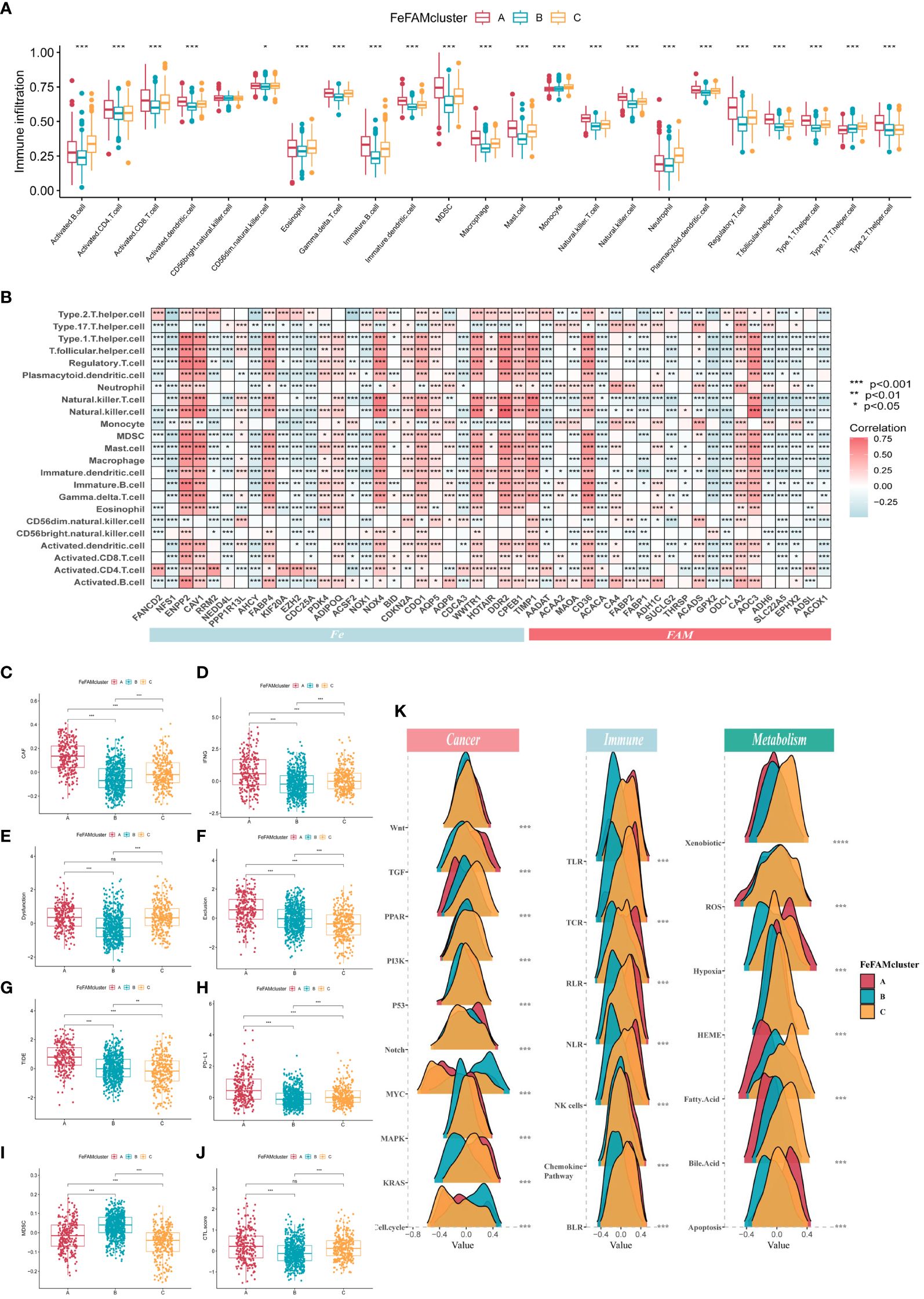
Determine 2 Characterization of tumor microenvironment, signing and immune pathways in numerous FeFAM patterns. (A) Traits of immune infiltrating cells in numerous FeFAMclusters. (B) Traits of immune infiltrating cells in numerous FeFAM regulators. (C-J) Field plots confirmed the numerous distinction in CAF (C), IFNG (D), Dyfunction (E), Exclusion (F), TIDE (G), PD-L1 (H), MDSC (I), and CTL.rating (J). (Ok) The cancer-related, immune-related and metabolism-related pathways between the three FeFAM subtypes. * p < 0.05, ** p < 0.01, *** p < 0.001, ns, not vital. FeFAM, ferroptosis and fatty acid metabolism regulators.
3.4 Signaling and immune pathway variations between the FeFAM patterns
Using the “gsva” bundle, we executed GSVA-enrichment experiments to analyze varied cancer-related signaling pathways throughout the three patterns throughout the Hallmarker and KEGG pathways (Supplementary Determine S4D). The findings point out that FeFAM cluster A was considerably enriched in immune- and tumor-related pathways, similar to “apoptosis”, “epithelial mesenchymal transition (EMT)”, “inflammatory response”, and “VEGF signaling pathway”. The FeFAM clusters B and C are two distinct teams with particular associations by way of their organic capabilities and metabolic pathways. FeFAM cluster B, as an example, is considerably related to “DNA restore and replication”, “protein export”, and “spliceosome”, whereas FeFAM cluster C is related to the metabolic pathway “fatty acid metabolism”, “linoleic acid metabolism”, “nicotinate and nicotinamide metabolism” and “nitrogen metabolism”. Consequently, we additional investigated the carcinogen-signaling, immune-related, and metabolic pathways to match the variations among the many three patterns. Wnt, TGF, Notch, MAPK, KRAS, TLR, TCR, RLR, NK cells, chemokine pathway, hypoxia, and apoptosis have been activated in cluster A. PPAR, PI3K, P53, xenobiotics, ROS, HEME, fatty acids, and bile acids have been activated in cluster C, indicating that ferroptosis and fatty acid metabolism could also be upgraded to enhance prognosis and stop immune escape (Determine 2K). These analyses offered extra proof that FeFAM molecules regulate the immune microenvironment and facilitate immune evasion in sufferers with CRC by various signaling pathways. This underscores the potential of FeFAM as a promising goal for immunotherapy.
3.5 Built-in building and constant prognostic worth of the FeFAMscore
Primarily based on the various expression ranges of FeFAM genes among the many three patterns, we subjected the 50 FeFAM-related genes to our machine learning-based integrative method to assemble an FeFAM-related signature, termed the FeFAMscore. Within the coaching cohort, 117 algorithms generated from a random permutation of 10 machine-learning algorithms have been employed to compute the C-index utilizing a 10-fold cross-validation framework. The mannequin was subsequently evaluated throughout 4 cohorts to gauge its predictive efficacy and to find out its consistency throughout completely different datasets. Following this analysis, the mannequin with the very best common C-index among the many validation cohorts was recognized. Particularly, the mixture of CoxBoost and StepCox (backward and each) yielded the very best common C-index of 0.689, establishing it because the optimum mannequin (Determine 3A; Supplementary Desk S2). Fifteen genes have been first screened utilizing the CoxBoost mannequin after which subjected to backward Cox proportional hazards regression. A closing set of 15 genes was recognized, together with KIF20A, ACSF2, NOX1, BID, AADAT, ACAA2, FABP1, CA2, SLC22A5, PPP1R13L, AQP5, HOTAIR, DDR2, TIMP1, and CD36 (Determine 3B). Subsequently, the FeFAM rating for every affected person was decided by using the expression ranges of 15 genes, which have been weighted utilizing the regression coefficients obtained from a Cox mannequin (Determine 3B). Subsequently, all sufferers have been dichotomized into high- and low-FeFAMScore teams. These 15 genes considerably distinguished high-risk people from low-risk people (Figures 3C–H). Sufferers categorized into the excessive FeFAMscore group exhibited considerably poorer total survival (OS) in comparison with these within the low FeFAMscore group, as decided by Kaplan–Meier survival evaluation in each the mixed coaching (N=1029, P<0.001) and 4 validation datasets: GSE17536 (N=177, P<0.001), GSE17537 (N=55, P=0.003), GSE29621 (N=65, P=0.007), and GSE38832 (N=122, P<0.001). The same final result was noticed within the meta-cohort (N=1448), thereby affirming the predictive accuracy and reliability of the mannequin. An alluvial diagram illustrates the correlation between FeFAMcluster and FeFAMscore (Determine 3I).
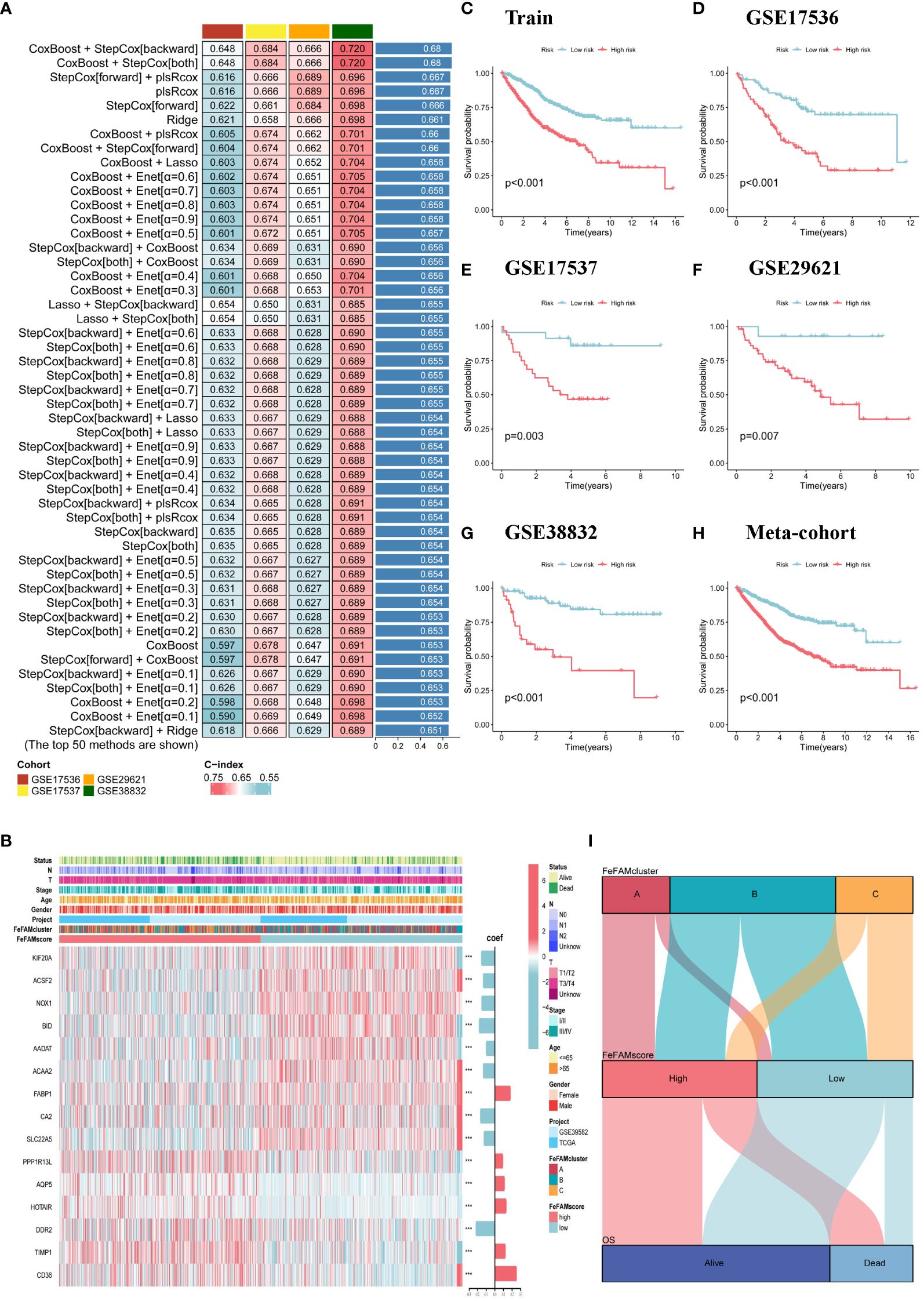
Determine 3 Development of a machine learning-based signature. (A) The highest C-index of fifty machine studying strategies in 4 validation cohorts. (B) The heatmap demonstrates the relationships between the three FeFAM phenotypes, clinicopathologic traits, coef worth and the expression variations of the FeFAM-related genes in practice cohort. (C–H) Kaplan-Meier curves of OS in accordance with the FeFAMscore in Prepare cohorts (log-rank check: p<0.001) (C). GSE17536 (Log-rank check: p<0.001) (D). GSE17537 (Log-rank check: p = 0.003) (E). GSE29621 (Log-rank check: p = 0.007) (F). GSE38832 (Log-rank check: p < 0.001) (G). Meta-cohort (Log-rank check: p < 0.001) (H). (I) Alluvial diagram exhibiting the correlation of FeFAMclusters and FeFAMscore. *** p < 0.001. FeFAM, ferroptosis and fatty acid metabolism regulators.
3.6 Constant prognostic worth of FeFAMscore
Receiver working attribute (ROC) curve evaluation was performed to guage the discriminative potential of the FeFAMscores. Within the coaching cohort, the areas underneath the ROC curve (AUC) for 1-, 3-, and 5-year survival have been 0.701, 0.712, and 0.668, respectively. Moreover, wonderful outcomes have been additionally indicated within the check cohorts, together with 0.738, 0.718, and 0.64 in GSE17536; 0.729, 0.684, and 0.687 in GSE17537; 0.769, 0.670, and 0.649 in GSE29621; and 0.799, 0.773, and 0.708 in GSE38832 (Determine 4A). The meta-cohort of those sufferers confirmed AUC values of 0.687, 0.683, and 0.644, indicating that the FeFAMscore mannequin is predictive and dependable in a number of impartial CRC cohorts (Determine 4A).
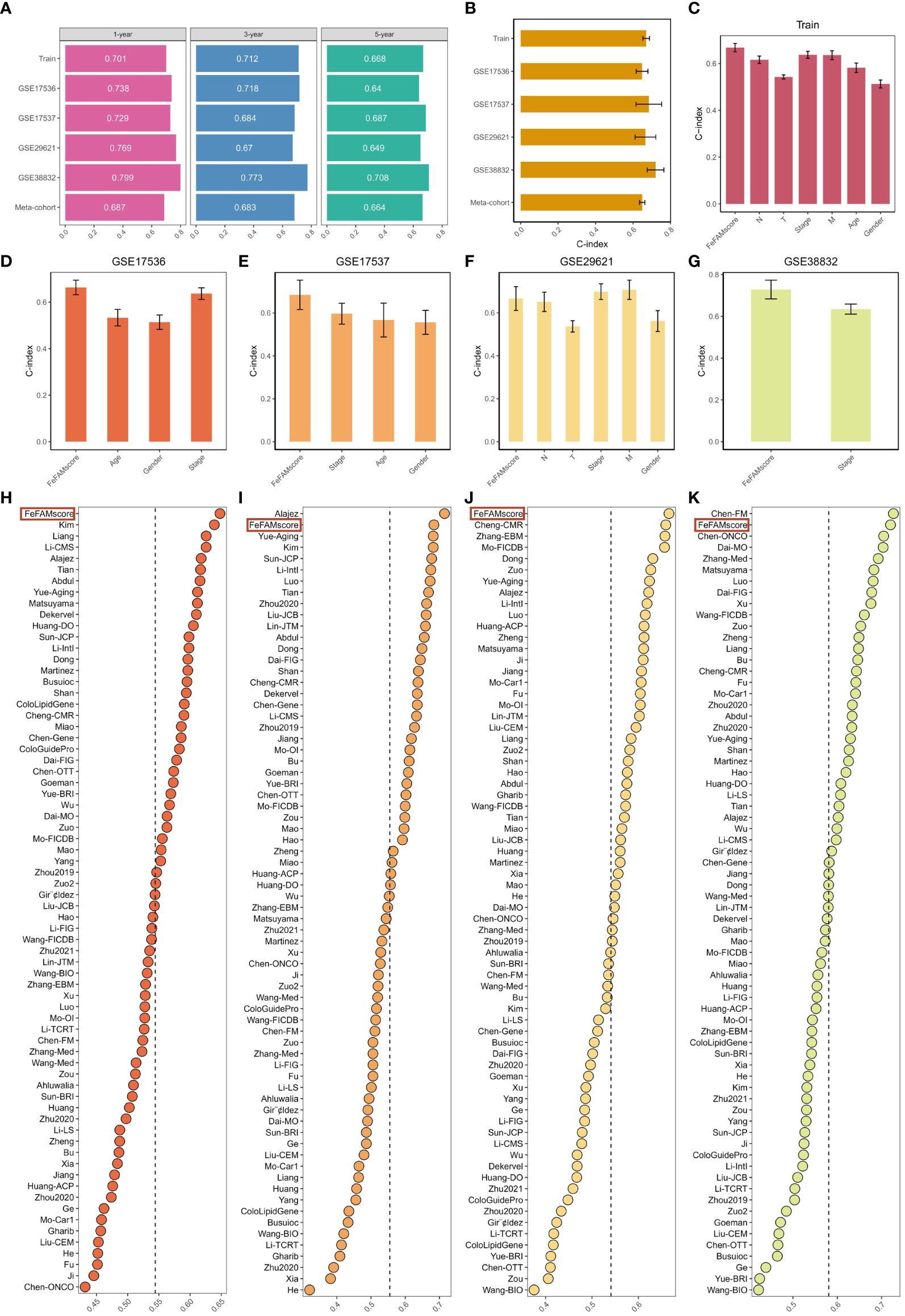
Determine 4 Comparability between the FeFAMscore and the opposite 69 signatures in colorectal most cancers. (A) Time-dependent ROC evaluation for predicting OS at 1,3, and 5 years in practice cohort (n = 1029), GSE17536 (n = 177), GSE17537 (n = 55), GSE29621 (n = 65), and GSE38832 (n = 122). (B) C-index of FeFAMscore throughout all datasets. (C-G) The efficiency of FeFAMscore was in contrast with different scientific variables in predicting prognosis. Prepare (C), GSE17536 (D), GSE17537 (E), GSE29621 (F), and GSE38832 (G). (H-Ok) C-index of FeFAMscore and 69 revealed signatures in GSE17536 (H), GSE17537 (I), GSE29621 (J), and GSE38832 (Ok). *** p < 0.001. FeFAM, ferroptosis and fatty acid metabolism regulators.
The C-index [95% confidence interval] was 0.67 [0.652–0.688], 0.648 [0.617–0.679], 0.684 [0.639–0.807], 0.666 [0.614–0.790], 0.720 [0.649–0.804], 0.649 [0.646–0.711] within the 4 impartial validation cohorts and meta-cohorts, respectively (Determine 4B). To foretell affected person prognosis, scientific traits together with age, intercourse, T stage, N stage, M stage, and stage are broadly acknowledged. Subsequently, the C-index was utilized to measure the predictive accuracy between the FeFAMscore and scientific traits within the coaching and 4 impartial validation cohorts. The FeFAMscore exhibited considerably larger predictive accuracy than different scientific traits within the coaching, GSE17536, GSE17537, and GSE38832 cohorts (Figures 4C–G). In distinction, the efficiency of the FeFAMscore in GSE29621 cohort was just like that of the M stage and phases, which can have been because of the small pattern measurement and information bias. These outcomes point out that FeFAMScore could also be a potential different biomarker for predicting survival danger in scientific apply.
3.7 Resilient predictive efficiency of FeFAMscore
Because the sequencing depth regularly will increase, CRC therapy outcomes are effectively predicted. Machine learning-based prognostic fashions for CRC have been more and more developed in recent times. To quantify the efficiency of the FeFAMscore, we systematically retrieved mRNA signatures from CRC analysis over the previous decade and at last acquired 69 mRNA signatures. We in contrast the predictive potential of the FeFAMscore utilizing the C-index worth within the 4 impartial validation cohorts. The FeFAMscore ranked first within the GSE17536 and GSE29621 datasets, adopted by GSE17537 and GSE38832 (Figures 4D-G). Nevertheless, some fashions exhibited considerable predictive efficiency for the GSE17537 and GSE38832 datasets and carried out reasonably in different cohorts, additional proving the individuality and reliability of our fashions. The Chen-FM mannequin, as an example, confirmed a greater C-index than the FeFAMscore in GSE38832 and was poorly displayed in GSE17536, GSE17537, and GSE38832 with a C-index of lower than 0.6. The above outcomes demonstrated the great predictive efficiency of the FeFAMscore (Figures 4H–Ok).
3.8 ACAA2 is related to tumor development in CRC
To additional consider the expression and performance of the FeFAMscore, we first carried out RT-qPCR in cell traces from sufferers with CRC for the six genes. The opposite 9 genes concerned within the FeFAM rating have been demonstrated by different researchers (33–41). In contrast with these in regular human colonic cells (NCM460 cells), the expression of ACAA2 was considerably larger in HCT116 and CACO2 cells, whereas the expression of ACSF2, DDR2 and SLC22A5 was considerably elevated within the CRC cells (Figures 5A, B). Among the many expression and correlation of the genes, ACAA2 was considerably overexpressed within the tumor tissues and powerful correlated with ferroptosis regulators. We then used two small interfering RNAs (siRNAs) to downregulate ACAA2 expression in HCT116 and CACO2 CRC cell traces (Determine 5C). The western blot additional demonstrated the outcomes (Determine 5D). Cell viability was decreased by ACAA2 downregulation after 72 h (Figures 5E, F). As well as, Cell colony formation experiments demonstrated a major discount in colony numbers in HCT116 and CACO2 cell traces following ACAA2 knockdown. (Determine 5G). Transwell assays additionally confirmed that ACAA2 knockdown considerably decreased the migratory potential of CRC cells (Figures 5H–J). Taken collectively, these outcomes point out that ACAA2 not solely induces CRC cell proliferation, but in addition promotes CRC cell migration.
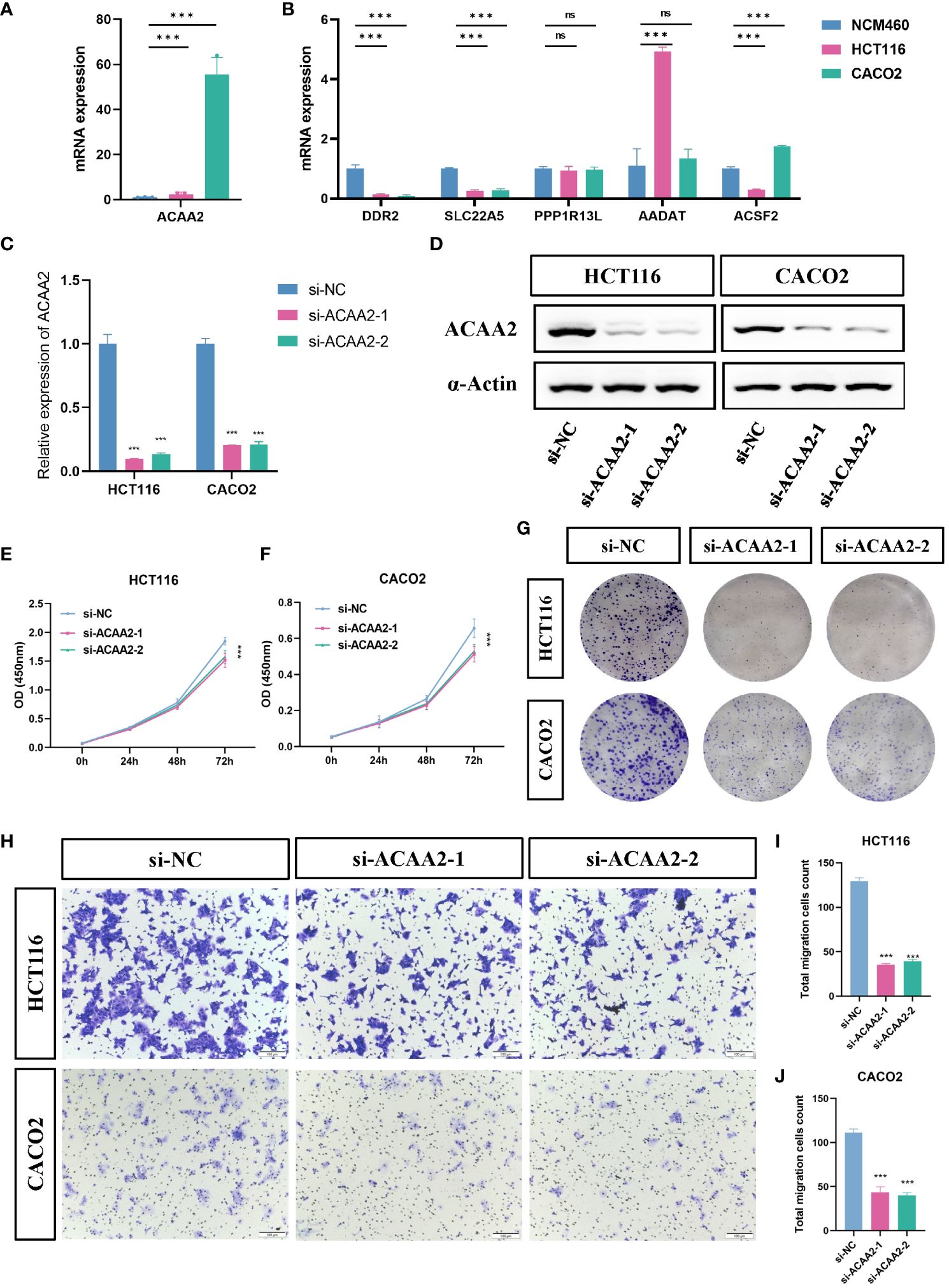
Determine 5 ACAA2 promotes colorectal most cancers development. (A, B) Comparability of the expression ranges of ACAA2, DDR2, SLC22A5, PPP1R13L, AADAT and ACSF2 between NCM460 cells, HCT116 cells, and CACO2 cells. (C) The knockdown effectivity of ACAA2 in HCT116 cells and CACO2. (D) Consultant western blots examined the expression of ACAA2 protein ranges after the downregulation of ACAA2 of HCT116 and CACO2 cell traces. (E, F) The CCK8 assay detected cell viability after decreased ACAA2 expression in HCT116 (E) and CACO2 (F) cell traces. (G) Knockdown of ACAA2 considerably decreased the variety of clones in HCT116 and CACO2 cell traces. (H) The transwell assay detected the migration potential of HCT116 and CACO2 cells after decreased ACAA2 expression. (I-J) Quantification outcomes of numbers of relative migration charges in transwell assay in HCT116 (I) and CACO2 (J) cells. *** p < 0.001, ns, not vital.
3.9 Mutation standing in excessive and low FeFAMscore teams
To discover the mechanisms underlying the FeFAMScore, somatic mutations within the sufferers with CRC within the TCGA cohort have been additional analyzed. As anticipated, extra mutations in high 15 genes have been noticed within the excessive FeFAMscore group than that within the low FeFAMscore group (Figures 6A, B). As well as, co-occurrence and mutual exclusion have been noticed amongst these genes (Figures 6C, D). The forest plot additionally revealed that the BRAF gene, which is mostly considered a possible prognostic danger issue, had extra mutations than the low one, which signifies poor prognostic survival and worse ICI efficacy (Determine 6E). Furthermore, the excessive FeFAMscore teams exhibited a better TMB than the low FeFAMscore teams (Determine 6F). Poor prognosis was additionally demonstrated by excessive TMB mixed with a excessive FeFAMscore (Determine 6G).
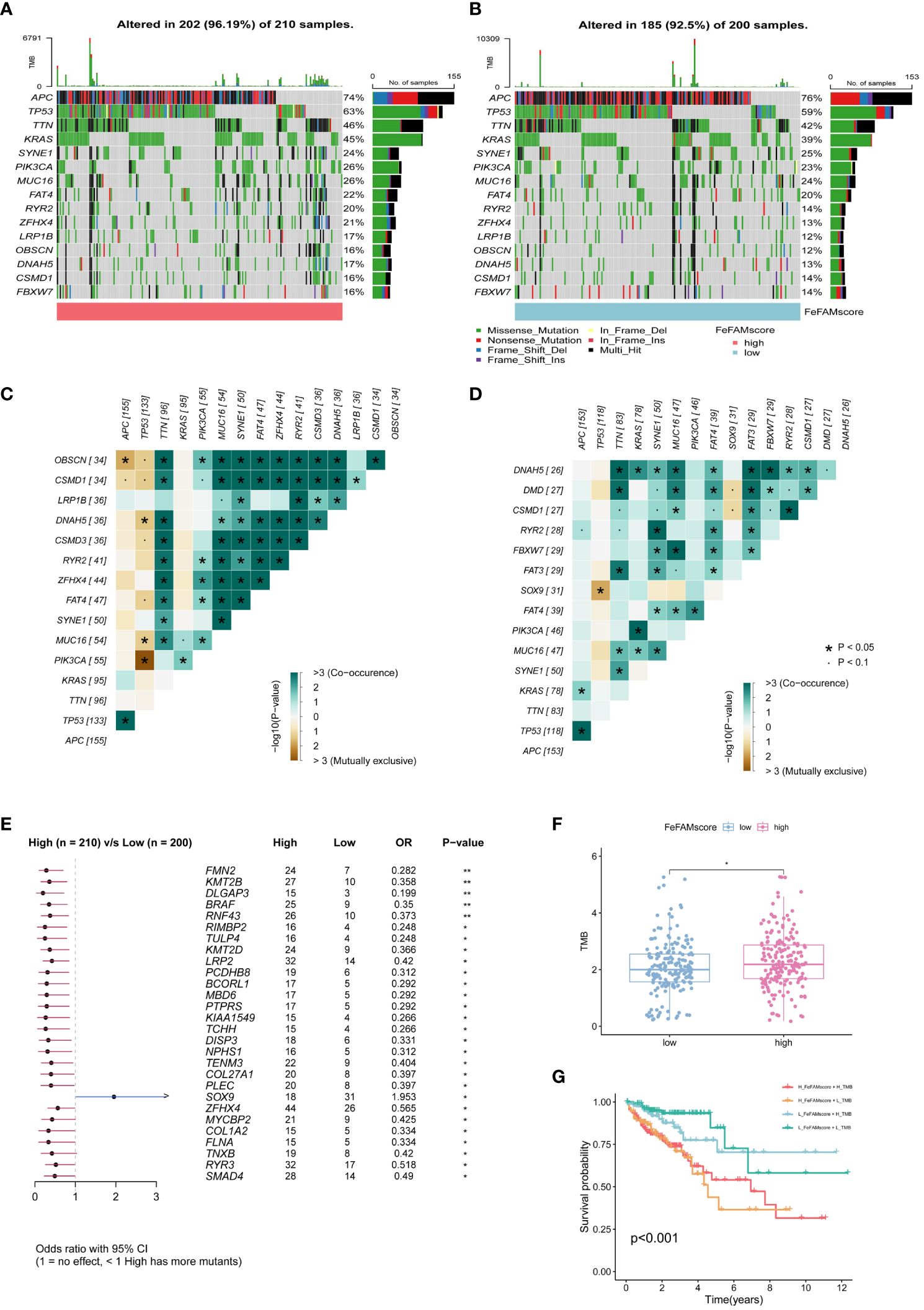
Determine 6 The FeFAMscore associated to the tumor mutation standing. (A, B) Visible abstract exhibiting widespread genetic alterations in low (A) and excessive (B) FeFAMscore teams. (C, D) Interplay impact of genes mutating within the low (C) and excessive (D) FeFAMscore teams. (E) Forest plot gene mutations within the CRC sufferers. (F) The TMB in high and low FeFAMscore teams. (G) Survival evaluation for CRC sufferers measured by each FeFAMscore and TMB utilizing Kaplan-Meier curves. * p < 0.05, ns, not vital. FeFAM, ferroptosis and fatty acid metabolism regulators.
3.10 Immune traits associated to FeFAMscore
We first adopted the ssGSEA algorithm to discover the correlation between tumor-infiltrating immune cells and the FeFAMscore within the coaching cohorts (Supplementary Determine S5A), which indicated that the FeFAMscore had a optimistic relationship with the immune cells. These findings counsel that the excessive FeFAMscore group, regardless of exhibiting a worse prognosis, harbored a better abundance of immunologically activated cells than the low FeFAMscore group. Moreover, the excessive FeFAMscore group demonstrated an elevated presence of immunosuppressive cells similar to MDSCs, macrophages, mast cells, and regulatory T cells (Supplementary Determine S5B). Subsequently, ssGSEA and 6 exterior algorithms, together with CIBERSORT, EPIC, MCPCOUNTER, TIMER, quantiseq, and XCELL have been additional used, yielding related outcomes: the excessive FeFAMscore group had a excessive ImmuneScore, StromaScore, and MicroenvironmentScore (Supplementary Determine S5C). Subsequent, we investigated the cancer-, immune-, and metabolic-related pathways between the 2 teams, which signifies that a excessive FeFAMscore prefers to be enriched in cancer-related and immune-related pathways (Supplementary Determine S5D). Apparently, fatty acid metabolism was considerably activated within the low FeFAMscore group, which can enhance the prognosis. Primarily based on these findings, the excessive FeFAMscore teams in all probability had a number of targets that will profit from particularly focused immunotherapy, although that they had a worse prognosis.
3.11 FeFAMscore predicts CRC response to immunotherapy
We first used TIDE and ESTIMATE algorithms to measure the microenvironment in sufferers with high and low FeFAMscores (Figures 7A–F; Supplementary Determine S5E). The outcomes indicated that the excessive FeFAMscore group was related to excessive immune infiltration however excessive TIDE, CTL.rating, dysfunction, MSI, and PD-L1, which signifies that though these people with poor survival prognosis comprise immunosuppressive cells, that is the primary motive for immune evasion and poor ICI efficacy in these people. This implies that the excessive FeFAMscore group with poor survival could also be a particular goal for activated immune cells, enhancing survival prognosis. Therefore, the submap algorithm was used to evaluate the feasibility of the FeFAMscore in predicting immunotherapy efficacy. These findings affirmed that people within the excessive FeFAMscore group could profit from each anti-PD-1 and anti-CTLA4 therapies (Determine 7G).
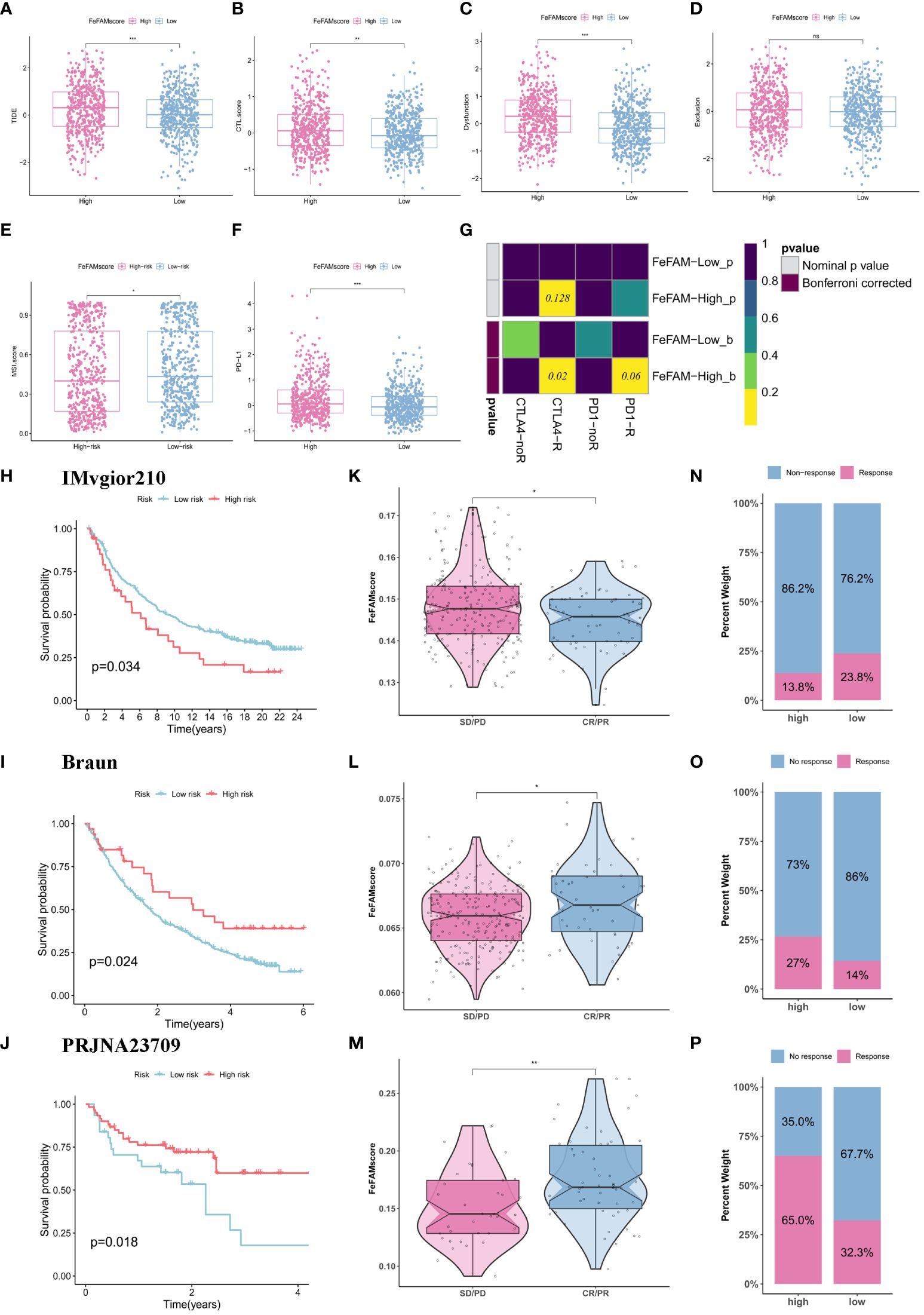
Determine 7 FeFAMscore predicts the response of colorectal to immunotherapy. (A-F) Field plots confirmed the numerous distinction in TIDE (A), CTL.rating (B), Dyfunction (C), Exclusion (D), MDSC (E), PD-L1 (F). (G) The submap algorithm predicts the likelihood of anti-PD1 and anti-CTLA4 immunotherapy response in excessive and low FeFAMscore teams. (H-J) The Kaplan–Meier curve exhibited a major distinction in survival charge between the excessive and low FeFAMscore teams within the IMVgior210 cohort (H), Braun (I), and PRJNA23709 (J). (Ok-M) The Wilcoxon rank-sum check of FeFAMscore variation within the IMVgior210 cohort (Ok), Braun (L), and PRJNA23709 (M). (N-P) The stacked histogram exhibits the distinction in immunotherapy responsiveness between the excessive and low FeFAMscore teams within the IMVgior210 cohort (N), Braun (O), and PRJNA23709 (P). FeFAM, ferroptosis and fatty acid metabolism regulators. * p < 0.05, ** p < 0.01, *** p < 0.001, ns: not vital.
Primarily based on the earlier evaluation, we decided the FeFAMscore within the IMvigor210 cohorts (anti-PD-L1 remedy) (42), Braun cohorts (anti-PD-1 remedy) (43) and PRJNA23709 (anti-PD-1 remedy + anti-CLTA4 remedy) (44). Within the IMvigor210 dataset, sufferers with low FeFAMscores exhibited higher prognoses than these with excessive FeFAMscores. Moreover, people with low FeFAMscores have been more likely to reply favorably to anti-PD-L1 immunotherapy (Figures 7H, Ok, N). Apparently, within the Braun cohort, sufferers with excessive FeFAMscores demonstrated the potential for profit from anti-PD-1 remedy (Figures 7I, L, O). As anticipated, sufferers with renal cell carcinoma and excessive FeFAMscore had considerably improved survival likelihood and have been possible to answer anti-PD-1 remedy. The outcomes have been proven in PRJNA23709 when sufferers with excessive FeFAMscore acquired mixed anti-PD-1 and anti-CTLA4 remedy (Figures 7J, M, P). They not solely drastically improved survival prognosis, but in addition acquired a exceptional response charge to immune remedy. These outcomes counsel that people with a low FeFAMscore could profit from immunotherapy, however that people with a excessive FeFAMscore get hold of wonderful response charges and survival with particular immunotherapies.
3.12 FeFAMscore predicts CRC sensitivity to chemotherapeutic response evaluation
To evaluate probably efficient medicine related to the FeFAMscore, we investigated chemotherapeutic brokers utilizing the “oncoPredict” bundle. With the in contrast variations between two danger FeFAM teams by the Wilcoxion check with p<0.05, we considerably filtered 316 (complete: 545) and 95 (complete: 224) compounds within the CTRP and GDSC, respectively. Subsequent, we investigated the drug intersections within the two databases (Supplementary Determine S6A). Spearman’s technique was used to measure the correlation between FeFAM genes and drug sensitivity. Some genes and medicines, similar to KIF20A and AADAT, interacted antagonistically (Supplementary Figures S6B, C). Nevertheless, the ACSF2 and FABP1 interacted synergistically. To validate the irreplaceable position of FeFAM molecules in chemotherapy, we utilized the FeFAMscore to information chemotherapeutic choice for CRC in scientific apply. Exploring within the CRC associated chemotherapy medicine, the osimertinib, oxaliplatin, gefitinib, eriotinib, navitoclax, and cyclophosphamide are helpful for the sufferers with excessive FeFAMscore, not like irinotecan, niraparib, gemcitabine, niraparib, dabrafenib, and selumetinib (Supplementary Figures S6D–O). These findings underscore the supply of various chemotherapy modalities tailor-made to particular sufferers with CRC, thereby paving the best way for precision chemotherapy and customized therapy approaches.
4 Dialogue
A number of therapeutic modalities, together with surgical procedure, chemotherapy, immunotherapy, radiotherapy, and focused remedy have emerged as key methods in CRC analysis (3, 5, 45, 46). These various therapy approaches symbolize a profound development in CRC administration, reflecting the multifaceted method necessitated by illness complexity (47). Amongst these, immunotherapy is a promising frontier that exploits the intricate interaction between the immune system and malignant cells to elicit therapeutic responses (48). Nevertheless, a subset of sufferers with CRC exhibiting poor mismatch restore or microsatellite instability-high (dMMR/MSI-H) represents a comparatively small fraction, comprising roughly 15% and 4% sufferers with CRC and metastatic colorectal most cancers (mCRC), respectively; a proportion of those sufferers swiftly progress to a state of immune resistance (38, 39). The AJCC staging system is a broadly accepted criterion for scientific administration and encompasses therapeutic decision-making and surveillance methods for CRC. The utility of the AJCC staging system is constrained by the variability in scientific outcomes noticed amongst sufferers labeled throughout the similar stage (49). This will likely not solely lead to overtreatment and undertreatment, but in addition make it troublesome to mirror the sensitivity of immunotherapy and chemotherapy as a result of it doesn’t mirror the TME. To bridge this hole, figuring out novel prognostic and therapeutic targets for CRC is significant.
Cell demise is a regulated course of in cells and could also be associated to metabolism throughout tumor development, metastasis, and drug resistance. Ferroptosis is an iron-related cell demise pathway characterised by lipid peroxide accumulation (50, 51). Fatty acid metabolism performs a pivotal position in tumorigenesis, illness development, and therapy resistance by facilitating augmented lipid synthesis, storage, and catabolism (52). Quite a few research have demonstrated that ferroptosis is considerably correlated with metabolism, significantly lipid metabolism (50–53). As an illustration, ASL4, a fatty acid metabolism-related gene, is induced by the T cell-derived interferon (IFN)-γ to vary the tumor lipid sample, which elevated arachidonic acid (AA) manufacturing to advertise ferroptosis (32). SLC47A1, which regulates lipid transforming and survival throughout ferroptosis, inhibits the anticancer exercise of ferroptosis inducers (54). Furthermore, they work together to modulate drug sensitivity (32, 55). Consequently, the affect of ferroptosis regulators and fatty acid molecules on the TME in addition to their predictive capability for prognosis and response to immunotherapy in CRC, stay unclear.
This research elucidated the genetic and transcriptomic range of FeFAMs throughout 33 most cancers species utilizing a multi-omics method. Related frequencies noticed among the many ferroptosis and fatty acid metabolism regulators point out their interconnectedness. Spearman’s rank correlation coefficient additional demonstrated a robust correlation of ferroptosis and fatty acid metabolism regulators between 33 most cancers species, particularly in CRC (R= -0.93; p<0.001). Moreover, after screening 50 genes utilizing the “limma” bundle and univariate Cox regression evaluation, the sufferers have been stratified into three distinct phenotypes, every exhibiting vital disparities in genetic profiles and immune infiltration throughout the clusters.
We then distinguished three ferroptosis- and fatty acid metabolism-related patterns, named FeFAM clusters A/B/C. The TME traits within the three patterns indicated differential immune cell compositions. FeFAM clusters A and C have been correlated with immune cell abundance; nonetheless, they displayed disparate prognostic outcomes. The FeFAM cluster A processed with excessive immune activate, StromaSocre, PD-L1 expression, CAF expression, and excessive activation of TGF-β signaling pathway proved a robust relation with immune-exclude subtypes, whereas the FeFAM cluster C related to wonderful prognosis and considerable immune cell was considered an immune-inflamed phenotype. Apparently, FeFAM cluster C exhibited the identical ranges of dysfunction and CTL.rating and excessive metabolism ranges, such because the ROS and fatty acids, than FeFAM cluster A, which indicated that the interplay of ferroptosis and fatty acid metabolism could coordinate with T cell dysfunction (32). MDSC density was the very best in FeFAM cluster B, which is outlined because the immune desert subtype. These outcomes display that FeFAM molecules play a significant position within the TME and should set off extrinsic immune escape.
Additional FeFAM molecule characterization in CRC is crucial. Growing options related to FeFAM molecules will facilitate prognosis prediction and immune response analysis in CRC. To keep away from mannequin choice bias and ensure mannequin accuracy, we randomly mixed 10 classical algorithms and ultimately obtained 117 mixed algorithms. Subsequently, we developed FeFAMscore, a machine-learning-based FeFAM-related mannequin, which exhibited one of the best efficiency among the many 117 signatures. Recognizing the heterogeneity usually noticed within the sufferers with CRC, we externally validated the FeFAMscore utilizing 4 extra CRC databases. The very best C-index among the many validations not solely confirmed the collection of the optimum mannequin, CoxBoost mixed with stepwise Cox (backward route), but in addition underscored the accuracy and generalizability of the mannequin. Furthermore, a comparability of 69 revealed CRC signatures confirmed improved accuracy, exhibiting robustness. To validate mannequin accuracy, we recognized ACAA2 as a key FeFAMscore regulator and performed mobile experiments, which revealed that ACAA2 promotes CRC proliferation and migration.
Moreover, FeFAMscore demonstrated a sturdy affiliation with survival outcomes. The opposed prognosis noticed within the excessive FeFAMscore group could also be attributed to improved activation of anti-immune elements, probably fostering a TME conducive to immune evasion. Apparently, the FeFAMscore and tumor immune infiltration extent in CRC is positively correlated. Furthermore, mutations resulting in tumor neoantigens, together with a excessive tumor mutational burden (TMB), enhance tumor immunogenic neoantigen abundance (56). The sufferers with excessive TMB could profit from immunotherapy, however many sufferers don’t obtain the specified outcomes (42). Related outcomes have been noticed on this research. Tide, a computational technique developed by Peng Jiang, fashions T cell dysfunction and exclusion mechanisms of tumor immune evasion by infiltration of cytotoxic T lymphocytes (CTL), confirmed the identical outcomes (42). Nevertheless, owing to the variances in immune-related pathways between the 2 cohorts and the first mechanism of immune evasion being dysfunction, it’s believable that there could also be particular therapeutic advantages for the excessive FeFAMscore group. The Submap algorithm additional supported these outcomes, exhibiting that the excessive FeFAMscore group processes have been extra delicate to anti-PD-1 and anti-CTLA4 therapies. In keeping with the outcomes of earlier research, the predominant mechanism means that each excessive and low fatty acid metabolism can have an effect on the expression stage of iron demise, consequently affecting the mode of motion of CTLs (30, 32). The excessive FeFAMscore group exhibited diminished fatty acid metabolism, probably regulating ferroptosis to enhance CTL sensitivity. To validate these outcomes, we analyzed the FeFAMscores in immunotherapy cohorts receiving anti-PD-L1 remedy, anti-PD-1 remedy, and anti-PD-1 mixed with anti-CTLA4 remedy. Related outcomes have been noticed in these cohorts, additional demonstrating the constraints of TMB and TIDE. Moreover, concerning chemotherapeutic brokers, the FeFAMscore exhibited promising predictive capabilities. Collectively, these findings point out that FeFAMscore holds promise as a worthwhile instrument for formulating efficacious CRC therapy methods.
This research had some limitations. First, the intricate regulatory mechanisms governing ferroptosis and fatty acid metabolism stay unclear. Furthermore, retrospective cohorts sourced from publicly obtainable on-line databases have been used. Giant multicenter potential scientific investigations are warranted to corroborate these findings. Lastly, to validate the predictive utility of the FeFAMscore in immunotherapy response, extra indicators are required, together with potential cohorts of sufferers with glioma present process immunotherapeutic interventions.
In conclusion, by a complete method integrating multicenter evaluation and machine studying algorithms, we developed a steady and dependable prognostic and immunotherapeutic response predictor, the FeFAMscore, for CRC. Notably, the excessive FeFAMscore group demonstrated an elevated sensitivity to anti-PD-1 and anti-CTLA4 therapies. The FeFAMscore holds promise as a worthwhile instrument for tailoring efficacious therapy regimens for CRC.
Knowledge availability assertion
The datasets offered on this research might be present in on-line repositories. The names of the repository/repositories and accession quantity(s) might be discovered under: https://www.ncbi.nlm.nih.gov/, TCGA-COAD, https://www.ncbi.nlm.nih.gov/, TCGA-READ, https://www.ncbi.nlm.nih.gov/, GSE17536, https://www.ncbi.nlm.nih.gov/, GSE17537, https://www.ncbi.nlm.nih.gov/, GSE29621, https://www.ncbi.nlm.nih.gov/, GSE39582, https://www.ncbi.nlm.nih.gov/, GSE38832.
Writer contributions
JCZ: Conceptualization, Knowledge curation, Formal Evaluation, Investigation, Methodology, Venture administration, Sources, Software program, Supervision, Visualization, Writing – authentic draft, Writing – evaluate & enhancing. JYZ: Conceptualization, Knowledge curation, Formal Evaluation, Investigation, Methodology, Validation, Writing – evaluate & enhancing. YL: Funding acquisition, Investigation, Venture administration, Software program, Supervision, Writing – evaluate & enhancing. YZ: Conceptualization, Knowledge curation, Venture administration, Writing – evaluate & enhancing. XZ: Formal Evaluation, Investigation, Supervision, Writing – authentic draft. WC: Knowledge curation, Formal Evaluation, Visualization, Writing – evaluate & enhancing. LY: Funding acquisition, Venture administration, Supervision, Visualization, Writing – evaluate & enhancing. QZ: Knowledge curation, Investigation, Methodology, Sources, Supervision, Validation, Visualization, Writing – evaluate & enhancing.
Funding
The writer(s) declare monetary help was acquired for the analysis, authorship, and/or publication of this text. The Nationwide Pure Science Basis of China (No. 82172816)
Acknowledgments
We appreciated the help of the First Affiliated Hospital of Wenzhou Medical College.
Battle of curiosity
The authors declare that the analysis was performed within the absence of any business or monetary relationships that could possibly be construed as a possible battle of curiosity.
Writer’s be aware
All claims expressed on this article are solely these of the authors and don’t essentially symbolize these of their affiliated organizations, or these of the writer, the editors and the reviewers. Any product that could be evaluated on this article, or declare that could be made by its producer, isn’t assured or endorsed by the writer.
Supplementary materials
The Supplementary Materials for this text might be discovered on-line at: https://www.frontiersin.org/articles/10.3389/fimmu.2024.1416443/full#supplementary-material
References
2. Bretthauer M, Løberg M, Wieszczy P, Kalager M, Emilsson L, Garborg Ok, et al. Impact of colonoscopy screening on dangers of colorectal most cancers and associated demise. N Engl J Med. (2022) 387:1547–56. doi: 10.1056/NEJMoa2208375
PubMed Summary | CrossRef Full Textual content | Google Scholar
4. Cercek A, Lumish M, Sinopoli J, Weiss J, Shia J, Lamendola-Essel M, et al. Pd-1 blockade in mismatch repair-deficient, domestically superior rectal most cancers. N Engl J Med. (2022) 386:2363–76. doi: 10.1056/NEJMoa2201445
PubMed Summary | CrossRef Full Textual content | Google Scholar
5. André T, Shiu Ok-Ok, Kim TW, Jensen BV, Jensen LH, Punt C, et al. Pembrolizumab in microsatellite-instability-high superior colorectal most cancers. N Engl J Med. (2020) 383:2207–18. doi: 10.1056/NEJMoa2017699
PubMed Summary | CrossRef Full Textual content | Google Scholar
6. Hsieh RC-E, Krishnan S, Wu R-C, Boda AR, Liu A, Winkler M, et al. Atr-mediated cd47 and pd-L1 up-regulation restricts radiotherapy-induced immune priming and abscopal responses in colorectal most cancers. Sci Immunol. (2022) 7:eabl9330. doi: 10.1126/sciimmunol.abl9330
PubMed Summary | CrossRef Full Textual content | Google Scholar
7. Liu C, Wang X, Qin W, Tu J, Li C, Zhao W, et al. Combining radiation and the atr inhibitor berzosertib prompts sting signaling and enhances immunotherapy by way of inhibiting shp1 perform in colorectal most cancers. Most cancers Commun (Lond). (2023) 43:435–54. doi: 10.1002/cac2.12412
PubMed Summary | CrossRef Full Textual content | Google Scholar
9. Chan TA, Yarchoan M, Jaffee E, Swanton C, Quezada SA, Stenzinger A, et al. Improvement of tumor mutation burden as an immunotherapy biomarker: utility for the oncology clinic. Ann Oncol. (2019) 30:44–56. doi: 10.1093/annonc/mdy495
PubMed Summary | CrossRef Full Textual content | Google Scholar
10. Cortes-Ciriano I, Lee S, Park W-Y, Kim T-M, Park PJ. A molecular portrait of microsatellite instability throughout a number of cancers. Nat Commun. (2017) 8:15180. doi: 10.1038/ncomms15180
PubMed Summary | CrossRef Full Textual content | Google Scholar
12. Berod L, Friedrich C, Nandan A, Freitag J, Hagemann S, Harmrolfs Ok, et al. De novo fatty acid synthesis controls the destiny between regulatory T and T helper 17 cells. Nat Med. (2014) 20:1327–33. doi: 10.1038/nm.3704
PubMed Summary | CrossRef Full Textual content | Google Scholar
13. Bachem A, Makhlouf C, Binger KJ, de Souza DP, Tull D, Hochheiser Ok, et al. Microbiota-derived short-chain fatty acids promote the reminiscence potential of antigen-activated cd8+ T cells. Immunity. (2019) 51:285–97. doi: 10.1016/j.immuni.2019.06.002
PubMed Summary | CrossRef Full Textual content | Google Scholar
14. Conche C, Finkelmeier F, Pešić M, Nicolas AM, Böttger TW, Kennel KB, et al. Combining ferroptosis induction with mdsc blockade renders main tumours and metastases in liver delicate to immune checkpoint blockade. Intestine. (2023) 72:1774–82. doi: 10.1136/gutjnl-2022-327909
PubMed Summary | CrossRef Full Textual content | Google Scholar
15. Fang H, Wu Y, Chen L, Cao Z, Deng Z, Zhao R, et al. Regulating the obesity-related tumor microenvironment to enhance most cancers immunotherapy. ACS Nano. (2023) 17:4748–63. doi: 10.1021/acsnano.2c11159
PubMed Summary | CrossRef Full Textual content | Google Scholar
16. Zhang Q, Deng T, Zhang H, Zuo D, Zhu Q, Bai M, et al. Adipocyte-derived exosomal mttp suppresses ferroptosis and promotes chemoresistance in colorectal most cancers. Adv Sci (Weinh). (2022) 9:e220335. doi: 10.1002/advs.202203357
17. Qiu B, Zandkarimi F, Bezjian CT, Reznik E, Soni RK, Gu W, et al. Phospholipids with two polyunsaturated fatty acyl tails promote ferroptosis. Cell. (2024) 187:1177–90. doi: 10.1016/j.cell.2024.01.030
PubMed Summary | CrossRef Full Textual content | Google Scholar
18. Chen C, Yang Y, Guo Y, He J, Chen Z, Qiu S, et al. Cyp1b1 inhibits ferroptosis and induces anti-pd-1 resistance by degrading acsl4 in colorectal most cancers. Cell Dying Dis. (2023) 14:271. doi: 10.1038/s41419-023-05803-2
PubMed Summary | CrossRef Full Textual content | Google Scholar
19. Zeng Ok, Li W, Wang Y, Zhang Z, Zhang L, Zhang W, et al. Inhibition of cdk1 overcomes oxaliplatin resistance by regulating acsl4-mediated ferroptosis in colorectal most cancers. Adv Sci (Weinh). (2023) 10:e2301088. doi: 10.1002/advs.202301088
PubMed Summary | CrossRef Full Textual content | Google Scholar
20. Ruan R, Li L, Li X, Huang C, Zhang Z, Zhong H, et al. Unleashing the potential of mixing fgfr inhibitor and immune checkpoint blockade for fgf/fgfr signaling in tumor microenvironment. Mol Most cancers. (2023) 22:60. doi: 10.1186/s12943-023-01761-7
PubMed Summary | CrossRef Full Textual content | Google Scholar
21. Gao Z, Bai Y, Lin A, Jiang A, Zhou C, Cheng Q, et al. Gamma delta T-cell-based immune checkpoint remedy: engaging candidate for antitumor therapy. Mol Most cancers. (2023) 22:31. doi: 10.1186/s12943-023-01722-0
PubMed Summary | CrossRef Full Textual content | Google Scholar
22. Zhou N, Yuan X, Du Q, Zhang Z, Shi X, Bao J, et al. Ferrdb V2: replace of the manually curated database of ferroptosis regulators and ferroptosis-disease associations. Nucleic Acids Res. (2023) 51:D571–D82. doi: 10.1093/nar/gkac935
PubMed Summary | CrossRef Full Textual content | Google Scholar
23. Wilkerson MD, Hayes DN. Consensusclusterplus: A category discovery instrument with confidence assessments and merchandise monitoring. Bioinformatics. (2010) 26:1572–3. doi: 10.1093/bioinformatics/btq170
PubMed Summary | CrossRef Full Textual content | Google Scholar
25. Dhanasekaran R, Deutzmann A, Mahauad-Fernandez WD, Hansen AS, Gouw AM, Felsher DW. The myc oncogene – the grand orchestrator of most cancers progress and immune evasion. Nat Rev Clin Oncol. (2022) 19:23–36. doi: 10.1038/s41571-021-00549-2
PubMed Summary | CrossRef Full Textual content | Google Scholar
26. Zhou B, Lin W, Lengthy Y, Yang Y, Zhang H, Wu Ok, et al. Notch signaling pathway: structure, illness, and therapeutics. Sign Transduct Goal Ther. (2022) 7:95. doi: 10.1038/s41392-022-00934-y
PubMed Summary | CrossRef Full Textual content | Google Scholar
28. Liu Z, Liu L, Weng S, Guo C, Dang Q, Xu H, et al. Machine learning-based integration develops an immune-derived lncrna signature for enhancing outcomes in colorectal most cancers. Nat Commun. (2022) 13:816. doi: 10.1038/s41467-022-28421-6
PubMed Summary | CrossRef Full Textual content | Google Scholar
29. Mariathasan S, Turley SJ, Nickles D, Castiglioni A, Yuen Ok, Wang Y, et al. Tgfβ Attenuates tumour response to pd-L1 blockade by contributing to exclusion of T cells. Nature. (2018) 554:544–8. doi: 10.1038/nature25501
PubMed Summary | CrossRef Full Textual content | Google Scholar
30. Ma X, Xiao L, Liu L, Ye L, Su P, Bi E, et al. Cd36-mediated ferroptosis dampens intratumoral cd8+ T cell effector perform and impairs their antitumor potential. Cell Metab. (2021) 33:1001–12. doi: 10.1016/j.cmet.2021.02.015
PubMed Summary | CrossRef Full Textual content | Google Scholar
31. Wang Y, Tian Q, Hao Y, Yao W, Lu J, Chen C, et al. The kinase advanced mtorc2 promotes the longevity of virus-specific reminiscence cd4+ T cells by stopping ferroptosis. Nat Immunol. (2022) 23:303–17. doi: 10.1038/s41590-021-01090-1
PubMed Summary | CrossRef Full Textual content | Google Scholar
32. Liao P, Wang W, Wang W, Kryczek I, Li X, Bian Y, et al. Cd8+ T cells and fatty acids orchestrate tumor ferroptosis and immunity by way of acsl4. Most cancers Cell. (2022) 40:365–78. doi: 10.1016/j.ccell.2022.02.003
PubMed Summary | CrossRef Full Textual content | Google Scholar
33. Xiong M, Zhuang Ok, Luo Y, Lai Q, Luo X, Fang Y, et al. Kif20a promotes mobile Malignant habits and enhances resistance to chemotherapy in colorectal most cancers by regulation of the jak/stat3 signaling pathway. Getting old (Albany NY). (2019) 11:11905–21. doi: 10.18632/growing old.v11i24
PubMed Summary | CrossRef Full Textual content | Google Scholar
34. Banskota S, Regmi SC, Kim J-A. Nox1 to nox2 swap deactivates ampk and induces invasive phenotype in colon most cancers cells by overexpression of mmp-7. Mol Most cancers. (2015) 14:1–14. doi: 10.1186/s12943-015-0379-0
PubMed Summary | CrossRef Full Textual content | Google Scholar
35. Seenath MM, Roberts D, Cawthorne C, Saunders MP, Armstrong GR, O’Dwyer ST, et al. Reciprocal relationship between expression of hypoxia inducible issue 1alpha (Hif-1alpha) and the pro-apoptotic protein bid in ex vivo colorectal most cancers. Br J Most cancers. (2008) 99:459–63. doi: 10.1038/sj.bjc.6604474
PubMed Summary | CrossRef Full Textual content | Google Scholar
36. Wooden SM, Gill AJ, Brodsky AS, Lu S, Friedman Ok, Karashchuk G, et al. Fatty acid-binding protein 1 is preferentially misplaced in microsatellite instable colorectal carcinomas and is immune modulated by way of the interferon Γ Pathway. Mod Pathol. (2017) 30:123–33. doi: 10.1038/modpathol.2016.170
PubMed Summary | CrossRef Full Textual content | Google Scholar
37. Choi SS, Lee SK, Kim JK, Park H-Ok, Lee E, Jang J, et al. Flightless-1 inhibits er stress-induced apoptosis in colorectal most cancers cells by regulating ca2+ Homeostasis. Exp Mol Med. (2020) 52:940–50. doi: 10.1038/s12276-020-0448-3
PubMed Summary | CrossRef Full Textual content | Google Scholar
38. Chen C, Ma T, Zhang C, Zhang H, Bai L, Kong L, et al. Down-regulation of aquaporin 5-mediated epithelial-mesenchymal transition and anti-metastatic impact by pure product cairicoside E in colorectal most cancers. Mol Carcinog. (2017) 56:2692–705. doi: 10.1002/mc.22712
PubMed Summary | CrossRef Full Textual content | Google Scholar
39. Liu B, Liu Q, Pan S, Huang Y, Qi Y, Li S, et al. The hotair/mir-214/st6gal1 crosstalk modulates colorectal most cancers procession by mediating sialylated C-met by way of jak2/stat3 cascade. J Exp Clin Most cancers Res. (2019) 38:455. doi: 10.1186/s13046-019-1468-5
PubMed Summary | CrossRef Full Textual content | Google Scholar
40. Tune G, Xu S, Zhang H, Wang Y, Xiao C, Jiang T, et al. Timp1 is a prognostic marker for the development and metastasis of colon most cancers by fak-pi3k/akt and mapk pathway. J Exp Clin Most cancers Res. (2016) 35:148. doi: 10.1186/s13046-016-0427-7
PubMed Summary | CrossRef Full Textual content | Google Scholar
41. Fang Y, Shen Z-Y, Zhan Y-Z, Feng X-C, Chen Ok-L, Li Y-S, et al. Cd36 inhibits B-catenin/C-myc-mediated glycolysis by ubiquitination of gpc4 to repress colorectal tumorigenesis. Nat Commun. (2019) 10:3981. doi: 10.1038/s41467-019-11662-3
PubMed Summary | CrossRef Full Textual content | Google Scholar
42. Balar AV, Galsky MD, Rosenberg JE, Powles T, Petrylak DP, Bellmunt J, et al. Atezolizumab as first-line therapy in cisplatin-ineligible sufferers with domestically superior and metastatic urothelial carcinoma: A single-arm, multicentre, part 2 trial. Lancet. (2017) 389:67–76. doi: 10.1016/S0140-6736(16)32455-2
PubMed Summary | CrossRef Full Textual content | Google Scholar
43. Braun DA, Hou Y, Bakouny Z, Ficial M, Sant’ Angelo M, Forman J, et al. Interaction of somatic alterations and immune infiltration modulates response to pd-1 blockade in superior clear cell renal cell carcinoma. Nat Med. (2020) 26:909–18. doi: 10.1038/s41591-020-0839-y
PubMed Summary | CrossRef Full Textual content | Google Scholar
44. Gide TN, Quek C, Menzies AM, Tasker AT, Shang P, Holst J, et al. Distinct immune cell populations outline response to anti-pd-1 monotherapy and anti-pd-1/anti-ctla-4 mixed remedy. Most cancers Cell. (2019) 35. doi: 10.1016/j.ccell.2019.01.003
PubMed Summary | CrossRef Full Textual content | Google Scholar
46. Zhang N, Ng AS, Cai S, Li Q, Yang L, Kerr D. Novel therapeutic methods: concentrating on epithelial-mesenchymal transition in colorectal most cancers. Lancet Oncol. (2021) 22:e358–e68. doi: 10.1016/S1470-2045(21)00343-0
PubMed Summary | CrossRef Full Textual content | Google Scholar
47. Andrei P, Battuello P, Grasso G, Rovera E, Tesio N, Bardelli A. Built-in approaches for precision oncology in colorectal most cancers: the extra you recognize, the higher. Semin Most cancers Biol. (2022) 84:199–213. doi: 10.1016/j.semcancer.2021.04.007
PubMed Summary | CrossRef Full Textual content | Google Scholar
48. Ganesh Ok, Stadler ZK, Cercek A, Mendelsohn RB, Shia J, Segal NH, et al. Immunotherapy in colorectal most cancers: rationale, challenges and potential. Nat Rev Gastroenterol Hepatol. (2019) 16:361–75. doi: 10.1038/s41575-019-0126-x
PubMed Summary | CrossRef Full Textual content | Google Scholar
52. Hoy AJ, Nagarajan SR, Butler LM. Tumour fatty acid metabolism within the context of remedy resistance and weight problems. Nat Rev Most cancers. (2021) 21:753–66. doi: 10.1038/s41568-021-00388-4
PubMed Summary | CrossRef Full Textual content | Google Scholar
54. Lin Z, Liu J, Lengthy F, Kang R, Kroemer G, Tang D, et al. The lipid flippase slc47a1 blocks metabolic vulnerability to ferroptosis. Nat Commun. (2022) 13:7965. doi: 10.1038/s41467-022-35707-2
PubMed Summary | CrossRef Full Textual content | Google Scholar
55. Yang Y, Gu H, Zhang Ok, Guo Z, Wang X, Wei Q, et al. Exosomal acadm sensitizes gemcitabine-resistance by modulating fatty acid metabolism and ferroptosis in pancreatic most cancers. BMC Most cancers. (2023) 23:789. doi: 10.1186/s12885-023-11239-w
PubMed Summary | CrossRef Full Textual content | Google Scholar
56. Chabanon RM, Pedrero M, Lefebvre C, Marabelle A, Soria J-C, Postel-Vinay S. Mutational panorama and sensitivity to immune checkpoint blockers. Clin Most cancers Res. (2016) 22:4309–21. doi: 10.1158/1078-0432.CCR-16-0903
PubMed Summary | CrossRef Full Textual content | Google Scholar

